SARS-CoV-2 virus PCR tiling and sequencing from RNA using EXP-MRT001 and SQK-RBK114.96 (MRT_9186_v114_revL_8Dec2025)
MinION: Protocol
SARS-CoV-2 virus PCR tiling and sequencing from RNA using EXP-MRT001 and SQK-RBK114.96 V MRT_9186_v114_revL_8Dec2025
This is a legacy method. Please refer to our new equivalent method: SARS-CoV-2 virus PCR tiling and sequencing from RNA using SQK-RBK114.96
This protocol:
- Uses extracted RNA samples
- Includes reverse transcription and tiled PCR amplification
- Is for multiplexing 1-96 samples
- Has a library preparation time of ~315 minutes
- Includes fragmentation
- Is compatible with R10.4.1 flow cells
For Research Use Only
This is an Early Access product.
For more information about our Early Access programmes, please see this article on product release phases.
FOR RESEARCH USE ONLY.
Contents
Introduction to the protocol
Library preparation
- 3. Reverse transcription
- 4. PCR
- 5. Addition of rapid barcodes
- 6. Pooling samples and clean-up
- 7. Priming and loading the MinION and GridION Flow Cell
Sequencing and data analysis
Troubleshooting
Overview
This is a legacy method. Please refer to our new equivalent method: SARS-CoV-2 virus PCR tiling and sequencing from RNA using SQK-RBK114.96
This protocol:
- Uses extracted RNA samples
- Includes reverse transcription and tiled PCR amplification
- Is for multiplexing 1-96 samples
- Has a library preparation time of ~315 minutes
- Includes fragmentation
- Is compatible with R10.4.1 flow cells
For Research Use Only
This is an Early Access product.
For more information about our Early Access programmes, please see this article on product release phases.
1. Overview of the protocol
This method will be moved into legacy phase in 2026.
The Midnight RT PCR Expansion (EXP-MRT001) used in this method is scheduled for discontinuation in 2026, and will not be available for purchase from June 2026.
An equivalent method using 3rd party reagents and customer-ordered RT-PCR primers can be found in the documentation space of the Nanopore Communit or by following this link:
Please note, this method is equivalent and uses the same reagents and RT-PCR primers provided in the Midnight RT PCR Expansion (EXP-MRT001).
If you have any concerns or require further support, please contact your Oxford Nanopore Technologies representative.
This protocol is a work in progress and some details are expected to change over time. Please make sure you always use the most recent version of the protocol.
The PCR tiling of SARS-CoV-2 virus with Rapid Barcoding Kit 96 V14 and Midnight RT PCR Expansion (SQK-RBK114.96 and EXP-MRT001) protocol is an updated version of the PCR tiling of SARS-CoV-2 virus with rapid barcoding and Midnight RT PCR Expansion (SQK-RBK110.96 and EXP-MRT001) using our most recent Kit 14 chemistry and an updated downstream analysis.
Introduction to the protocol
To enable support for the rapidly expanding user requests, the team at Oxford Nanopore Technologies have put together an updated workflow based on the ARTIC Network protocols and analysis methods. The protocol uses Oxford Nanopore Technologies' Rapid Barcoding Kit 96 V14 (SQK-RBK114.96) and Midnight RT PCR Expansion (EXP-MRT001) for barcoding and library preparation.
While this protocol is available in the Nanopore Community, we kindly ask users to ensure they are citing the members of the ARTIC network who have been behind the development of these methods.
This protocol is similar to the ARTIC amplicon sequencing protocol for MinION for SARS-CoV-2 v3 (LoCost) by Josh Quick and the method used in Freed et al., 2020. The protocol generates amplicons in a tiled fashion across the whole SARS-CoV-2 genome.
To generate tiled PCR amplicons from the SARS-CoV-2 viral cDNA for use with the Rapid Barcoding Kit 96 V14 (SQK-RBK114.96), primers were designed by Freed *et al.*, 2020 using Primal Scheme. These primers are in the Midnight RT PCR Expansion (EXP-MRT001) and are designed to generate 1.2 kb amplicons. Primer sequences can be found here.
As mutations in SARS-CoV-2 variants emerge, amplicon drop out may be observed. For users wishing to design their own primer spike-ins to address this, we suggest adding to the appropriate primer pool at a final concentration between 3.33 µM and 6.66 µM.
Steps in the sequencing workflow:
Prepare for your experiment
You will need to:
- Extract your RNA.
- Ensure you have your sequencing kit, the correct equipment and reagents.
- Download the software for acquiring and analysing your data.
- Check your flow cell to ensure it has enough pores for a good sequencing run.
Prepare your library
You will need to:
- Reverse transcribe your RNA samples with random hexamers.
- Amplify the samples by tiled PCR using separate primer pools.
- Combine the primer pools.
- Attach Rapid Barcodes supplied in the kit to the DNA ends, pool the samples and SPRI purify.
- Prime the flow cell and load your DNA library into the flow cell.
Sequencing and analysis
You will need to:
- Start a sequencing run using the MinKNOW software, selecting SQK-RBK114.96 in kit selection, which will collect raw data from the device and convert it into basecalled reads.
- (Optional): Perform downstream analysis of the data using the wf-artic analysis workflow integrated within the EPI2ME Labs application.
Before starting
This protocol outlines how to carry out PCR tiling of SARS-CoV-2 viral RNA samples on a 96-well plate using the Rapid Barcoding Kit 96 V14 (SQK-RBK114.96) with the Midnight RT PCR Expansion (EXP-MRT001).
It is required to use total RNA extracted from samples that have been screened by a suitable qPCR assay.
When processing multiple samples at once, we recommend making master mixes with an additional 10% of the volume. We also recommend using a template-free pre-PCR hood for making up the master mixes, and a separate template pre-PCR hood for handling the samples. It is important to clean and/or UV irradiate these hoods between sample batches. Furthermore, to track and monitor cross-contamination events, it is important to run a negative control reaction at the reverse transcription stage using nuclease-free water instead of sample, and carrying this control through the rest of the prep.
All post-PCR procedures must be carried out in a separate area to the pre-PCR preparation, with dedicated equipment for liquid handling in each area.
Compatibility of this protocol
This protocol should only be used in combination with:
- Rapid Barcoding Kit 96 V14 (SQK-RBK114.96)
- Midnight RT PCR Expansion (EXP-MRT001)
- R10.4.1 flow cells (FLO-MIN114)
- Flow Cell Wash Kit (EXP-WSH004)
- MinION Mk1D - MinION Mk1D IT requirements document
- GridION - GridION IT requirements document
2. Equipment and consumables
Materials
- Input RNA in 10 mM Tris-HCl, pH 8.0
- Rapid Barcoding Kit 96 V14 (SQK-RBK114.96)
- Midnight RT PCR Expansion (EXP-MRT001)
Consumables
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- Freshly prepared 80% ethanol in nuclease-free water
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- 1.5 ml Eppendorf DNA LoBind tubes
- 2 ml Eppendorf DNA LoBind tubes
- 5 ml Eppendorf DNA LoBind tubes
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Eppendorf, 0030129504) with PCR seals
- Bovine Serum Albumin (BSA) (50 mg/ml) (e.g. Invitrogen™ UltraPure™ BSA 50 mg/ml, AM2616)
Equipment
- Hula mixer (gentle rotator mixer)
- Magnetic separation rack
- Centrifuge capable of taking 96-well plates
- Microfuge
- Vortex mixer
- Thermal cycler
- Multichannel pipettes suitable for dispensing 0.5–10 μl, 2–20 μl and 20–200 μl, and tips
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- Ice bucket with ice
- Timer
- Qubit fluorometer (or equivalent)
Optional equipment
- Eppendorf 5424 centrifuge (or equivalent)
- PCR hood with UV steriliser (optional but recommended to reduce cross-contamination)
- PCR-Cooler (Eppendorf)
- Stepper pipette and tips
Check your flow cell
We highly recommend that you check the number of pores in your flow cell prior to starting a sequencing experiment. This should be done within 12 weeks of purchasing your MinION/GridION Flow Cells. Oxford Nanopore Technologies will replace any unsed flow cell with fewer than the number of pores listed in the Table below, when the result is reported within two days of performing the flow cell check, and when the storage recommendations have been followed. To do the flow cell check, please follow the instructions in the Flow Cell Check document.
| Flow cell | Minimum number of active pores covered by warranty |
|---|---|
| MinION/GridION Flow Cell | 800 |
For this protocol, you will need your extracted RNA in 8 µl 10 mM Tris-HCl, pH 8.0.
The Rapid Adapter (RA) used in this kit and protocol is not interchangeable with other sequencing adapters.
Rapid Barcoding Kit 96 V14 (SQK-RBK114.96) contents
| Name | Acronym | Cap colour | No. of vials | Fill volume per vial (µl) |
|---|---|---|---|---|
| Rapid Adapter | RA | Green | 2 | 15 |
| Adapter Buffer | ADB | Clear | 1 | 100 |
| AMPure XP Beads | AXP | Amber | 3 | 1,200 |
| Elution Buffer | EB | Black | 1 | 1,500 |
| Sequencing Buffer | SB | Red | 1 | 1,700 |
| Library Beads | LIB | Pink | 1 | 1,800 |
| Library Solution | LIS | White cap, pink label | 1 | 1,800 |
| Flow Cell Flush | FCF | Clear | 1 | 15,500 |
| Flow Cell Tether | FCT | Purple | 2 | 200 |
| Rapid Barcodes | RB01-96 | - | 3 plates | 8 µl per well |
This product contains AMPure XP reagent manufactured by Beckman Coulter, Inc. and can be stored at -20°C with the kit without detriment to reagent stability.
Midnight RT PCR Expansion (EXP-MRT001) contents
| Name | Acronym | Cap colour | Number of vials | Fill volume per vial (µl) |
|---|---|---|---|---|
| LunaScript RT SuperMix | LS RT | Blue | 3 | 500 |
| Q5 HS Master Mix | Q5 | Orange | 6 | 1,500 |
| Midnight Primer Pool A | MP A | White | 3 | 15 |
| Midnight Primer Pool B | MP B | Clear | 3 | 15 |
Midnight Primer sequences
As mutations in SARS-CoV-2 variants emerge, amplicon drop out may be observed. For users wishing to design their own primer spike-ins to address this, we suggest adding to the appropriate primer pool at a final concentration between 3.33 µM and 6.66 µM.
Below are the sequences for the V3 primer scheme used in the Midnight RT PCR Expansion.
Pool A
| Primer name | Primer Sequence |
|---|---|
SARSCoV_1200_1_LEFT | ACCAACCAACTTTCGATCTCTTGT |
SARSCoV_1200_1_RIGHT | GGTTGCATTCATTTGGTGACGC |
SARSCoV_1200_3_LEFT | GGCTTGAAGAGAAGTTTAAGGAAGGT |
SARSCoV_1200_3_RIGHT | GATTGTCCTCACTGCCGTCTTG |
SARSCoV_1200_5_LEFT | ACCTACTAAAAAGGCTGGTGGC |
SARSCoV_1200_5_RIGHT | AGCATCTTGTAGAGCAGGTGGA |
SARSCoV_1200_7_LEFT | ACCTGGTGTATACGTTGTCTTTGG |
SARSCoV_1200_7_RIGHT | GCTGAAATCGGGGCCATTTGTA |
SARSCoV_1200_9_LEFT | AGAAGTTACTGGCGATAGTTGTAATAACT |
SARSCoV_1200_9_RIGHT | TGCTGATATGTCCAAAGCACCA |
SARSCoV_1200_11_LEFT | AGACACCTAAGTATAAGTTTGTTCGCA |
SARSCoV_1200_11_RIGHT | GCCCACATGGAAATGGCTTGAT |
SARSCoV_1200_13_LEFT | ACCTCTTACAACAGCAGCCAAAC |
SARSCoV_1200_13_RIGHT | CGTCCTTTTCTTGGAAGCGACA |
SARSCoV_1200_15_LEFT | TTTTAAGGAATTACTTGTGTATGCTGCT |
SARSCoV_1200_15_RIGHT | ACACACAACAGCATCGTCAGAG |
SARSCoV_1200_17_LEFT | TCAAGCTTTTTGCAGCAGAAACG |
SARSCoV_1200_17_RIGHT | CCAAGCAGGGTTACGTGTAAGG |
SARSCoV_1200_19_LEFT | GGCACATGGCTTTGAGTTGACA |
SARSCoV_1200_19_RIGHT | CCTGTTGTCCATCAAAGTGTCCC |
SARSCoV_1200_21_LEFT | TCTGTAGTTTCTAAGGTTGTCAAAGTGA |
SARSCoV_1200_21_RIGHT | GCAGGGGGTAATTGAGTTCTGG |
21_right_spike | GTGTATGATTGAGTTCTGGTTGTAAG |
SARSCoV_1200_23_LEFT | ACTTTAGAGTCCAACCAACAGAATCT |
23_left_spike | ACTTTAGAGTTCAACCAACAGAATCT |
SARSCoV_1200_23_RIGHT | TGACTAGCTACACTACGTGCCC |
SARSCoV_1200_25_LEFT | TGCTGCTACTAAAATGTCAGAGTGT |
SARSCoV_1200_25_RIGHT | CATTTCCAGCAAAGCCAAAGCC |
SARSCoV_1200_27_LEFT | TGGATCACCGGTGGAATTGCTA |
SARSCoV_1200_27_RIGHT | TGTTCGTTTAGGCGTGACAAGT |
SARSCoV_1200_29_LEFT | TGAGGGAGCCTTGAATACACCA |
SARSCoV_1200_29_RIGHT | TAGGCAGCTCTCCCTAGCATTG |
Pool B
| Primer name | Primer sequences |
|---|---|
SARSCoV_1200_2_LEFT | CCATAATCAAGACTATTCAACCAAGGGT |
SARSCoV_1200_2_RIGHT | ACAGGTGACAATTTGTCCACCG |
SARSCoV_1200_4_LEFT | GGAATTTGGTGCCACTTCTGCT |
SARSCoV_1200_4_RIGHT | CCTGACCCGGGTAAGTGGTTAT |
SARSCoV_1200_6_LEFT | ACTTCTATTAAATGGGCAGATAACAACTG |
SARSCoV_1200_6_RIGHT | GATTATCCATTCCCTGCGCGTC |
SARSCoV_1200_8_LEFT | CAATCATGCAATTGTTTTTCAGCTATTTTG |
SARSCoV_1200_8_RIGHT | TGACTTTTTGCTACCTGCGCAT |
SARSCoV_1200_10_LEFT | TTTACCAGGAGTTTTCTGTGGTGT |
SARSCoV_1200_10_RIGHT | TGGGCCTCATAGCACATTGGTA |
SARSCoV_1200_12_LEFT | ATGGTGCTAGGAGAGTGTGGAC |
SARSCoV_1200_12_RIGHT | GGATTTCCCACAATGCTGATGC |
SARSCoV_1200_14_LEFT | ACAGGCACTAGTACTGATGTCGT |
SARSCoV_1200_14_RIGHT | GTGCAGCTACTGAAAAGCACGT |
SARSCoV_1200_16_LEFT | ACAACACAGACTTTATGAGTGTCTCT |
SARSCoV_1200_16_RIGHT | CTCTGTCAGACAGCACTTCACG |
SARSCoV_1200_18_LEFT | GCACATAAAGACAAATCAGCTCAATGC |
SARSCoV_1200_18_RIGHT | TGTCTGAAGCAGTGGAAAAGCA |
SARSCoV_1200_20_LEFT | ACAATTTGATACTTATAACCTCTGGAACAC |
SARSCoV_1200_20_RIGHT | GATTAGGCATAGCAACACCCGG |
SARSCoV_1200_22_LEFT | GTGATGTTCTTGTTAACAACTAAACGAACA |
SARSCoV_1200_22_RIGHT | AACAGATGCAAATCTGGTGGCG |
22_right_spike | AACAGATGCAAATTTGGTGGCG |
SARSCoV_1200_24_LEFT | GCTGAACATGTCAACAACTCATATGA |
24_left_spike | GCTGAATATGTCAACAACTCATATGA |
SARSCoV_1200_24_RIGHT | ATGAGGTGCTGACTGAGGGAAG |
SARSCoV_1200_26_LEFT | GCCTTGAAGCCCCTTTTCTCTA |
SARSCoV_1200_26_RIGHT | AATGACCACATGGAACGCGTAC |
SARSCoV_1200_28_LEFT | TTTGTGCTTTTTAGCCTTTCTGCT |
SARSCoV_1200_28_RIGHT | GTTTGGCCTTGTTGTTGTTGGC |
SARSCoV_1200_28_LEFT_27837T | TTTGTGCTTTTTAGCCTTTCTGTT |
Rapid barcode sequences
| Component | Sequence |
|---|---|
| RB01 | AAGAAAGTTGTCGGTGTCTTTGTG |
| RB02 | TCGATTCCGTTTGTAGTCGTCTGT |
| RB03 | GAGTCTTGTGTCCCAGTTACCAGG |
| RB04 | TTCGGATTCTATCGTGTTTCCCTA |
| RB05 | CTTGTCCAGGGTTTGTGTAACCTT |
| RB06 | TTCTCGCAAAGGCAGAAAGTAGTC |
| RB07 | GTGTTACCGTGGGAATGAATCCTT |
| RB08 | TTCAGGGAACAAACCAAGTTACGT |
| RB09 | AACTAGGCACAGCGAGTCTTGGTT |
| RB10 | AAGCGTTGAAACCTTTGTCCTCTC |
| RB11 | GTTTCATCTATCGGAGGGAATGGA |
| RB12 | CAGGTAGAAAGAAGCAGAATCGGA |
| RB13 | AGAACGACTTCCATACTCGTGTGA |
| RB14 | AACGAGTCTCTTGGGACCCATAGA |
| RB15 | AGGTCTACCTCGCTAACACCACTG |
| RB16 | CGTCAACTGACAGTGGTTCGTACT |
| RB17 | ACCCTCCAGGAAAGTACCTCTGAT |
| RB18 | CCAAACCCAACAACCTAGATAGGC |
| RB19 | GTTCCTCGTGCAGTGTCAAGAGAT |
| RB20 | TTGCGTCCTGTTACGAGAACTCAT |
| RB21 | GAGCCTCTCATTGTCCGTTCTCTA |
| RB22 | ACCACTGCCATGTATCAAAGTACG |
| RB23 | CTTACTACCCAGTGAACCTCCTCG |
| RB24 | GCATAGTTCTGCATGATGGGTTAG |
| RB25 | GTAAGTTGGGTATGCAACGCAATG |
| RB26 | CATACAGCGACTACGCATTCTCAT |
| RB27 | CGACGGTTAGATTCACCTCTTACA |
| RB28 | TGAAACCTAAGAAGGCACCGTATC |
| RB29 | CTAGACACCTTGGGTTGACAGACC |
| RB30 | TCAGTGAGGATCTACTTCGACCCA |
| RB31 | TGCGTACAGCAATCAGTTACATTG |
| RB32 | CCAGTAGAAGTCCGACAACGTCAT |
| RB33 | CAGACTTGGTACGGTTGGGTAACT |
| RB34 | GGACGAAGAACTCAAGTCAAAGGC |
| RB35 | CTACTTACGAAGCTGAGGGACTGC |
| RB36 | ATGTCCCAGTTAGAGGAGGAAACA |
| RB37 | GCTTGCGATTGATGCTTAGTATCA |
| RB38 | ACCACAGGAGGACGATACAGAGAA |
| RB39 | CCACAGTGTCAACTAGAGCCTCTC |
| RB40 | TAGTTTGGATGACCAAGGATAGCC |
| RB41 | GGAGTTCGTCCAGAGAAGTACACG |
| RB42 | CTACGTGTAAGGCATACCTGCCAG |
| RB43 | CTTTCGTTGTTGACTCGACGGTAG |
| RB44 | AGTAGAAAGGGTTCCTTCCCACTC |
| RB45 | GATCCAACAGAGATGCCTTCAGTG |
| RB46 | GCTGTGTTCCACTTCATTCTCCTG |
| RB47 | GTGCAACTTTCCCACAGGTAGTTC |
| RB48 | CATCTGGAACGTGGTACACCTGTA |
| RB49 | ACTGGTGCAGCTTTGAACATCTAG |
| RB50 | ATGGACTTTGGTAACTTCCTGCGT |
| RB51 | GTTGAATGAGCCTACTGGGTCCTC |
| RB52 | TGAGAGACAAGATTGTTCGTGGAC |
| RB53 | AGATTCAGACCGTCTCATGCAAAG |
| RB54 | CAAGAGCTTTGACTAAGGAGCATG |
| RB55 | TGGAAGATGAGACCCTGATCTACG |
| RB56 | TCACTACTCAACAGGTGGCATGAA |
| RB57 | GCTAGGTCAATCTCCTTCGGAAGT |
| RB58 | CAGGTTACTCCTCCGTGAGTCTGA |
| RB59 | TCAATCAAGAAGGGAAAGCAAGGT |
| RB60 | CATGTTCAACCAAGGCTTCTATGG |
| RB61 | AGAGGGTACTATGTGCCTCAGCAC |
| RB62 | CACCCACACTTACTTCAGGACGTA |
| RB63 | TTCTGAAGTTCCTGGGTCTTGAAC |
| RB64 | GACAGACACCGTTCATCGACTTTC |
| RB65 | TTCTCAGTCTTCCTCCAGACAAGG |
| RB66 | CCGATCCTTGTGGCTTCTAACTTC |
| RB67 | GTTTGTCATACTCGTGTGCTCACC |
| RB68 | GAATCTAAGCAAACACGAAGGTGG |
| RB69 | TACAGTCCGAGCCTCATGTGATCT |
| RB70 | ACCGAGATCCTACGAATGGAGTGT |
| RB71 | CCTGGGAGCATCAGGTAGTAACAG |
| RB72 | TAGCTGACTGTCTTCCATACCGAC |
| RB73 | AAGAAACAGGATGACAGAACCCTC |
| RB74 | TACAAGCATCCCAACACTTCCACT |
| RB75 | GACCATTGTGATGAACCCTGTTGT |
| RB76 | ATGCTTGTTACATCAACCCTGGAC |
| RB77 | CGACCTGTTTCTCAGGGATACAAC |
| RB78 | AACAACCGAACCTTTGAATCAGAA |
| RB79 | TCTCGGAGATAGTTCTCACTGCTG |
| RB80 | CGGATGAACATAGGATAGCGATTC |
| RB81 | CCTCATCTTGTGAAGTTGTTTCGG |
| RB82 | ACGGTATGTCGAGTTCCAGGACTA |
| RB83 | TGGCTTGATCTAGGTAAGGTCGAA |
| RB84 | GTAGTGGACCTAGAACCTGTGCCA |
| RB85 | AACGGAGGAGTTAGTTGGATGATC |
| RB86 | AGGTGATCCCAACAAGCGTAAGTA |
| RB87 | TACATGCTCCTGTTGTTAGGGAGG |
| RB88 | TCTTCTACTACCGATCCGAAGCAG |
| RB89 | ACAGCATCAATGTTTGGCTAGTTG |
| RB90 | GATGTAGAGGGTACGGTTTGAGGC |
| RB91 | GGCTCCATAGGAACTCACGCTACT |
| RB92 | TTGTGAGTGGAAAGATACAGGACC |
| RB93 | AGTTTCCATCACTTCAGACTTGGG |
| RB94 | GATTGTCCTCAAACTGCCACCTAC |
| RB95 | CCTGTCTGGAAGAAGAATGGACTT |
| RB96 | CTGAACGGTCATAGAGTCCACCAT |
3. Reverse transcription
Materials
- Input RNA in 10 mM Tris-HCl, pH 8.0
- LunaScript RT SuperMix (LS RT)
Consumables
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Eppendorf, 0030129504) with PCR seals
Equipment
- Multichannel pipettes suitable for dispensing 0.5–10 μl, 2–20 μl and 20–200 μl, and tips
- Thermal cycler
- Centrifuge capable of taking 96-well plates
- Ice bucket with ice
Optional equipment
- PCR-Cooler (Eppendorf)
- PCR hood with UV steriliser (optional but recommended to reduce cross-contamination)
- Stepper pipette and tips
Keep the RNA sample on ice as much as possible to prevent nucleolytic degradation, which may affect sensitivity.
In a clean pre-PCR hood, place a fresh 96-well plate (RT plate) into a PCR Cooler (if using). Using a stepper pipette, or multichannel pipette, add 2 µl of LunaScript RT SuperMix (LS RT) per well.
Depending on the number of samples, fill each well per column as follows:
| Plate location | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Columns | 1-3 | 1-6 | 1-12 |
To each well containing LunaScript RT SuperMix (LS RT), add 8 µl of sample and gently mix by pipetting. If adding less than 8 µl, make up the rest of the volume with nuclease-free water.
Example for X48 samples:
We recommend having a negative control and a positive control for every plate of samples.
Seal the RT plate and spin down.
Incubate the samples in the thermal cycler using the following program:
| Step | Temperature | Time | Cycles |
|---|---|---|---|
| Primer annealing | 25°C | 2 min | 1 |
| cDNA synthesis | 55°C | 10 min | 1 |
| Heat inactivation | 95°C | 1 min | 1 |
| Hold | 4°C | ∞ |
While the reverse transcription reaction is running, prepare the master mixes as described in the next section.
4. PCR
Materials
- Q5 HS Master Mix (Q5)
- Midnight Primer Pool A (MP A)
- Midnight Primer Pool B (MP B)
Consumables
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- 1.5 ml Eppendorf DNA LoBind tubes
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Eppendorf, 0030129504) with PCR seals
Equipment
- Multichannel pipettes suitable for dispensing 0.5–10 μl, 2–20 μl and 20–200 μl, and tips
- P1000 pipette and tips
- P200 pipette and tips
- Thermal cycler
- Microfuge
- Centrifuge capable of taking 96-well plates
- Ice bucket with ice
Optional equipment
- PCR-Cooler (Eppendorf)
- PCR hood with UV steriliser (optional but recommended to reduce cross-contamination)
- Stepper pipette and tips
Primer design
To generate tiled PCR amplicons from the SARS-CoV-2 viral cDNA, primers were designed by Freed et al., 2020 using Primal Scheme. These primers are designed to generate 1200 bp amplicons that overlap by approximately 20 bp. These primer sequences can be found here.
We recommend handling the primers in a clean template-free PCR hood.
In the template-free pre-PCR hood, prepare the following master mixes in Eppendorf DNA LoBind tubes and mix thoroughly as follows:
Volume per sample:
| Reagent | Pool A | Pool B |
|---|---|---|
| Nuclease-free water | 3.7 µl | 3.7 µl |
| Midnight Primer Pool A (MP A) | 0.05 µl | - |
| Midnight Primer Pool B (MP B) | - | 0.05 µl |
| Q5 HS Master Mix (Q5) | 6.25 µl | 6.25 µl |
| Total | 10 µl | 10 µl |
For x24 samples:
| Reagent | Pool A | Pool B |
|---|---|---|
| Nuclease-free water | 102 µl | 102 µl |
| Midnight Primer Pool A (MP A) | 2 µl | - |
| Midnight Primer Pool B (MP B) | - | 2 µl |
| Q5 HS Master Mix (Q5) | 172 µl | 172 µl |
| Total | 276 µl | 276 µl |
For x48 samples:
| Reagent | Pool A | Pool B |
|---|---|---|
| Nuclease-free water | 203 µl | 203 µl |
| Midnight Primer Pool A (MP A) | 3 µl | - |
| Midnight Primer Pool B (MP B) | - | 3 µl |
| Q5 HS Master Mix (Q5) | 344 µl | 344 µl |
| Total | 550 µl | 550 µl |
For x96 samples:
| Reagent | Pool A | Pool B |
|---|---|---|
| Nuclease-free water | 407 µl | 407 µl |
| Midnight Primer Pool A (MP A) | 6 µl | - |
| Midnight Primer Pool B (MP B) | - | 6 µl |
| Q5 HS Master Mix (Q5) | 687 µl | 687 µl |
| Total | 1,100 µl | 1,100 µl |
Using a stepper pipette or a multichannel pipette, aliquot 10 µl of Pool A and Pool B into a clean 96-well plate(s) as follows:
| Plate location | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Columns | Pool A: 1-3 Pool B: 4-6 | Pool A: 1-6 Pool B: 7-12 | Pool A: 1-12 Pool B: 1-12 |
Note: For X96 samples, Pool A is a separate plate to Pool B.
Using a multichannel pipette, transfer 2.5 μl of each RT reaction from the RT plate to the corresponding well for both Pool A and Pool B in the PCR plate(s), taking care not to cross-contaminate different wells. Mix by pipetting the contents of each well up and down.
There should be two PCR reactions per sample.
Example for X48 samples:
Mix by pipetting the contents of each well up and down.
Carry forward the negative control from the reverse transcription reaction to monitor cross-contamination events.
We recommend having a negative control and a positive control for every plate of samples.
Seal the plate(s) and spin down briefly.
Incubate using the following program, with the heated lid set to 105°C:
| Step | Temperature | Time | Cycles |
|---|---|---|---|
| Initial denaturation | 98°C | 30 sec | 1 |
| Denaturation Annealing and extension | 98°C 61°C 65°C | 15 sec 2 min 3 min | 35 |
| Hold | 4°C | ∞ |
If necessary, the protocol can be paused at this point. The samples should be kept at 4°C and can be stored overnight.
5. Addition of rapid barcodes
Materials
- Rapid Barcode Plate (RB01-96)
Consumables
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Eppendorf, 0030129504) with PCR seals
Equipment
- Multichannel pipettes suitable for dispensing 2–20 μl and 20–200 μl, and tips
- Thermal cycler
- Centrifuge capable of taking 96-well plates
Check your flow cell.
We recommend performing a flow cell check before the addition of barcodes to samples to ensure you have a flow cell with enough pores for a good sequencing run.
See the flow cell check document for more information.
Spin down the Rapid Barcode Plate and PCR reactions prior to opening to collect material in the bottom of the wells.
The wells of the barcoding plate are intended for single use only. Please ensure your barcode well is sealed before use, and do not reuse the barcode well once pierced/opened.
Using a multichannel pipette or stepper pipette, transfer 2.5 μl nuclease-free water to the wells of a fresh 96-well plate (Barcode Attachment Plate).
Depending on the number of samples, aliquot into each well of the columns as follows:
| Plate location | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Columns | 1-3 | 1-6 | 1-12 |
Using a multichannel pipette, transfer the entire contents of each well of PCR Pool B to the corresponding well of PCR Pool A and mix by pipetting.
Depending on the number of samples, Pool B columns will correspond to different Pool A columns.
| No. of samples | Pool B column | Corresponding Pool A column |
|---|---|---|
| X24 | 4 5 6 | 1 2 3 |
| X48 | 7 8 9 10 11 12 | 1 2 3 4 5 6 |
| X96 | 1 2 3 4 5 6 7 8 9 10 11 12 | 1 2 3 4 5 6 7 8 9 10 11 12 |
Example for X48 samples:
Using a multichannel pipette, transfer 5 µl from each well of PCR Pool A (now containing pooled PCR products) to the corresponding well of the Barcode Attachment Plate and mix by pipetting.
Depending on the number of samples, PCR Pool A will be in each well of the following columns:
| Plate location | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Columns | 1-3 | 1-6 | 1-12 |
Example for X48 samples:
Using a multichannel pipette, transfer 2.5 μl from the Rapid Barcode Plate to the corresponding well of the Barcode Attachment Plate, taking care not to cross-contaminate different wells. Mix by pipetting.
Depending on the number of samples, aliquot into each well of the columns as follows:
| Plate location | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Columns | 1-3 | 1-6 | 1-12 |
Example for X48 samples:
Samples must be thoroughly mixed.
Seal the Barcode Attachment Plate and spin down.
Incubate the plate in a thermal cycler at 30°C for 2 minutes and then at 80°C for 2 minutes.
6. Pooling samples and clean-up
Materials
- AMPure XP Beads (AXP)
- Elution Buffer from the Oxford Nanopore kit (EB)
- Rapid Adapter (RA)
- Adapter Buffer (ADB)
Consumables
- Freshly prepared 80% ethanol in nuclease-free water
- 1.5 ml Eppendorf DNA LoBind tubes
- 5 ml Eppendorf DNA LoBind tubes
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- Qubit™ Assay Tubes (Invitrogen, Q32856)
Equipment
- Microfuge
- Centrifuge capable of taking 96-well plates
- Hula mixer (gentle rotator mixer)
- Magnetic separation rack
- Ice bucket with ice
- P1000 pipette and tips
- P200 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- Qubit fluorometer plate reader (or equivalent for QC check)
Briefly spin down the Barcode Attachment Plate to collect the liquid at the bottom of the wells prior to opening.
Pool the barcoded samples in a 1.5 ml Eppendorf DNA LoBind tube.
We expect to have about ~10 µl per sample.
| X24 samples | X48 samples | X96 samples | |
|---|---|---|---|
| Total volume | ~240 µl | ~480 µl | ~960 µl |
Mix pooled samples by vortexing.
Pooled barcoded samples must be thoroughly mixed.
Transfer half of the barcoded pooled sample to a clean 1.5 ml Eppendorf DNA LoBind tube.
Per sample, we expect to take forward ~5 µl.
| X24 samples | X48 samples | X96 samples | |
|---|---|---|---|
| Example volume | 120 µl | 240 µl | 480 µl |
Resuspend the AMPure XP Beads (AXP) by vortexing.
To the pooled barcoded sample, add an equal volume of resuspended AMPure XP Beads (AXP, or SPRI) and mix by pipetting.
| Example volume | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Volume of 1X AXP | 120 µl | 240 µl | 480 µl |
Incubate on a Hula mixer (rotator mixer) for 5 minutes at room temperature.
Prepare at least 3 ml of fresh 80% ethanol in nuclease-free water.
Spin down the sample and pellet on a magnet. Keep the tube on the magnet, and pipette off the supernatant when clear and colourless.
Keep the tube on the magnet and wash the beads with 1 ml of freshly-prepared 80% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
Repeat the previous step.
Briefly spin down and place the tube back on the magnet. Pipette off any residual ethanol. Allow to dry for 30 seconds, but do not dry the pellet to the point of cracking.
Remove the tube from the magnetic rack and resuspend the pellet by pipetting in 15 µl Elution Buffer (EB). Incubate for 10 minutes at room temperature.
Pellet the beads on a magnet until the eluate is clear and colourless.
Remove and retain 15 µl of eluate containing the DNA library into a clean 1.5 ml Eppendorf DNA LoBind tube.
Quantify the DNA concentration by using the Qubit dsDNA HS Assay Kit.
Take forward 11 µl of your eluted DNA library.
In a fresh 1.5 ml Eppendorf DNA LoBind tube, dilute the Rapid Adapter (RA) as follows and pipette mix:
| Reagent | Volume |
|---|---|
| Rapid Adapter (RA) | 1.5 μl |
| Adapter Buffer (ADB) | 3.5 μl |
| Total | 5 μl |
Add 1 µl of the diluted Rapid Adapter (RA) to the barcoded DNA.
Mix gently by flicking the tubes, and spin down.
Incubate the reaction for 5 minutes at room temperature.
The prepared library is used for loading into the flow cell. Store the library on ice until ready to load.
7. Priming and loading the MinION and GridION Flow Cell
Materials
- Flow Cell Flush (FCF)
- Flow Cell Tether (FCT)
- Library Solution (LIS)
- Library Beads (LIB)
- Sequencing Buffer (SB)
Consumables
- 1.5 ml Eppendorf DNA LoBind tubes
- MinION/GridION Flow Cell
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- Bovine Serum Albumin (BSA) (50 mg/ml) (e.g. Invitrogen™ UltraPure™ BSA 50 mg/ml, AM2616)
Equipment
- MinION or GridION device
- MinION/GridION Flow Cell Light Shield
- P1000 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
Please note, this kit is only compatible with R10.4.1 flow cells (FLO-MIN114).
Take the flow cell out of the fridge and leave it at room temperature for 20 minutes. This will improve visibility of the array during priming and sample loading.
Priming and loading a flow cell
We recommend all new users watch the 'Priming and loading your flow cell' video before your first run.
Using the Library Solution
For most sequencing experiments, use the Library Beads (LIB) for loading your library onto the flow cell. However, for viscous libraries it may be difficult to load with the beads and may be appropriate to load using the Library Solution (LIS).
Thaw the Sequencing Buffer (SB), Library Beads (LIB) or Library Solution (LIS, if using), Flow Cell Tether (FCT) and Flow Cell Flush (FCF) at room temperature before mixing by vortexing. Then spin down and store on ice.
For optimal sequencing performance and improved output on MinION R10.4.1 flow cells (FLO-MIN114), add Bovine Serum Albumin (BSA) to the flow cell priming mix at a final concentration of 0.2 mg/ml.
Note: We do not recommend using any other albumin type (e.g. recombinant human serum albumin).
Prepare the flow cell priming mix with BSA in a suitable tube for the number of flow cells to flush. Once combined, mix well by pipette mixing.
| Reagents | Volume per flow cell |
|---|---|
| Flow Cell Flush (FCF) | 1,170 µl |
| Bovine Serum Albumin (BSA) at 50 mg/ml | 5 µl |
| Flow Cell Tether (FCT) | 30 µl |
| Total volume | 1,205 µl |
Open the MinION or GridION device lid and slide the flow cell under the clip. Press down firmly on the priming port cover to ensure correct thermal and electrical contact.
Complete a flow cell check to assess the number of pores available before loading the library.
This step can be omitted if the flow cell has been checked previously.
See the flow cell check document for more information.
Slide the flow cell priming port cover clockwise to open the priming port.
Take care when drawing back buffer from the flow cell. Do not remove more than 20-30 µl, and make sure that the array of pores are covered by buffer at all times. Introducing air bubbles into the array can irreversibly damage pores.
After opening the priming port, check for a small air bubble under the cover. Draw back a small volume to remove any bubbles:
- Set a P1000 pipette to 200 µl
- Insert the tip into the priming port
- Turn the wheel until the dial shows 220-230 µl, to draw back 20-30 µl, or until you can see a small volume of buffer entering the pipette tip
Note: Visually check that there is continuous buffer from the priming port across the sensor array.
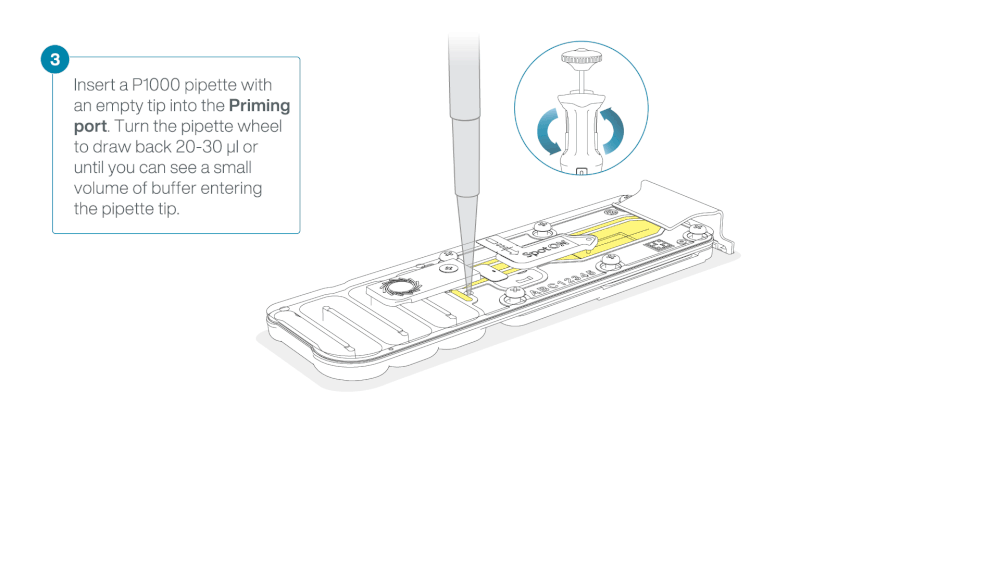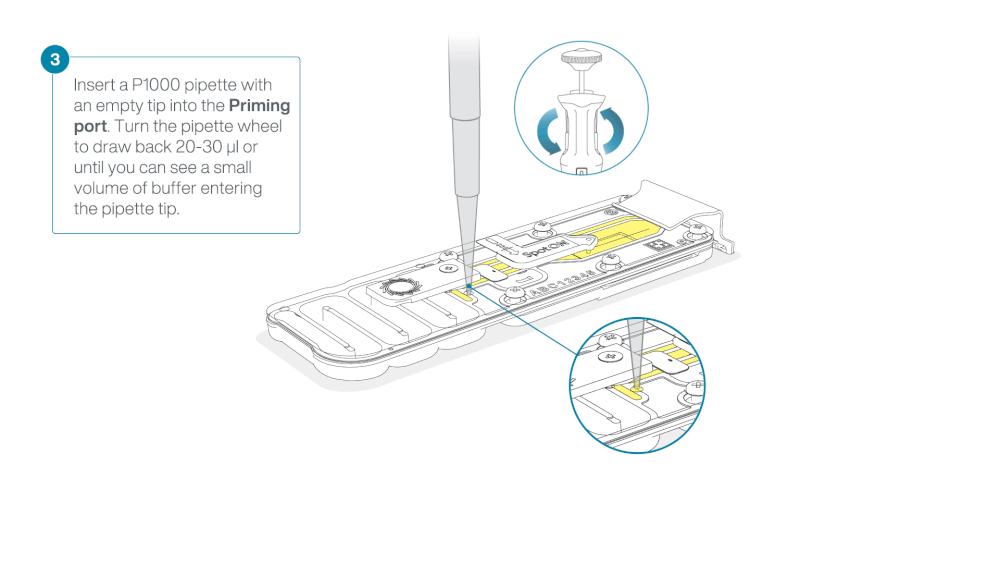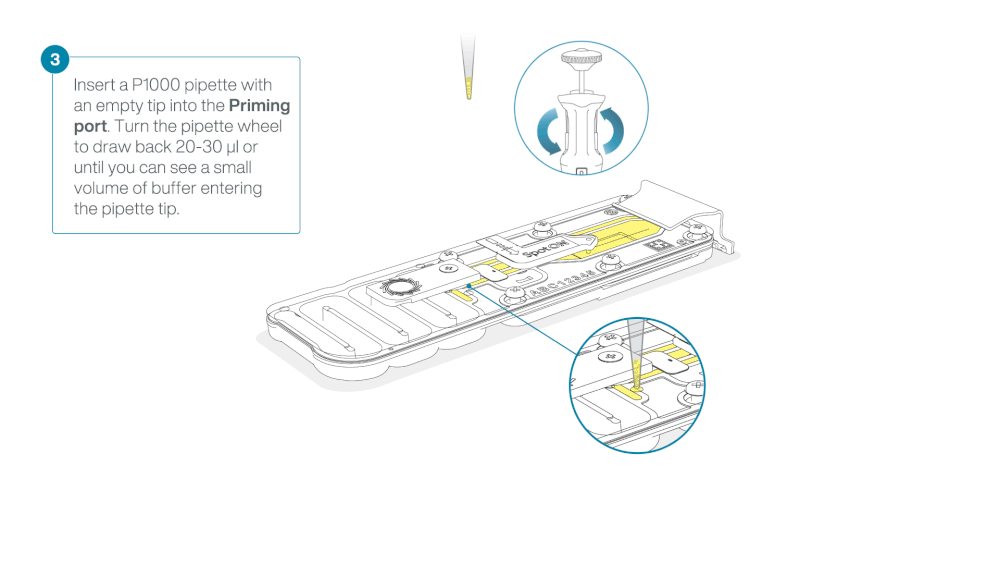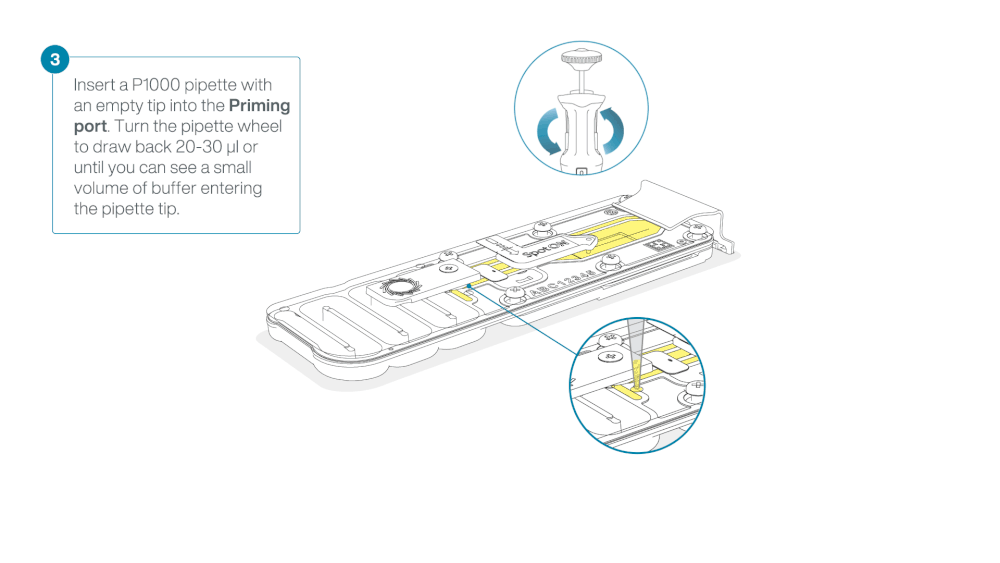
Load 800 µl of the priming mix into the flow cell via the priming port, avoiding the introduction of air bubbles. Wait for five minutes. During this time, prepare the library for loading by following the steps below.
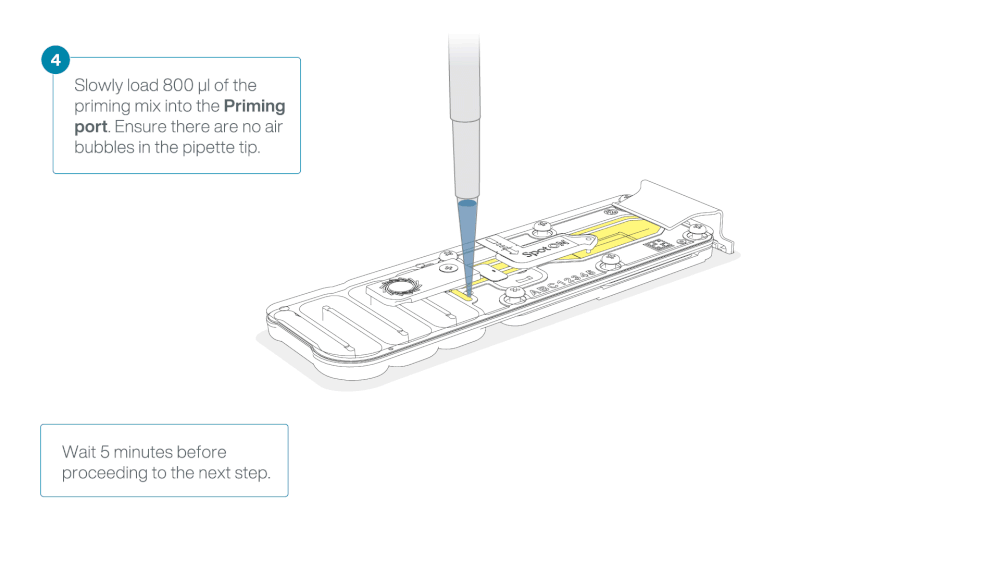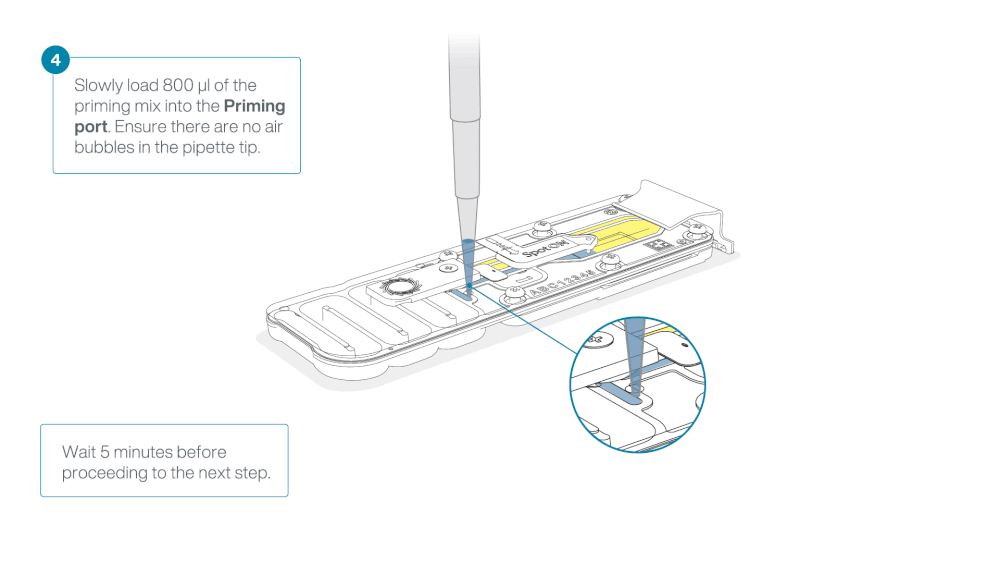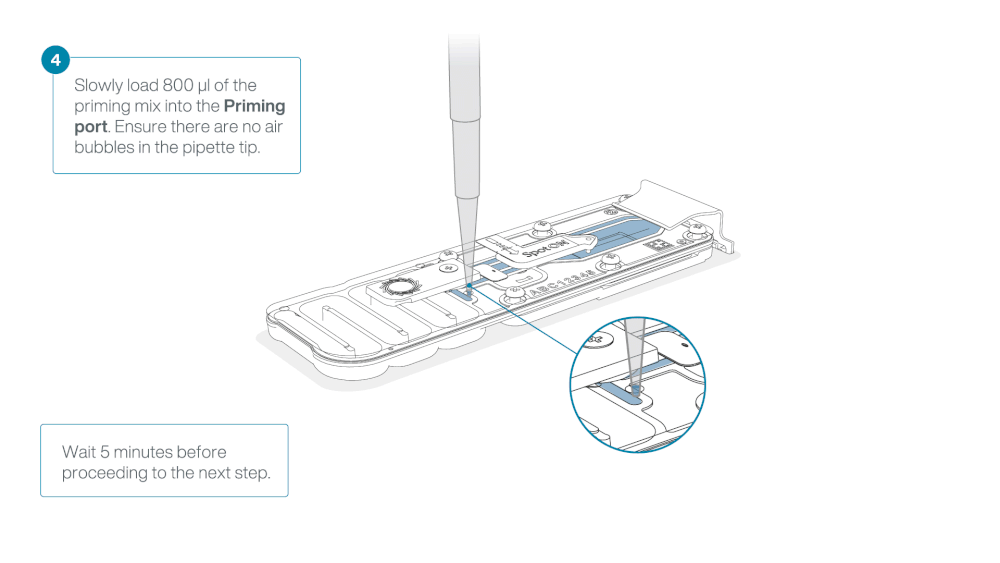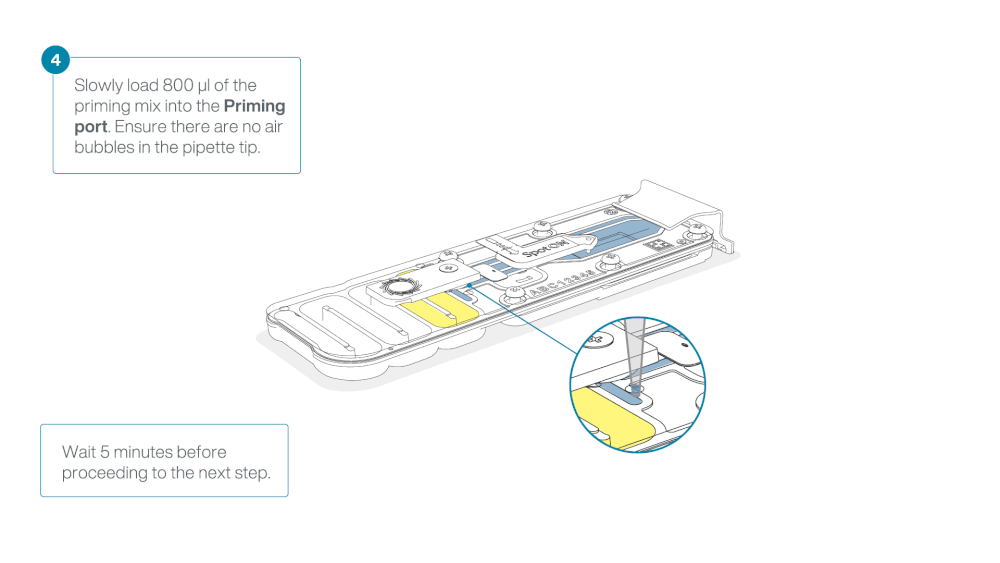
Thoroughly mix the contents of the Library Beads (LIB) by pipetting.
The Library Beads (LIB) tube contains a suspension of beads. These beads settle very quickly. It is vital that they are mixed immediately before use.
We recommend using the Library Beads (LIB) for most sequencing experiments. However, the Library Solution (LIS) is available for more viscous libraries.
In a new 1.5 ml Eppendorf DNA LoBind tube, prepare the library for loading as follows:
| Reagent | Volume per flow cell |
|---|---|
| Sequencing Buffer (SB) | 37.5 µl |
| Library Beads (LIB) mixed immediately before use, or Library Solution (LIS), if using | 25.5 µl |
| DNA library | 12 µl |
| Total | 75 µl |
Complete the flow cell priming:
- Gently lift the SpotON sample port cover to make the SpotON sample port accessible.
- Load 200 µl of the priming mix into the flow cell priming port (not the SpotON sample port), avoiding the introduction of air bubbles.

Mix the prepared library gently by pipetting up and down just prior to loading.
Add 75 μl of the prepared library to the flow cell via the SpotON sample port in a dropwise fashion. Ensure each drop flows into the port before adding the next.

Gently replace the SpotON sample port cover, making sure the bung enters the SpotON port and close the priming port.

For optimal sequencing output, install the light shield on your flow cell as soon as the library has been loaded.
We recommend leaving the light shield on the flow cell when library is loaded, including during any washing and reloading steps. The shield can be removed when the library has been removed from the flow cell.
Place the light shield onto the flow cell, as follows:
Carefully place the leading edge of the light shield against the clip. Note: Do not force the light shield underneath the clip.
Gently lower the light shield onto the flow cell. The light shield should sit around the SpotON cover, covering the entire top section of the flow cell.

The MinION Flow Cell Light Shield is not secured to the flow cell and careful handling is required after installation.
Close the device lid and set up a sequencing run on MinKNOW.
When a flow cell is inserted into the MinION Mk1D, the device lid will sit on top of the flow cell, leaving a small gap around the sides. This is normal and has no impact on the performance of the device.
Please refer to this FAQ regarding the device lid.

8. Data acquisition and basecalling
We do not recommend sequencing and performing data analysis simultaneously on your device.
To ensure the compute on your device can keep up with the requirements for sequencing and/or analysis, we strongly recommend against running both processes at the same time.
Ensure your sequencing run has completed before setting off data analysis. Data analysis will be performed post-sequencing.
Equally, we do not recommend starting a sequencing run if you are currently performing data analysis on your device.
How to start sequencing
Data acquisition and real-time basecalling are carried out by the MinKNOW software. Please ensure MinKNOW is installed on your computer or device. Further instructions for setting up a sequencing run can be found in the MinKNOW protocol.
We recommend setting up your sequencing run using the basecalling and barcoding recommendations outlined below. All other parameters can be left to their default settings.
MinKNOW settings
For fastest turnaround time and easiest analysis, basecalling should be performed live during the sequencing run using the High Accuracy (HAC) basecaller.
Below are the recommendations for MinKNOW settings:
Positions
Flow cell position: [user defined]
Experiment name: [user defined]
Flow cell type: FLO-MIN114
Sample ID: [user defined]
Kit
Kit selection: Rapid Barcoding Kit 96 (SQK-RBK114-96)
Run configuration
Sequencing and analysis
Basecalling: On [default]
Modified bases: Off
Model: High-accuracy basecalling (HAC) [default]
Barcoding: On [default]
Trim barcodes: Off [default]
Barcode both ends: On
Custom barcodes selection: Off [default]
Alignment: Off [default]
Adaptive sampling: Off [default]
Advanced options
Active channel selection: On [default]
Time between pore scans: 1.5 [default]
Reserve pores: On [default]
Data targets
Run limit: [user defined]*
*Sequencing time will depend on data requirements.
Output
Output format
.POD5: On [default]
.FASTQ: On [default]
.BAM: On
Filtering: On [default]
Qscore: 9 [default]
Minimum read length: 200 bp [default]
9. Downstream analysis
Recommended pipeline analysis
The wf-artic is a bioinformatics workflow for the analysis of ARTIC sequencing data prepared using the Midnight protocol. The bioinformatics workflow is orchestrated by the Nextflow software. Nextflow is a publicly available and open-source project that enables the execution of scientific workflows in a scalable and reproducible way. The use of the Nextflow software has been integrated into the EPI2ME Labs software that we recommend for running our downstream analysis methods.
Alternative methods for downstream analysis are available using your device terminal or command line, however we only suggest this for experienced users.
Demultiplexed sequence reads are processed using the ARTIC Field Bioinformatics software that has been modified for the analysis of FASTQ sequences prepared using Oxford Nanopore Rapid Sequencing kits. The other modification to the ARTIC workflow is the use of a primer scheme that defines the sequencing primers used by the Midnight protocol and their genomic locations on the SARS-CoV-2 genome.
The wf-artic workflow includes other analytical steps that include cladistic analysis using Nextclade and strain assignment using Pangolin. The data facets included in the report are parameterised and additional information such as plots of depth-of-coverage across the reference genome is optional.
The complete source for wf-artic is linked, and the Nextflow software will download the scripts and logic flow from this location.
The wf-artic workflow needs to be started manually as outlined below in 'Running a Midnight analysis using EPI2ME Labs'.
Software set-up and installation
The EPI2ME application provides a clean interface to accessing bioinformatics workflows, and is our recommended method in performing your post-sequencing analysis.
Follow the instructions in the EPI2ME Installation guide to install the application on your device.
For more information on how to use EPI2ME, refer to the EPI2ME Quick Start guide.
Installing and updating the wf-artic workflow in EPI2ME Labs:
Ensure you have installed the wf-artic workflow prior to the first analysis set-up.
In the EPI2ME Labs home page, scroll down to the "Install workflows" section and click on epi2me-labs/wf-artic:
If you have already installed the wf-artic workflow, ensure you are using the latest version.
Updating the workflow can be done directly through EPI2ME Labs by navigating to the wf-artic workflow page and clicking Update Workflow:
Demultiplexing of multiple barcoded samples
The wf-artic analysis requires FASTQ sequence data that has already been demultiplexed.
Reads will be demultiplexed during sequencing if you are following the recommended "Required settings in MinKNOW". However, demultiplexing can also be done post-sequencing using the MinKNOW software.
For more information and guides on demultiplexing using MinKNOW, refer to the "Post-run analysis" section in our MinKNOW Protocol.
The expected input for wf-artic is a folder of folders as shown below. Each of the barcode folders should contain the FASTQ sequence data and files may either be uncompressed or gzipped.
$ tree -d MidnightFastq/
MidnightFastq/
├── barcode01
├── barcode02
├── barcode03
├── barcode04
├── barcode05
├── barcode06
└── unclassified
Basecalling model
The basecalling model should be specified when setting up the wf-artic analysis. This should reflect the basecalling model selected during your run set-up as follows:
- If using the default model, High-accuracy basecalling (HAC): r1041_e82_400bps_hac_variant_g615
- If you have used Super accurate basecalling (SUP), please use: r1041_e82_400bps_sup_variant_g615
- If you have used FAST basecalling, please use: r1041_e82_400bps_fast_variant_g615
Running a Midnight analysis using EPI2ME Labs
Open the EPI2ME Labs application on your device.
Open the "Workflows" tab in the EPI2ME Labs application and click on the "wf-artic" workflow:
In the "wf-artic" workflow page, select "Run this workflow" to open analysis set-up:
Complete the wf-artic run set-up:
Select your data input file location. Please note, this folder must contain the demultiplexed FASTQ files of your sequencing run.
Expand the Primer Scheme Selection tab and set the Scheme version to Midnight-ONT/V3.
Expand the Advanced Options tab and set the Medaka model to the basecalling model used in your sequencing run.
Expand the Extra configuration tab and set the Run name for your wf-artic analysis.
Click Launch workflow at the bottom of the page to begin your analysis.
Navigate to the "Analysis" tab in the EPI2ME Labs application to monitor your run:
Completed analysis and result files
The wf-artic analysis outputs will be written to the Working Directory folder specified in the EPI2ME Labs Settings tab.
The location of this folder is specified in the wf-artic run Instance parameters preceeded by out_dir.
However, these files can also be accessed directly in the EPI2ME Labs application from the completed analysis page for your run:
These outputs include:
all_consensus.fastaA multi-FASTA format sequence file containing the consensus sequence for each of the samples investigated. This consensus sequence has been prepared for the whole SARS-CoV-2 genome, not just the spike protein region. The consensus sequence masks the non-spike regions and regions of low sequence coverage with N residues.all_variants.vcf.gzA gzipped VCF file that describes all high-quality genetic variants called by medaka from the sequenced samples.all_variants.vcf.gz.tbiAn index file for the gzipped VCF file.consensus_status.txtA tab delimited file that reports whether a consensus sequence has been successfully prepared for a sample, or not.wf-artic-report.htmlA report summarising these data. This HTML format report also includes the output of the Nextclade software that can be used for a visual inspection of, for example, primer drop out or other qualitative consensus sequence aspects.
Other files are included in the work-directory. This includes per sample VCF files of all genetic variants prior to filtering and other sequences.
Housekeeping and disk usage
The "Working Directory" can be specified in the EPI2ME Labs "Settings" tab and defines where the workflow intermediate files and outputs are stored.
This folder will accumulate a significant number of files that correspond to raw BAM files, other larger intermediates and analysis results files. We recommend this folder to be routinely cleared.
10. Flow cell reuse and returns
Materials
- Flow Cell Wash Kit (EXP-WSH004)
After your sequencing experiment is complete, if you would like to reuse the flow cell, please follow the Flow Cell Wash Kit protocol and store the washed flow cell at +2°C to +8°C.
The Flow Cell Wash Kit protocol is available on the Nanopore Community.
We recommend you to wash the flow cell as soon as possible after you stop the run. However, if this is not possible, leave the flow cell on the device and wash it the next day.
Alternatively, follow the returns procedure to send the flow cell back to Oxford Nanopore.
Instructions for returning flow cells can be found here.
If you encounter issues or have questions about your sequencing experiment, please refer to the Troubleshooting Guide that can be found in this protocol.
11. Issues during DNA/RNA extraction and library preparation
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Low sample quality
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Low DNA purity (Nanodrop reading for DNA OD 260/280 is <1.8 and OD 260/230 is <2.0–2.2) | The DNA extraction method does not provide the required purity | The effects of contaminants are shown in the Contaminants document. Please try an alternative extraction method that does not result in contaminant carryover. Consider performing an additional SPRI clean-up step. |
| Low RNA integrity (RNA integrity number <9.5 RIN, or the rRNA band is shown as a smear on the gel) | The RNA degraded during extraction | Try a different RNA extraction method. For more info on RIN, please see the RNA Integrity Number document. Further information can be found in the DNA/RNA Handling page. |
| RNA has a shorter than expected fragment length | The RNA degraded during extraction | Try a different RNA extraction method. For more info on RIN, please see the RNA Integrity Number document. Further information can be found in the DNA/RNA Handling page. We recommend working in an RNase-free environment, and to keep your lab equipment RNase-free when working with RNA. |
Low DNA recovery after AMPure bead clean-up
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Low recovery | DNA loss due to a lower than intended AMPure beads-to-sample ratio | 1. AMPure beads settle quickly, so ensure they are well resuspended before adding them to the sample. 2. When the AMPure beads-to-sample ratio is lower than 0.4:1, DNA fragments of any size will be lost during the clean-up. |
| Low recovery | DNA fragments are shorter than expected | The lower the AMPure beads-to-sample ratio, the more stringent the selection against short fragments. Please always determine the input DNA length on an agarose gel (or other gel electrophoresis methods) and then calculate the appropriate amount of AMPure beads to use. 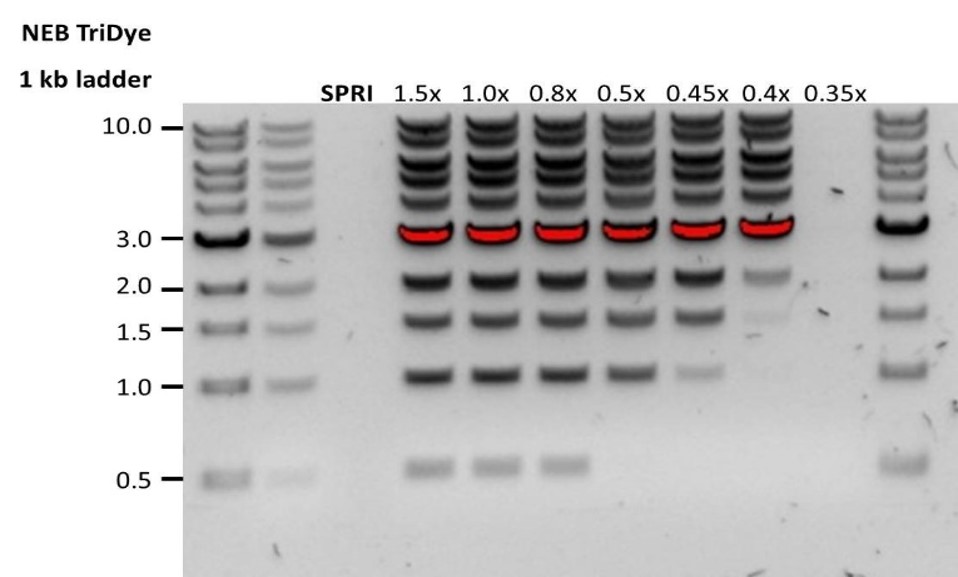 |
| Low recovery after end-prep | The wash step used ethanol <70% | DNA will be eluted from the beads when using ethanol <70%. Make sure to use the correct percentage. |
12. Issues during the sequencing run
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Fewer pores at the start of sequencing than after Flow Cell Check
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | An air bubble was introduced into the nanopore array | After the Flow Cell Check it is essential to remove any air bubbles near the priming port before priming the flow cell. If not removed, the air bubble can travel to the nanopore array and irreversibly damage the nanopores that have been exposed to air. The best practice to prevent this from happening is demonstrated in this video. |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | The flow cell is not correctly inserted into the device | Stop the sequencing run, remove the flow cell from the sequencing device and insert it again, checking that the flow cell is firmly seated in the device and that it has reached the target temperature. If applicable, try a different position on the device (GridION/PromethION). |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | Contaminations in the library damaged or blocked the pores | The pore count during the Flow Cell Check is performed using the QC DNA molecules present in the flow cell storage buffer. At the start of sequencing, the library itself is used to estimate the number of active pores. Because of this, variability of about 10% in the number of pores is expected. A significantly lower pore count reported at the start of sequencing can be due to contaminants in the library that have damaged the membranes or blocked the pores. Alternative DNA/RNA extraction or purification methods may be needed to improve the purity of the input material. The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
MinKNOW script failed
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Script failed" | Restart the computer and then restart MinKNOW. If the issue persists, please collect the MinKNOW log files and contact Technical Support. If you do not have another sequencing device available, we recommend storing the flow cell and the loaded library at 4°C and contact Technical Support for further storage guidance. |
Pore occupancy below 40%
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Pore occupancy <40% | Not enough library was loaded on the flow cell | Ensure you load the recommended amount of good quality library in the relevant library prep protocol onto your flow cell. Please quantify the library before loading and calculate mols using tools like the Promega Biomath Calculator, choosing "dsDNA: µg to pmol" |
| Pore occupancy close to 0 | The Ligation Sequencing Kit was used, and sequencing adapters did not ligate to the DNA | Make sure to use the NEBNext Quick Ligation Module (E6056) and Oxford Nanopore Technologies Ligation Buffer (LNB, provided in the sequencing kit) at the sequencing adapter ligation step, and use the correct amount of each reagent. A Lambda control library can be prepared to test the integrity of the third-party reagents. |
| Pore occupancy close to 0 | The Ligation Sequencing Kit was used, and ethanol was used instead of LFB or SFB at the wash step after sequencing adapter ligation | Ethanol can denature the motor protein on the sequencing adapters. Make sure the LFB or SFB buffer was used after ligation of sequencing adapters. |
| Pore occupancy close to 0 | No tether on the flow cell | Tethers are adding during flow cell priming (FLT/FCT tube). Make sure FLT/FCT was added to FB/FCF before priming. |
Shorter than expected read length
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Shorter than expected read length | Unwanted fragmentation of DNA sample | Read length reflects input DNA fragment length. Input DNA can be fragmented during extraction and library prep. 1. Please review the Extraction Methods in the Nanopore Community for best practice for extraction. 2. Visualise the input DNA fragment length distribution on an agarose gel before proceeding to the library prep. 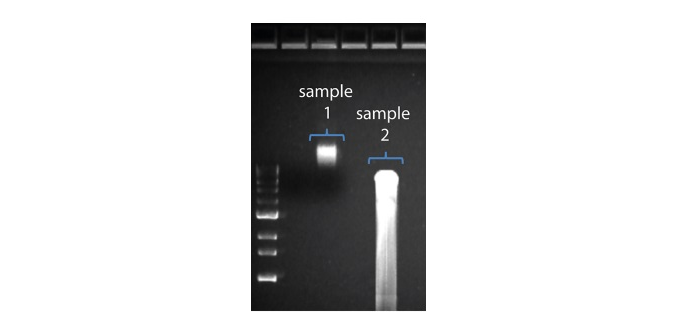 In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented. In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented.3. During library prep, avoid pipetting and vortexing when mixing reagents. Flicking or inverting the tube is sufficient. |
Large proportion of unavailable pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
Large proportion of unavailable pores (shown as blue in the channels panel and pore activity plot) 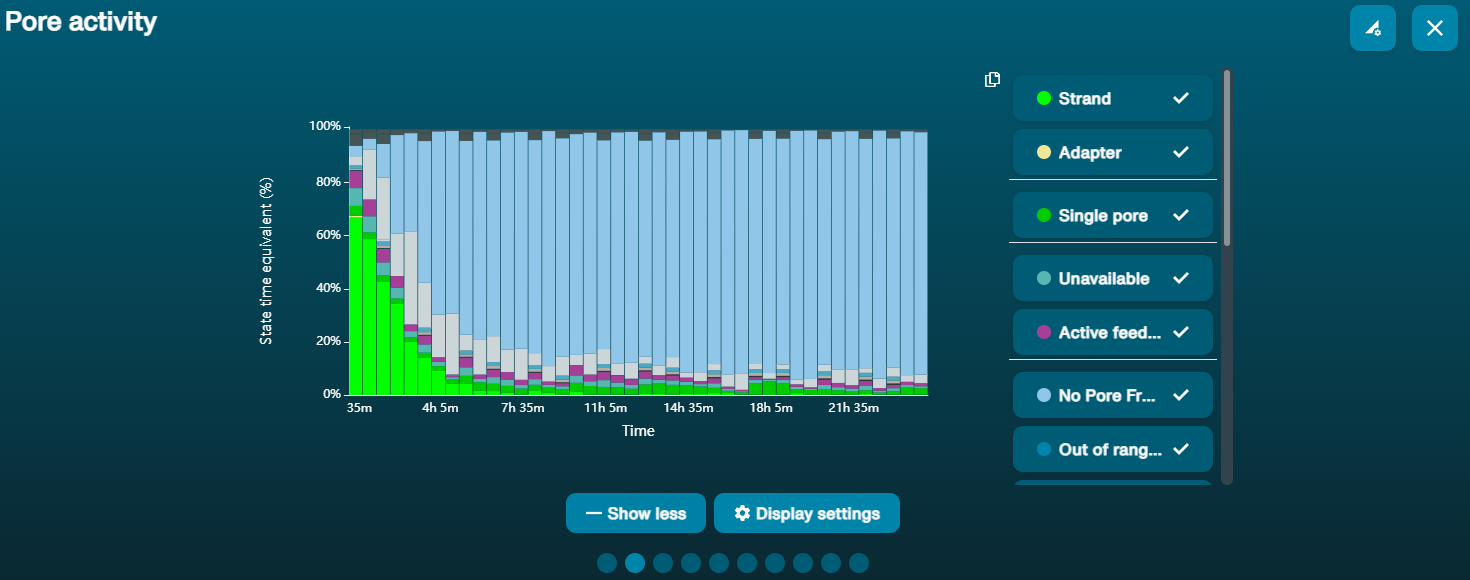 The pore activity plot above shows an increasing proportion of "unavailable" pores over time. The pore activity plot above shows an increasing proportion of "unavailable" pores over time. | Contaminants are present in the sample | Some contaminants can be cleared from the pores by the unblocking function built into MinKNOW. If this is successful, the pore status will change to "sequencing pore". If the portion of unavailable pores stays large or increases: 1. A nuclease flush using the Flow Cell Wash Kit (EXP-WSH004) can be performed, or 2. Run several cycles of PCR to try and dilute any contaminants that may be causing problems. |
Large proportion of inactive pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Large proportion of inactive/unavailable pores (shown as light blue in the channels panel and pore activity plot. Pores or membranes are irreversibly damaged) | Air bubbles have been introduced into the flow cell | Air bubbles introduced through flow cell priming and library loading can irreversibly damage the pores. Watch the Priming and loading your flow cell video for best practice |
| Large proportion of inactive/unavailable pores | Certain compounds co-purified with DNA | Known compounds, include polysaccharides, typically associate with plant genomic DNA. 1. Please refer to the Plant leaf DNA extraction method. 2. Clean-up using the QIAGEN PowerClean Pro kit. 3. Perform a whole genome amplification with the original gDNA sample using the QIAGEN REPLI-g kit. |
| Large proportion of inactive/unavailable pores | Contaminants are present in the sample | The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
Temperature fluctuation
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Temperature fluctuation | The flow cell has lost contact with the device | Check that there is a heat pad covering the metal plate on the back of the flow cell. Re-insert the flow cell and press it down to make sure the connector pins are firmly in contact with the device. If the problem persists, please contact Technical Services. |
Failed to reach target temperature
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Failed to reach target temperature" | The instrument was placed in a location that is colder than normal room temperature, or a location with poor ventilation (which leads to the flow cells overheating) | MinKNOW has a default timeframe for the flow cell to reach the target temperature. Once the timeframe is exceeded, an error message will appear and the sequencing experiment will continue. However, sequencing at an incorrect temperature may lead to a decrease in throughput and lower q-scores. Please adjust the location of the sequencing device to ensure that it is placed at room temperature with good ventilation, then re-start the process in MinKNOW. Please refer to this link for more information on MinION temperature control. |







