Metagenomic sequencing
Enhance genome assemblies
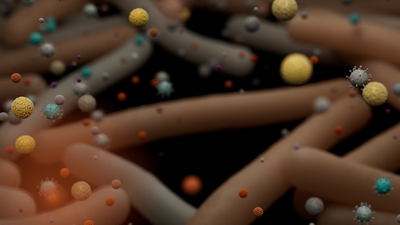
Metagenomics is the genomic analysis of multiple organisms from a single sample, revealing insight into the genetic composition of microbial communities. Any-length nanopore reads deliver enhanced genome assemblies, accurate identification of closely related species, and resolve mobile genetic elements from metagenomic samples.
With Oxford Nanopore sequencing, real-time data streaming unlocks immediate access to results, such as species identification, relative abundance, and antimicrobial resistance (AMR) analysis. Furthermore, combining long reads with targeted approaches enables sequencing of informative genes, such as 16S ribosomal RNA (rRNA) in their entirety, improving resolution of identification.
Featured content

Pathogen metagenomics
Metagenomic sequencing is a key technique for the identification of potential pathogens. This end-to-end workflow overview introduces how to rapidly identify bacterial, fungal, and viral pathogens from respiratory research samples using metagenomic sequencing.

Addressing the challenges of metagenomics with Oxford Nanopore sequencing
Explore how nanopore sequencing is addressing the challenges of metagenomics to obtain complete or nearly complete genome sequences from uncultured microorganisms — providing important means to study their biology, ecology, and evolution.
Related content
Recommended device for metagenomic sequencing
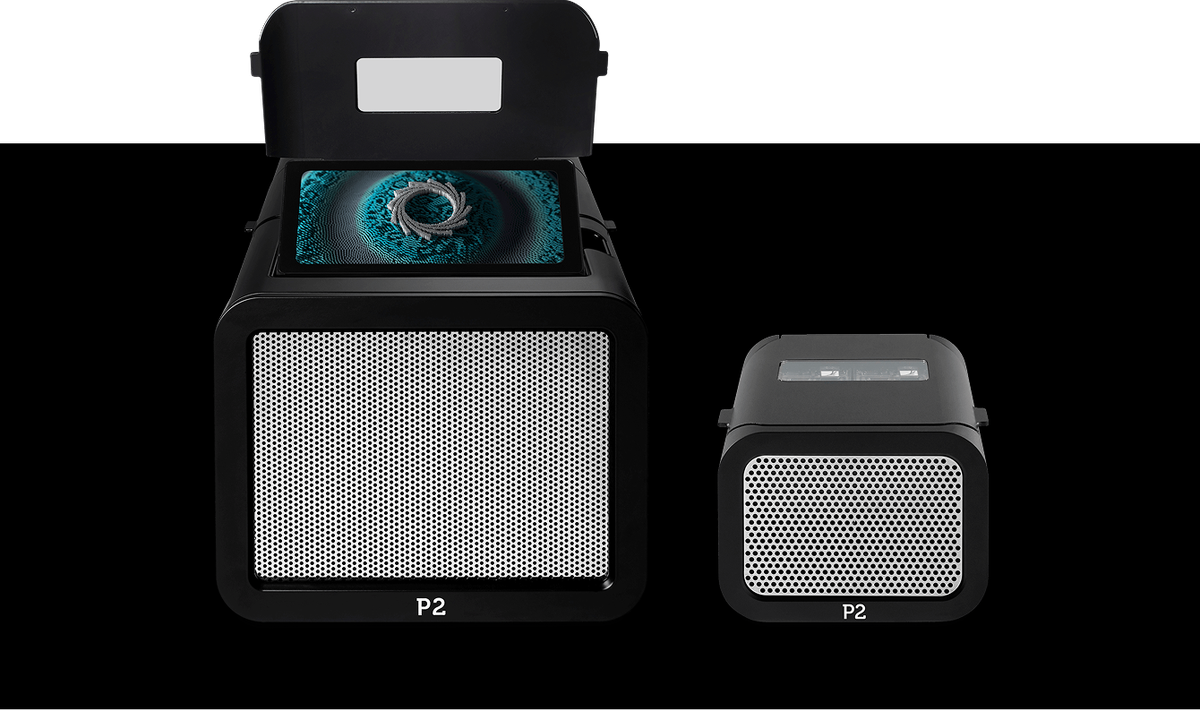

PromethION 2 devices
Offering flexible, affordable, high-output nanopore devices for every lab in a compact, accessible form, the PromethION 2 Solo and PromethION 2 Integrated devices are ideal for obtaining complete circular genomes from complex metagenomic samples in real time.