Whole-genome sequencing
Get comprehensive whole-genome analysis on a single platform
Whole-genome sequencing aims to provide complete analysis of an organism’s genome, but legacy short-read sequencing technologies are known to miss many important genomic regions and variants — even in the exome.
Oxford Nanopore sequencing, with its capacity to produce ultra-long reads (exceeding 4 Mb), can span complex structural variants (SVs) and repeat regions that legacy technologies cannot access, while the facility to directly sequence native DNA (and RNA), without amplification, further enables simultaneous detection of epigenetic modifications alongside the nucleotide sequence to deliver deeper genomic insights.
Featured content

Comprehensive human genomic variant and methylation analysis
From sample to answer, discover how nanopore sequencing delivers comprehensive human variation detection in this best practice, end-to-end workflow. Get accurate SNV, SV, STR, and methylation results in a single streamlined assay, with integrated tertiary analysis.

Generate reference-quality bacterial genome assemblies
Discover how our rapid, end-to-end nanopore sequencing workflow delivers streamlined access to complete, reference-quality bacterial genome assemblies.
Recommended device for whole-genome sequencing
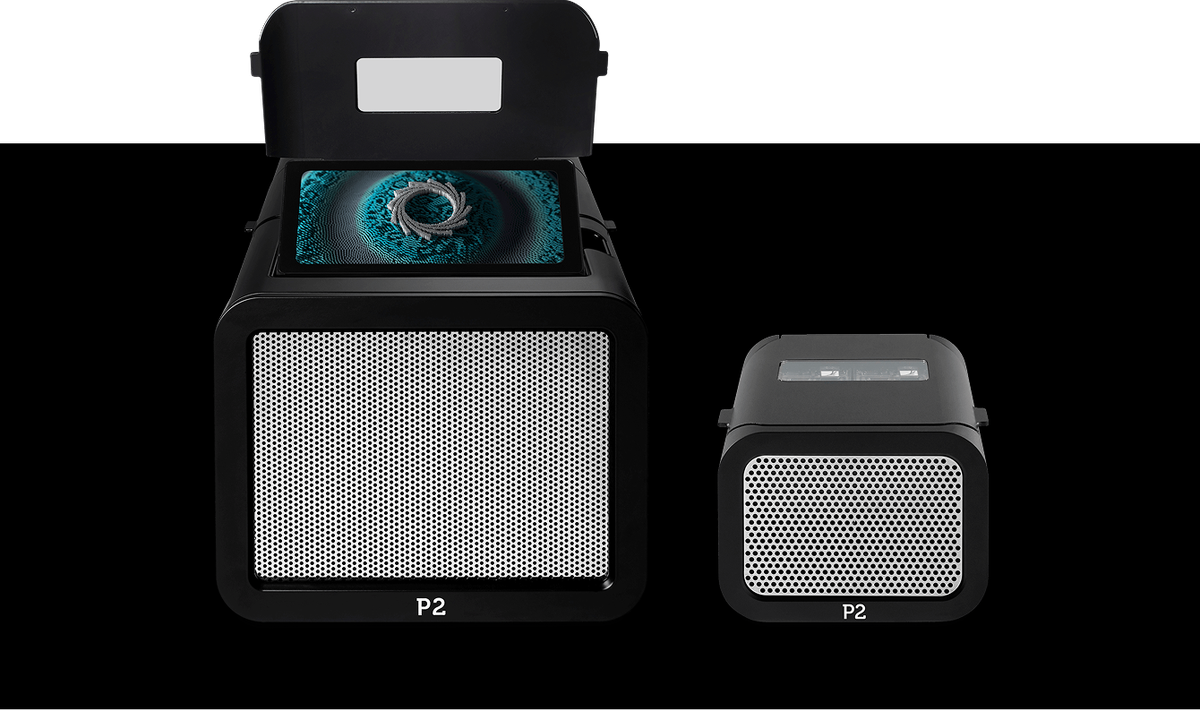

PromethION 2 devices
Offering the flexibility of two independent, high-output PromethION Flow Cells, the compact PromethION 2 devices bring the benefits of high-coverage, real-time nanopore sequencing to every lab. Ideal for low-cost access to highly accurate whole-genome and metagenome assemblies.