Human genomics
Generate unprecedented insights for human genomics research
With real-time, multiomic Oxford Nanopore sequencing, you can discover previously hidden human genomic, epigenomic, and transcriptomic variation — from the population level down to the single-cell level.
Fully characterise challenging regions that cannot be resolved with legacy short-read sequencing technologies. Detect single nucleotide variants (SNVs), short tandem repeats (STRs), and DNA methylation, and generate highly contiguous genomes by spanning repeat regions and structural variants (SVs) — in one go. Furthermore, with RNA sequencing, interrogate full-length RNA transcript isoforms and identify RNA base modifications, such as methylation, as standard.
With Oxford Nanopore technology, there is no limit to read length (current record >4 Mb), allowing you to reveal critical insights for human genomics research — from developmental biology and rare disease genomics to common complex diseases.
Featured content

A guide to human genomics with Oxford Nanopore
This getting started guide introduces how to sequence human genomes with Oxford Nanopore, from the construction of new, highly complete reference assemblies to the comprehensive identification of variants.

Comprehensive human genomic variant and methylation analysis
From sample to answer, discover how nanopore sequencing delivers comprehensive human variation detection in this best practice, end-to-end workflow. Get accurate SNV, SV, STR, and methylation results in a single streamlined assay, with integrated tertiary analysis.
Highlighted application
Pharmacogenomics with Oxford Nanopore sequencing
Pharmacogenomics (PGx), the study of genomic-driven response to treatments, includes some of the genome's most challenging genes. Conventional array and legacy sequencing technologies are incapable of fully resolving the problems presented by pseudogene homology, haplotyping requirements, and complex structural variants.
Oxford Nanopore sequencing delivers the capabilities to fully resolve all the PGx variants with a single technology and a single assay. You can choose from two different methods to achieve full resolution PGx: (1) hybrid capture; and (2) adaptive sampling.
Recommended device for human genome sequencing
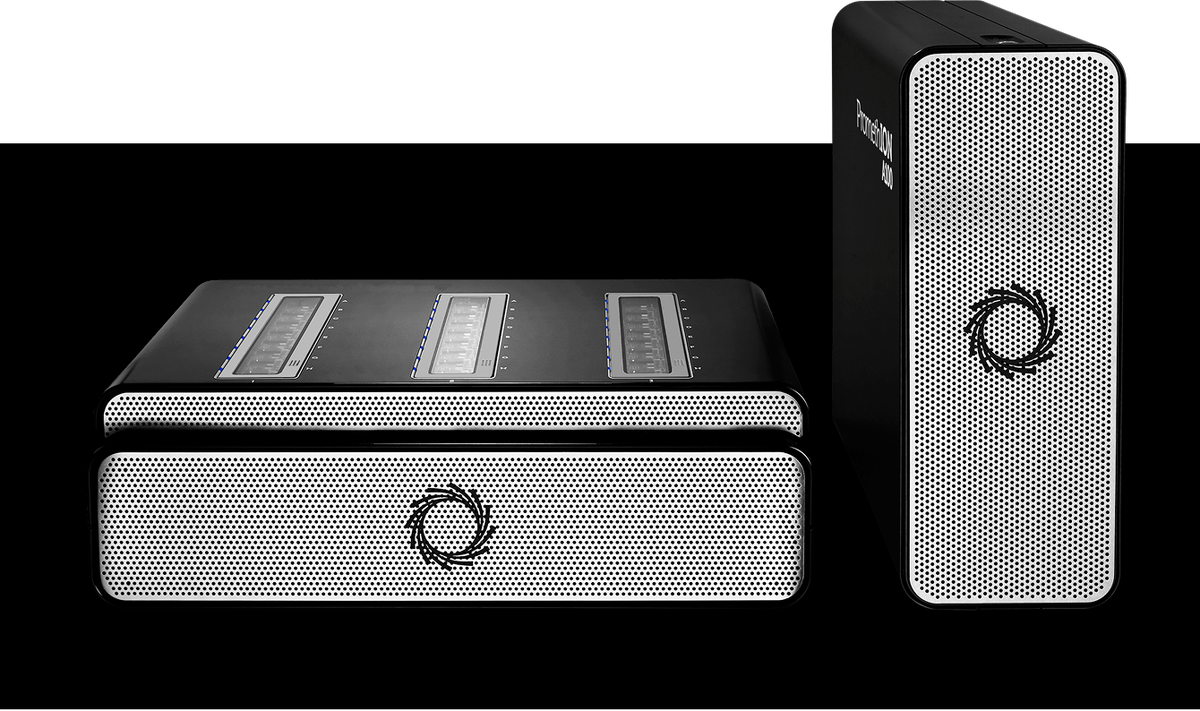

PromethION 24
The PromethION 24 device combines 24 independently addressable, high-capacity flow cells with powerful, integrated compute. The device delivers flexible, on-demand access to terabases of sequencing data — ideal for cost-effective, high-throughput sequencing of human genomes and transcriptomes.