Multiomics whole-genome characterisation of a cancer genome using Oxford Nanopore sequencing
- Published on: May 20 2025
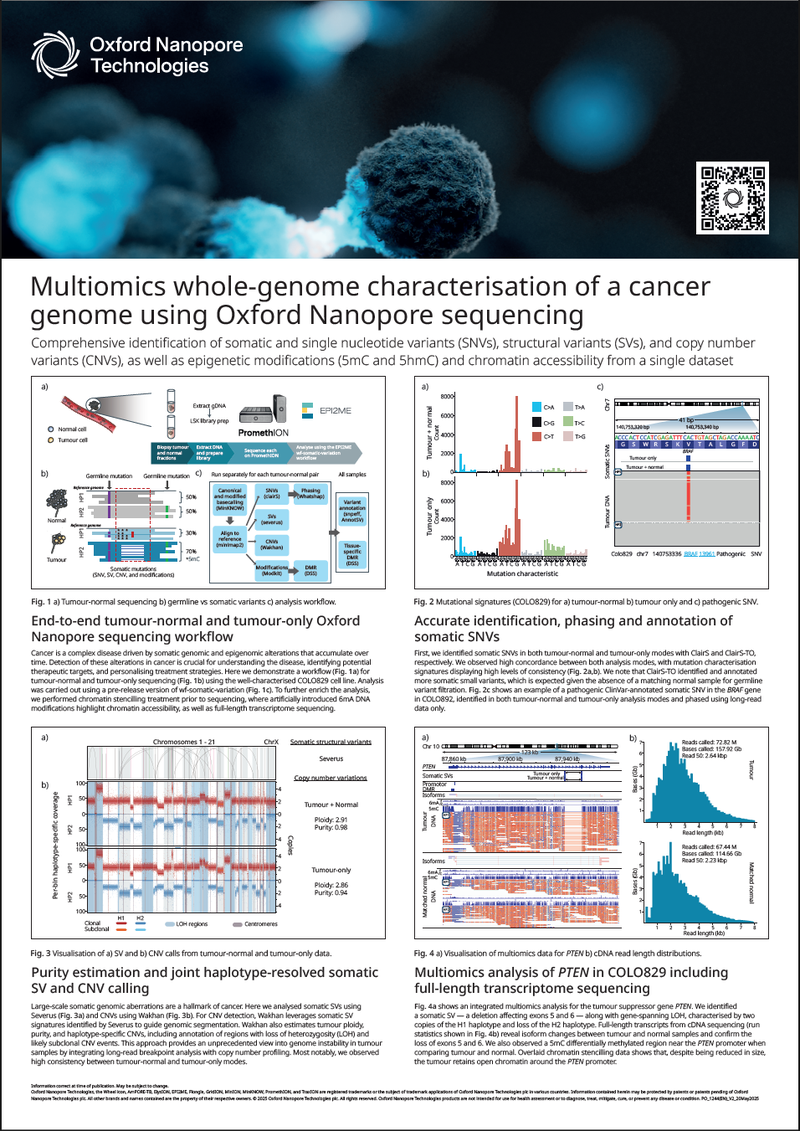
Comprehensive identification of somatic and single nucleotide variants (SNVs), structural variants (SVs), and copy number variants (CNVs), as well as epigenetic modifications (5mC and 5hmC) and chromatin accessibility from a single dataset.
Download the poster to find out about:
- Our end-to-end tumour-normal and tumour-only Oxford Nanopore sequencing workflow
- Accurate identification, phasing and annotation of somatic SNVs
- Purity estimation and joint haplotype-resolved somatic SV and CNV calling
- Multiomics analysis of PTEN in COLO829 including full-length transcriptome sequencing