Cancer research and sequencing
Reveal more cancer biology with ultra-rich Oxford Nanopore sequencing data
The genetic underpinnings of cancer are diverse and many types of genomic aberration — from single nucleotide variants (SNVs) to structural variants (SVs), copy number variants (CNVs), fusion transcripts, and epigenetic modifications (e.g. DNA/RNA methylation) — can cause, contribute to, or indicate disease. As a result, researchers traditionally relied on multiple techniques to identify and analyse different facets of cancer.
Now, with Oxford Nanopore technology, researchers are going beyond next-generation sequencing (NGS), generating sequencing reads of any length, including ultra-long reads (>4 Mb achieved) that can span complex genomic regions. This, combined with integrated base modification detection and real-time results, means that nanopore oncology sequencing delivers a streamlined and rapid solution for complete characterisation of cancer and tumour samples.
Featured content

Accelerating cancer research through comprehensive genomics
Discover how cancer researchers are using Oxford Nanopore sequencing for comprehensive characterisation of cancer samples, delivering accurate and rapid analysis of SVs, SNVs, CNVs, methylation, full-length isoforms, fusion transcripts, and splice variants — all from a single technology.

Hereditary Cancer Panel: targeted sequencing via adaptive sampling
Accurately resolve complex structural variants (SVs), single nucleotide variants (SNVs), and insertions/deletions (indels) with the Oxford Nanopore Hereditary Cancer Panel — a comprehensive sequencing assay. In this flyer, discover how to utilise the Hereditary Cancer Panel to investigate 258 key genes associated with hereditary cancer risk.
Related content
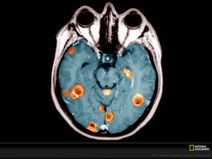
ROBIN: a unified nanopore-based assay integrating intraoperative methylome classification and next-day comprehensive profiling for ultra-rapid tumour diagnosis

Exploring the future of rapid leukaemia diagnosis with a single-platform workflow

Obtaining full-length isoforms from single cells with Oxford Nanopore sequencing
The most comprehensive insight into cancer genomes
Featured product for oncology
Hereditary Cancer Panel
The Oxford Nanopore Hereditary Cancer Panel is a comprehensive sequencing assay targeting 258 full-length genes — covering exons, introns, and promoters — linked to inherited cancer risk. The panel uses adaptive sampling: a fast and flexible on-sequencer target enrichment methodology. No lengthy library preparation, baits, or primers required.
Recommended device for cancer research and sequencing

PromethION 24
PromethION Flow Cells offer the highest yield for nanopore sequencing, translating into high coverage of cancer genomes and high resolution of full-length transcripts. PromethION 24 is ideal for comprehensive whole-genome characterisation and biomarker discovery across any number of cancer samples.







