Human cfDNA multiplex sequencing from blood using SQK-NBD114.24 (CFM_9208_v114_revH_31Mar2026)
PromethION: Protocol
Human cfDNA multiplex sequencing from blood using SQK-NBD114.24 V CFM_9208_v114_revH_31Mar2026
This document describes the steps for sample extraction, library preparation, sequencing, and data analysis.
The protocol:
- Uses human cfDNA
- Enables multiplexing of 12 samples
- Uses the Native Barcoding Kit 24 V14 (SQK-NBD114.24)
- Is compatible with R10.4.1 flow cells
For Research Use Only
FOR RESEARCH USE ONLY
Contents
Introduction to the protocol
Sample preparation
Library preparation
- 4. DNA repair and end-prep
- 5. Native barcode ligation
- 6. Adapter ligation and clean-up
- 7. Priming and loading the PromethION Flow Cell
Sequencing and data analysis
Troubleshooting
Overview
This document describes the steps for sample extraction, library preparation, sequencing, and data analysis.
The protocol:
- Uses human cfDNA
- Enables multiplexing of 12 samples
- Uses the Native Barcoding Kit 24 V14 (SQK-NBD114.24)
- Is compatible with R10.4.1 flow cells
For Research Use Only
1. Overview of the protocol
Introduction to the multiplex human cfDNA sequencing protocol
This protocol describes how to carry out preparation and sequencing of 12 human cell-free DNA (cfDNA) samples using the Native Barcoding Kit 24 V14 (SQK-NBD114.24). Typically, we obtain ~3 Gb of aligned data (1x coverage) for each of the 12 human cfDNA samples processed with this protocol.
Please note: This method has been developed for multiplexing 12 samples. We do not recommend deviating from the outlined method.
Reducing the number of samples multiplexed may lead to an increase in coverage, but increasing the number of samples multiplexed will lead to a reduction in coverage of aligned data per sample.
Prior to library preparation, the sample extraction is carried out using the QIAGEN QIAamp MinElute ccfDNA Midi Kit, following our Human blood cell-free DNA (cfDNA) extraction for multiplex sequencing method.
Note: We recommend that blood samples are processed while fresh, as we have observed potential gDNA contamination arising from blood that has been stored in certain types of collection tubes.
For more information on the development and performance of this method, please refer to our Updated method for cell-free DNA (cfDNA) methylation profiling know-how document. An additional know-how document is also available for the optimisation of library preparation for longer cell-free DNA (cfDNA).
Steps in the workflow
Prepare for your experiment
You will need to:
- Extract your DNA, and check its length, quantity and purity. The quality checks performed during the protocol are essential in ensuring experimental success.
- Ensure you have your sequencing kit, the correct equipment and third-party reagents.
- Download the software for acquiring and analysing your data.
- Check your flow cell to ensure it has enough pores for a good sequencing run.
Sample preparation
Using the outlined extraction method, extract the cfDNA from your human blood samples, and quantify the DNA:
Library preparation
The Table below is an overview of the steps required in the library preparation, including timings and optional stopping points.
| Library preparation | Process | Time | Stop option |
|---|---|---|---|
| DNA repair and end-prep | Repair the cfDNA and prepare the DNA ends for adapter attachment. | 125 minutes | 4°C overnight |
| Native barcode ligation | Ligate the native barcodes to the DNA ends. | 60 minutes | 4°C overnight |
| Adapter ligation and clean-up | Attach the sequencing adapters to the DNA ends. | 50 minutes | 4°C short-term storage or for repeated use, such as re-loading your flow cell. -80°C for single-use, long-term storage. We strongly recommend sequencing your library as soon as it is adapted. |
| Priming and loading the flow cell | Prime the flow cell and load the prepared library for sequencing. | 10 minutes |
Sequencing and analysis
You will need to:
- Start a sequencing run using the MinKNOW software which will collect raw data from the device and basecall reads.
- (Optional) Raw sequencing data can be basecalled and aligned to a reference using Dorado.
- (Optional) Start the EPI2ME software and select a bioinformatics workflow to analyse your data. Alternatively, external tools can be used to further analyse and explore your data.
Compatibility of this protocol
This protocol should only be used in combination with:
- Native Barcoding Kit 24 V14 (SQK-NBD114.24)
- R10.4.1 PromethION Flow Cells (FLO-PRO114M)
- Flow Cell Wash Kit (EXP-WSH004)
- Sequencing Auxiliary Vials V14 (EXP-AUX003)
- Native Barcoding Expansion V14 (EXP-NBA114)
- PromethION 24/48 device - PromethION IT requirements document
- PromethION 2 Solo device - PromethION 2 Solo IT requirements document
- PromethION 2 Integrated - PromethION 2 Integrated IT requirements document
2. Equipment and consumables
Materials
- (FOR EXTRACTION) ≥3.5 ml blood in EDTA K2 vacuum tube or ≥1ml plasma, per sample
- (FOR LIBRARY PREPARATION) ≥6 ng of recovered human cfDNA per sample
- Native Barcoding Kit 24 V14 (SQK-NBD114.24)
Consumables
- PromethION Flow Cells
- QIAamp MinElute ccfDNA Midi Kit (QIAGEN, 55284)
- Agencourt AMPure XP beads (Beckman Coulter™, A63881)
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- NEBNext® FFPE DNA Repair v2 Module (NEB, E7360)
- NEBNext Ultra II End repair/dA-tailing Module (NEB, E7546)
- NEB Blunt/TA Ligase Master Mix (NEB, M0367)
- NEBNext Quick Ligation Module (NEB, E6056)
- Ethanol, 100% (e.g. Fisher, 16606002)
- Isopropanol
- Freshly prepared 80% ethanol in nuclease-free water
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- 5 ml Eppendorf DNA LoBind tubes
- 15 ml Falcon tubes
- 1.5 ml Eppendorf DNA LoBind tubes
- 0.2 ml thin-walled PCR tubes
Equipment
- PromethION device
- PromethION Flow Cell Light Shield
- Centrifuge with capacity for 5 ml and 15 ml tubes, and a swing out and fixed angle rotors
- Hula mixer (gentle rotator mixer)
- Magnetic rack for 15 ml tubes
- Magnetic separation rack, suitable for 1.5 ml Eppendorf tubes
- Thermomixer, or other shaker for microcentrifuge tubes, with capacity to heat at 56°C
- Microfuge
- Vortex mixer
- Thermal cycler
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- P2 pipette and tips
- Ice bucket with ice
- Timer
- Qubit™ fluorometer (or equivalent for QC check)
The above list of materials, consumables, and equipment is for the extraction method in the sample preparation section, as well as the library preparation section of the protocol. If you have pre-extracted sample(s), you will only require the materials for the library preparation section of this protocol.
This protocol has been developed to process and sequence 12 samples. The following inputs are required:
Input requirements per sample for the extraction method:
- ≥3.5 ml of fresh blood in EDTA K2 vacuum tube or ≥1ml plasma, per sample
Note: We recommend that blood samples are processed while fresh, as we have observed potential gDNA contamination arising from blood that has been stored in certain types of collection tubes.
Input requirements per sample for the library preparation:
- Use ≥6 ng of recovered cfDNA, per sample as input. Inputs should be normalised so the same mass of sample is used in each reaction.
Note: This method has been developed for multiplexing 12 samples. We do not recommend deviating from the outlined method.
Input DNA
How to QC your input DNA
It is important that the input DNA meets the quantity and quality requirements. Using too little or too much DNA, or DNA of poor quality (e.g. highly fragmented or containing RNA or chemical contaminants) can affect your library preparation.
For instructions on how to perform quality control of your DNA sample, please read the Input DNA/RNA QC protocol.
Chemical contaminants
Depending on how the DNA is extracted from the raw sample, certain chemical contaminants may remain in the purified DNA, which can affect library preparation efficiency and sequencing quality. Read more about contaminants on the Contaminants page of the Community.
Third-party reagents
We have validated and recommend the use of all the third-party reagents used in this protocol. Alternatives have not been tested by Oxford Nanopore Technologies.
For all third-party reagents, we recommend following the manufacturer's instructions to prepare the reagents for use.
Check your flow cell
We highly recommend that you check the number of pores in your flow cell prior to starting a sequencing experiment. This should be done within 12 weeks of purchasing your PromethION Flow Cells. Oxford Nanopore Technologies will replace any unused flow cell with fewer than the number of pores listed in the Table below, when the result is reported within two days of performing the flow cell check, and when the storage recommendations have been followed. To do the flow cell check, please follow the instructions in the Flow Cell Check document.
| Flow cell | Minimum number of active pores covered by warranty |
|---|---|
| PromethION Flow Cell | 5000 |
We do not recommend mixing barcoded libraries with non-barcoded libraries prior to sequencing.
The Native Adapter (NA) included in this kit and protocol is not interchangeable with other sequencing adapters.
Native Barcoding Kit 24 V14 (SQK-NBD114.24) contents
Note: We are in the process of updating our native barcoding kits with an increased volume of Short Fragment Buffer (SFB). If you have an old format kit and/or require additional volume of Short Fragment Buffer (SFB), this can be purchased via our SFB Expansion (EXP-SFB001).
New format: increased volume of Short Fragment Buffer (SFB)
| Name | Acronym | Cap colour | No. of vials | Fill volume per vial (µl) |
|---|---|---|---|---|
| DNA Control Sample | DCS | Yellow | 2 | 35 |
| Native Adapter | NA | Green | 1 | 40 |
| Sequencing Buffer | SB | Red | 1 | 700 |
| Library Beads | LIB | Pink | 1 | 600 |
| Library Solution | LIS | White cap, pink label | 1 | 600 |
| Elution Buffer | EB | Black | 2 | 500 |
| AMPure XP Beads | AXP | Clear cap, light teal label | 1 | 6,000 |
| Long Fragment Buffer | LFB | Orange | 1 | 1,800 |
| Short Fragment Buffer | SFB | Clear | 1 | 13,000 |
| EDTA | EDTA | Blue | 1 | 700 |
| Flow Cell Flush | FCF | Clear cap, light blue label | 1 | 8,000 |
| Flow Cell Tether | FCT | Purple | 1 | 200 |
| Native Barcode plate | NB01-24 | - | 2 plates, 3 sets of barcodes per plate | 5 µl per well |
Old format: lower volume of Short Fragment Buffer (SFB)
| Name | Acronym | Cap colour | No. of vials | Fill volume per vial (µl) |
|---|---|---|---|---|
| DNA Control Sample | DCS | Yellow | 2 | 35 |
| Native Adapter | NA | Green | 1 | 40 |
| Sequencing Buffer | SB | Red | 1 | 700 |
| Library Beads | LIB | Pink | 1 | 600 |
| Library Solution | LIS | White cap, pink label | 1 | 600 |
| Elution Buffer | EB | Black | 2 | 500 |
| AMPure XP Beads | AXP | Clear cap, light teal label | 1 | 6,000 |
| Long Fragment Buffer | LFB | Orange | 1 | 1,800 |
| Short Fragment Buffer | SFB | Clear | 1 | 1,800 |
| EDTA | EDTA | Blue | 1 | 700 |
| Flow Cell Flush | FCF | Clear cap, light blue label | 1 | 8,000 |
| Flow Cell Tether | FCT | Purple | 1 | 200 |
| Native Barcode plate | NB01-24 | - | 2 plates, 3 sets of barcodes per plate | 5 µl per well |
Note: This product contains AMPure XP reagent manufactured by Beckman Coulter, Inc. and can be stored at -20°C with the kit without detriment to reagent stability.
Note: The DNA Control Sample (DCS) is a 3.6 kb standard amplicon mapping the 3' end of the Lambda genome.
3. Sample extraction method for multiplex sequencing of human cfDNA
Materials
- ≥3.5 ml fresh human blood in EDTA K2 vacuum tube or ≥1ml plasma, per sample
Consumables
- QIAamp MinElute ccfDNA Midi Kit (QIAGEN, 55284)
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- Ethanol, 100% (e.g. Fisher, 16606002)
- Isopropanol
- Freshly prepared 80% ethanol in nuclease-free water
- Nuclease-free water (e.g. ThermoFisher, AM9937)
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- 5 ml Eppendorf DNA LoBind tubes
- 15 ml Falcon tubes
- 0.2 ml thin-walled PCR tubes
- 1.5 ml Eppendorf DNA LoBind tubes
Equipment
- Centrifuge with capacity for 5 ml and 15 ml tubes, and a swing out and fixed angle rotors
- Microfuge
- Hula mixer (gentle rotator mixer)
- Magnetic rack for 15 ml tubes
- Magnetic separation rack
- Vortex mixer
- P1000 pipette and tips
- P100 pipette and tips
- Thermal cycler
- Ice bucket with ice
- Qubit™ fluorometer (or equivalent for QC check)
Optional equipment
- Agilent Femto Pulse System (or equivalent for read length QC)
Optimised extraction: Human blood cell-free DNA (cfDNA) extraction for multiplex sequencing
This extraction method can also be found in the Extraction Protocols tab in the Documentation space on the Nanopore Community: Human blood cell-free DNA (cfDNA) extraction for multiplex sequencing.
These instructions describe a method to extract cell-free DNA (cfDNA) from 12 human blood samples collected in EDTA K2 vacuum tubes (step 1), or human plasma (step 3). The extraction is performed using the QIAGEN QIAamp MinElute ccfDNA Midi Kit.
Note: The yield, DIN and sequencing read length of extracted DNA may vary depending on initial sample quality. Please ensure you are following the correct method and using high-quality sample inputs.
Alternatively, if you have previously extracted and stored your cfDNA sample(s), this can be used directly in the Library preparation section of this protocol.
We recommend that blood samples are processed while fresh, as we have observed potential gDNA contamination arising from blood that has been stored in certain types of collection tubes.
Preparation of plasma from fresh blood
Centrifuge ≥3.5 ml of fresh blood (overnight chilled delivery) in the EDTA K2 vacuum tube at 1,900 x g for 10 minutes at 4°C in a swing out rotor centrifuge.
Pipette and transfer the supernatant (this is the plasma fraction) to a fresh 5 ml DNA LoBind Eppendorf tube.
We recommend a minimum plasma volume of 1 ml is used, although volumes up to 4 ml have been validated.
To remove residual cells from the plasma, centrifuge the plasma at 16,000 x g for 10 minutes (or 6,000 x g for 30 minutes depending on the spin capacity of the centrifuge) at 4°C, in a fixed angle rotor.
It is important to remove residual cells from the sample when the blood/plasma is still fresh (from an overnight chilled delivery). Failing to do so will result in increased amounts of gDNA contamination in the sequencing library.
Aspirate the supernatant and transfer it to a fresh 15 ml tube.
Purification of cfDNA from 1–5 ml serum or plasma Using the QIAamp MinElute ccfDNA Midi Kit
Before starting the extraction:
- Prepare a shaker for microcentrifuge tubes at room temperature for use in step 14.
- Preheat a second shaker at 56°C for use in step 26. (Alternatively, equilibrate the first shaker to 56°C after step 14).
- Resuspend Magnetic Bead Suspension (from the QIAGEN QIAamp MinElute ccfDNA Midi Kit) by pulse-vortexing for 1 min.
Note: Do not let the suspension settle for more than 2 min before use. Pipette from the centre of the suspension.
Prepare the buffers for extraction:
- Add 8 ml isopropanol (100%) to 12 ml Buffer ACB concentrate to obtain 20 ml Buffer ACB. Mix well after adding isopropanol.
- Add 30 ml ethanol (96–100%) to 13 ml Buffer ACW2 concentrate to obtain 43 ml Buffer ACW2. Mix well after adding ethanol.
Mix the following components according to the instructions below in a 15 ml tube:
| Component | Volume for 1 ml plasma (µl) | Volume for 2 ml plasma (µl) | Volume for 3 ml plasma (µl) | Volume for 4 ml plasma (µl) | Volume for 5 ml plasma (µl) |
|---|---|---|---|---|---|
| Plasma | 1,000 | 2,000 | 3,000 | 4,000 | 5,000 |
| Magnetic Bead Suspension | 30 | 60 | 90 | 120 | 150 |
| Proteinase K | 55 | 110 | 165 | 220 | 275 |
| Bead Binding Buffer | 150 | 300 | 450 | 600 | 750 |
| Total volume | 1,235 | 2,470 | 3,705 | 4,940 | 6,175 |
Incubate the reaction for 10 min at room temperature while shaking (at a slow speed) end-over-end.
Spin the tube down briefly (30 seconds at 200 x g) to remove any solution in the cap.
Place the tube containing bead solution into a magnetic rack for 15 ml tubes. Let the tube stand for at least 1 min, until the solution is clear.
Remove and discard supernatant.
Remove the tube from the magnetic rack and add 200 µl of Bead Elution Buffer to the bead pellet. Vortex to resuspend beads, and pipette up and down to mix and rinse residual beads from the tube wall.
Transfer the full volume of mixture (including the beads) into a Bead Elution Tube.
Incubate for 5 min on a shaker for microcentrifuge tubes at room temperature and 300 rpm.
Note: If the same shaker for microcentrifuge tubes is to be used in step 26, remove the tubes after the room temperature incubation and equilibrate the shaker to 56°C
Place the Bead Elution Tube containing the bead solution into a magnetic rack for 2 ml tubes. Let the tube stand for at least 1 min, until the solution is clear.
Transfer the supernatant into a new Bead Elution tube. Discard the bead pellet.
Avoid transferring any magnetic beads in this step. Carryover may result in reduced cfDNA yield.
Add 300 µl Buffer ACB to the Bead Elution tube containing the supernatant, and vortex to mix. Briefly centrifuge the tube to remove drops from inside the lid.
Pipette the supernatant–Buffer ACB mixture from the previous step into a QIAamp UCP MinElute column.
Centrifuge for 1 min at 6,000 x g.
Place the QIAamp UCP MinElute column into a clean 2 ml collection tube, and discard the flow-through.
Add 500 µl Buffer ACW2 to the QIAamp UCP MinElute column.
Centrifuge for 1 min at 6,000 x g.
Place the QIAamp UCP MinElute column into a clean 2 ml collection tube, and discard the flow-through.
Centrifuge the QIAamp UCP MinElute column at 20,000 x g for 3 min.
Place the QIAamp UCP MinElute column into a new 1.5 ml elution tube and discard the 2 ml collection tube.
Open the lid of the tube and incubate the assembly in a shaker for microcentrifuge tubes at 56°C for 3 min to dry the membrane completely.
Carefully pipette 23 µl of ultra-clean water into the centre of the membrane. Close the lid and incubate at room temperature for 1 min.
Centrifuge at 20,000 x g for 1 min to elute the DNA.
To maximise yield from the elution, place the QIAamp UCP MinElute column in a clean 1.5 ml elution tube. Aspirate the eluate from the previous step and reload it onto the centre of the membrane. Close the lid and incubate for 1 min at room temperature.
Centrifuge at 20,000 x g for 1 min to elute the DNA.
Quantify 1 µl of eluted sample using a Qubit fluorometer.
From 1 ml of plasma, you can expect a yield of ≥ 6 ng cfDNA.
We recommend that the fragment length profiles of extracted cfDNA samples are analysed using a Femto Pulse (Agilent), or equivalent:

Fragment length profile of extracted cfDNA, run on a Femto Pule (Agilent). This example shows the characteristic nucleosome peaks with minimal gDNA contamination.
Take forward ≥6 ng of recovered cfDNA, for each of the 12 samples, to the library preparation stage of the protcol.
4. DNA repair and end-prep
Materials
- ≥6 ng of recovered human cfDNA per sample (12 samples)
- AMPure XP Beads (AXP)
Consumables
- NEBNext® FFPE DNA Repair v2 Module (NEB, E7360)
- NEBNext® Ultra™ II End Repair/dA-Tailing Module (NEB, E7546)
- Qubit dsDNA HS Assay Kit (Invitrogen, Q32851)
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- Freshly prepared 80% ethanol in nuclease-free water
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- 0.2 ml thin-walled PCR tubes
- 1.5 ml Eppendorf DNA LoBind tubes
Equipment
- P1000 pipette and tips
- P100 pipette and tips
- P10 pipette and tips
- Microfuge
- Thermal cycler
- Hula mixer (gentle rotator mixer)
- Magnetic separation rack
- Ice bucket with ice
Optional equipment
- Qubit™ fluorometer (or equivalent for QC check)
Prepare the NEBNext FFPE DNA Repair Mix, the NEBNext FFPE DNA Repair Buffer v2 and NEBNext Ultra II End Repair / dA-tailing Module reagents in accordance with manufacturer’s instructions, and place on ice.
For optimal performance, NEB recommend the following:
Thaw all reagents on ice.
Flick and/or invert the reagent tubes to ensure they are well mixed.
Note: Do not vortex the FFPE DNA Repair Mix, NEB Thermoliable Proteinase K or Ultra II End Prep Enzyme Mix.Always spin down tubes before opening for the first time each day.
The FFPE DNA Repair Buffer v2 may have a little precipitate. Allow the mixture to come to room temperature and pipette the buffer up and down several times to break up the precipitate, followed by vortexing the tube for 30 seconds to solubilise any precipitate.
Note: It is important the buffer is mixed well by vortexing.The FFPE DNA Repair Buffer v2 may have a yellow tinge and is fine to use if yellow.
Use ≥6 ng of recovered cfDNA per sample as input. Inputs should be normalised so that the same mass of sample is used in each reaction.
Prepare your cfDNA samples in nuclease-free water:
For each sample, ensure you have ≥6 ng of extracted cfDNA from the sample extraction, and transfer this into a 0.2 ml thin-walled PCR tube.
Note: You should have 12 sample tubes.Adjust the volume to 20 μl with nuclease-free water.
Mix thoroughly by pipetting up and down, or by flicking the tube.
Spin down briefly in a microfuge.
In each of the 0.2 ml thin-walled PCR tube containing your cfDNA samples, mix the following:
| Reagent | Volume |
|---|---|
| cfDNA from the previous step | 20 µl |
| NEBNext FFPE DNA Repair Buffer v2 | 3 µl |
| NEBNext FFPE DNA Repair Mix | 0.9 µl |
| Total | 23.9 µl |
Thoroughly mix the reaction by gently pipetting and briefly spinning down.
Using a thermal cycler with a heated lid set to 50°C, incubate the reaction at 37°C for 15 minutes and hold at 4°C.
Remove the reaction from the thermal cycler and place the tubes on ice.
Keeping the tubes on ice, add 0.9 µl of NEBNext Thermolabile Proteinase K directly to each of the repaired reaction mixtures.
Mix by pipetting 10 times, followed by spinning down quickly to collect all liquid from the sides of the tube.
Using a thermal cycler with a heated lid set to 75°C, incubate at 37°C for 15 minutes and 65°C for 5 minutes, then hold at 4°C.
Remove the reaction from the thermal cycler and place the tubes on ice.
Keeping the tubes on ice, add 1.3 µl of NEBNext Ultra II End Prep Enzyme Mix directly to the reaction mixture for a total volume of 26.1 µl.
Mix by pipetting 10 times, followed by spinning down quickly to collect all liquid from the sides of the tube.
Using a thermal cycler with a heated lid set to 75°C, incubate at 20°C for 30 minutes and 65°C for 30 minutes, then hold at 4°C.
Resuspend the AMPure XP Beads (AXP) by vortexing.
Transfer each cfDNA sample into a separate clean 1.5 ml Eppendorf DNA LoBind tube.
Add 80 µl of resuspended the AMPure XP Beads (AXP) to each end-prep reaction and mix by flicking the tube.
Incubate on a Hula mixer (rotator mixer) for 5 minutes at room temperature.
Prepare 5 ml of fresh 80% ethanol in nuclease-free water.
Note: Ensure you prepare sufficient 80% ethanol for your 12 samples.
Spin down the sample and pellet on a magnet until the supernatant is clear and colourless. Keep the tube on the magnet, and pipette off the supernatant.
Keep the tube on the magnet and wash the beads with 200 µl of freshly prepared 80% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
If the pellet was disturbed, wait for beads to pellet again before removing the ethanol.
Repeat the previous step.
Spin down and place the tube back on the magnet. Pipette off any residual ethanol. Allow to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
For each sample, remove the tube from the magnetic rack and resuspend the pellet in 10 µl nuclease-free water. Incubate for 2 minutes at room temperature.
Pellet the beads on a magnet until the eluate is clear and colourless, for at least 1 minute.
For each sample, remove and retain 10 µl of eluate into a separate clean 1.5 ml Eppendorf DNA LoBind tube.
Note: Ensure your samples are processed separately. At this stage they are not yet barcoded.
Quantify 1 µl of each eluted sample using a Qubit fluorometer.
Note: You should expect to recover approximately 75% of your input mass. For example, from 6 ng of cfDNA, a yield of approximately 4.5 ng is expected.
Take forward the repaired and end-prepped cfDNA into the native barcode ligation step. However, at this point it is also possible to store the sample at 4°C overnight.
5. Native barcode ligation
Materials
- Native Barcodes (NB01-24)
- AMPure XP Beads (AXP)
- EDTA (EDTA)
Consumables
- NEB Blunt/TA Ligase Master Mix (NEB, M0367)
- Freshly prepared 80% ethanol in nuclease-free water
- Nuclease-free water (e.g. Thermo Scientific, AM9937)
- 1.5 ml Eppendorf DNA LoBind tubes
- Eppendorf twin.tec® PCR plate 96 LoBind, semi-skirted (Merck, EP0030129504) with heat seals
- OR 0.2 ml thin-walled PCR tubes
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- Qubit™ dsDNA HS Assay Kit (ThermoFisher, Q32851)
Equipment
- Magnetic separation rack
- Vortex mixer
- Hula mixer (gentle rotator mixer)
- Microfuge
- Thermal cycler
- Ice bucket with ice
- Multichannel pipette and tips
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- P2 pipette and tips
- Qubit™ fluorometer (or equivalent for QC check)
Prepare the NEB Blunt/TA Ligase Master Mix according to the manufacturer's instructions, and place on ice:
Thaw the reagents at room temperature.
Spin down the reagent tubes for 5 seconds.
Ensure the reagents are fully mixed by performing 10 full volume pipette mixes.
Thaw the EDTA at room temperature and mix by vortexing. Then spin down and place on ice.
Thaw the Native Barcodes (NB01-24) at room temperature. Briefly spin down, individually mix the barcodes required for your number of samples by pipetting, and place them on ice.
The wells of the barcoding plate are intended for single use only. Please ensure your barcode well is sealed before use, and do not reuse the barcode well once pierced/opened.
Sample inputs should be normalised so the same mass of sample is used in each reaction.
Use the Qubit quantification results from the end of the "DNA repair and end-prep" stage of this protocol to ensure you are taking forward an equivalent mass for each sample.
Normalise and prepare your end-prepped cfDNA samples in nuclease-free water:
Inputs should be normalised so the same mass of sample is used in each reaction.
For each sample, take forward an equivalent mass of sample into a separate clean 0.2 ml thin-walled PCR tube.
Note: To take forward the maximum amount of DNA possible, we recommend taking the full volume of your lowest concentration sample and normalising the other samples to this.Adjust the volume of each sample to 7.5 μl with nuclease-free water.
Mix thoroughly by pipetting up and down, or by flicking the tube(s).
Spin down briefly in a microfuge.
Select a unique barcode for each sample to be run together on the same flow cell. 12 samples should be barcoded and combined in one experiment.
Please note: Only use one barcode per sample.
In the 0.2 ml PCR-tubes containing your normalised sample inputs, add the reagents in the following order for each sample:
| Reagent | Volume |
|---|---|
| End-prepped DNA | 7.5 µl |
| Native Barcode (NB01-24) | 2.5 µl |
| Blunt/TA Ligase Master Mix | 10 µl |
| Total | 20 µl |
Thoroughly mix the reaction by gently pipetting and briefly spinning down.
Incubate for 20 minutes at room temperature.
Add 4 µl of EDTA (blue cap) to each well and mix thoroughly by pipetting and spin down briefly.
EDTA is added at this step to stop the reaction.
Pool all the barcoded samples in a 1.5 ml Eppendorf DNA LoBind tube.
The pooled volume for the 12 samples should be 288 µl.
We recommend checking the base of your tubes/plate are all the same volume before pooling and after to ensure all the liquid has been taken forward.
Resuspend the AMPure XP Beads (AXP) by vortexing.
Add 1.2X (346 µl) AMPure XP Beads (AXP) to the pooled reaction, and mix by pipetting.
Incubate on a Hula mixer (rotator mixer) for 10 minutes at room temperature.
Prepare 2 ml of fresh 80% ethanol in nuclease-free water.
Spin down the sample and pellet on a magnet for 5 minutes. Keep the tube on the magnetic rack until the eluate is clear and colourless, and pipette off the supernatant.
Keep the tube on the magnetic rack and wash the beads with 700 µl of freshly prepared 80% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
If the pellet was disturbed, wait for beads to pellet again before removing the ethanol.
Repeat the previous step.
Spin down and place the tube back on the magnetic rack. Pipette off any residual ethanol. Allow the pellet to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
Remove the tube from the magnetic rack and resuspend the pellet in 32 µl nuclease-free water by gently flicking.
Incubate for 10 minutes at 37°C. Every 2 minutes, agitate the sample by gently flicking for 10 seconds to encourage DNA elution.
Pellet the beads on a magnetic rack until the eluate is clear and colourless.
Remove and retain 32 µl of eluate into a clean 1.5 ml Eppendorf DNA LoBind tube.
Quantify 1 µl of eluted sample using a Qubit fluorometer.
Note: You should expect to recover between 40–60 % of the input DNA mass. For example, if starting with 6 ng per sample and using 12x barcodes (72 ng total), a yield of approximately 30–40 ng, is expected.
Take forward the barcoded DNA library to the adapter ligation and clean-up step. However, you may store the sample at 4°C overnight.
6. Adapter ligation and clean-up
Materials
- S Fragment Buffer (SFB)
- Elution Buffer (EB)
- Native Adapter (NA)
- AMPure XP Beads (AXP)
Consumables
- NEBNext® Quick Ligation Module (NEB, E6056)
- 1.5 ml Eppendorf DNA LoBind tubes
- Qubit™ Assay Tubes (Invitrogen, Q32856)
- Qubit™ dsDNA HS Assay Kit (ThermoFisher, Q32851)
Equipment
- Microfuge
- Magnetic separation rack
- Vortex mixer
- Hula mixer (gentle rotator mixer)
- Thermal cycler
- P1000 pipette and tips
- P200 pipette and tips
- P100 pipette and tips
- P20 pipette and tips
- P10 pipette and tips
- Ice bucket with ice
- Qubit™ fluorometer (or equivalent for QC check)
The Native Adapter (NA) used in this kit and protocol is not interchangeable with other sequencing adapters.
Check your flow cell.
We recommend performing a flow cell check before starting adapter ligation and clean-up to ensure you have a flow cell with sufficient pores for a good sequencing run.
See the flow cell check document for more information.
Prepare the NEBNext Quick Ligation Reaction Module according to the manufacturer's instructions, and place on ice:
Thaw the reagents at room temperature.
Spin down the reagent tubes for 5 seconds.
Ensure the reagents are fully mixed by performing 10 full volume pipette mixes. Note: Do NOT vortex the Quick T4 DNA Ligase.
The NEBNext Quick Ligation Reaction Buffer (5x) may have a little precipitate. Allow the mixture to come to room temperature and pipette the buffer up and down several times to break up the precipitate, followed by vortexing the tube for several seconds to ensure the reagent is thoroughly mixed.
Do not vortex the Quick T4 DNA Ligase.
Spin down the Native Adapter (NA) and Quick T4 DNA Ligase, pipette mix and place on ice.
Thaw the Elution Buffer (EB) at room temperature and mix by vortexing. Then spin down and place on ice.
Thaw the Short Fragment Buffer (SFB) at room temperature and mix by vortexing. Then spin down and place on ice.
In a 1.5 ml Eppendorf LoBind tube, mix in the following order:
Between each addition, pipette mix 10 - 20 times.
| Reagent | Volume |
|---|---|
| Pooled barcoded sample | 30 µl |
| Native Adapter (NA) | 5 µl |
| NEBNext Quick Ligation Reaction Buffer (5X) | 10 µl |
| Quick T4 DNA Ligase | 5 µl |
| Total | 50 µl |
Thoroughly mix the reaction by gently pipetting and briefly spinning down.
Incubate the reaction for 20 minutes at room temperature.
The next clean-up step uses Short Fragment Buffer (SFB) rather than 80% ethanol to wash the beads. The use of ethanol will be detrimental to the sequencing reaction.
Resuspend the AMPure XP Beads (AXP) by vortexing.
Add 60 µl (1.2x) of resuspended AMPure XP Beads (AXP) to the reaction and mix by pipetting.
Incubate on a Hula mixer (rotator mixer) for 10 minutes at room temperature.
Spin down the sample and pellet on the magnetic rack. Keep the tube on the magnet and pipette off the supernatant.
Wash the beads by adding 250 μl of Short Fragment Buffer (SFB). Flick the beads to resuspend, spin down, then return the tube to the magnetic rack and allow the beads to pellet. Remove the supernatant using a pipette and discard.
Note: Take care when removing the supernatant, the viscosity of the buffer can contribute to loss of beads from the pellet.
Repeat the previous step.
Spin down and place the tube back on the magnet. Pipette off any residual supernatant. Allow to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
Remove the tube from the magnetic rack and resuspend the pellet in 33 µl of Elution Buffer (EB).
Spin down and incubate for 10 minutes at 37°C. Every 2 minutes, agitate the sample by gently flicking for 10 seconds to encourage DNA elution.
Pellet the beads on a magnet until the eluate is clear and colourless, for at least 1 minute.
Remove and retain 33 µl of eluate containing the DNA library into a clean 1.5 ml Eppendorf DNA LoBind tube.
Dispose of the pelleted beads
Quantify 1 µl of eluted sample using a Qubit fluorometer.
Note: You should expect to recover approximately 20% of input mass. For example, if starting with 6 ng per sample and using 12x barcodes (72 ng total), a yield of approximately 15 ng is expected.
The prepared library is used for loading into the flow cell. Store the library on ice or at 4°C until ready to load.
Library storage recommendations
We recommend storing libraries in Eppendorf DNA LoBind tubes at 4°C for short-term storage or repeated use, for example, re-loading flow cells between washes. For single use and long-term storage of more than 3 months, we recommend storing libraries at -80°C in Eppendorf DNA LoBind tubes.
7. Priming and loading the PromethION Flow Cell
Materials
- Sequencing Buffer (SB)
- Library Beads (LIB)
- Flow Cell Tether (FCT)
- Flow Cell Flush (FCF)
Consumables
- PromethION Flow Cells
- 1.5 ml Eppendorf DNA LoBind tubes
Equipment
- PromethION 2 Solo or PromethION 24/48 device
- PromethION device
- PromethION Flow Cell Light Shield
- P1000 pipette and tips
- P200 pipette and tips
- P20 pipette and tips
This method is only compatible with R10.4.1 flow cells (FLO-PRO114M).
After taking the flow cell out of the fridge, wait 20 minutes for the flow cell to reach room temperature before inserting it into the PromethION. Condensation can form on the flow cell in humid environments. Inspect the gold connector pins on the top and underside of the flow cell for condensation and wipe off with a lint-free wipe if any is observed. Ensure the heat pad (black pad) is present on the underside of the flow cell.
Thaw the Sequencing Buffer (SB), Library Beads (LIB), Flow Cell Tether (FCT) and Flow Cell Flush (FCF) at room temperature before mixing by vortexing. Then spin down and store on ice.
To prepare the flow cell priming mix, combine Flow Cell Tether (FCT) and Flow Cell Flush (FCF), as directed below. Mix by vortexing at room temperature.
In a clean suitable tube for the number of flow cells, combine the following reagents:
| Reagent | Volume per flow cell |
|---|---|
| Flow Cell Flush (FCF) | 1,170 µl |
| Flow Cell Tether (FCT) | 30 µl |
| Total volume | 1,200 µl |
For PromethION 2 Solo, load the flow cell(s) as follows:
Place the flow cell flat on the metal plate.
Slide the flow cell into the docking port until the gold pins or green board cannot be seen.

For the PromethION 24/48, load the flow cell(s) into the docking ports:
- Line up the flow cell with the connector horizontally and vertically before smoothly inserting into position.
- Press down firmly onto the flow cell and ensure the latch engages and clicks into place.
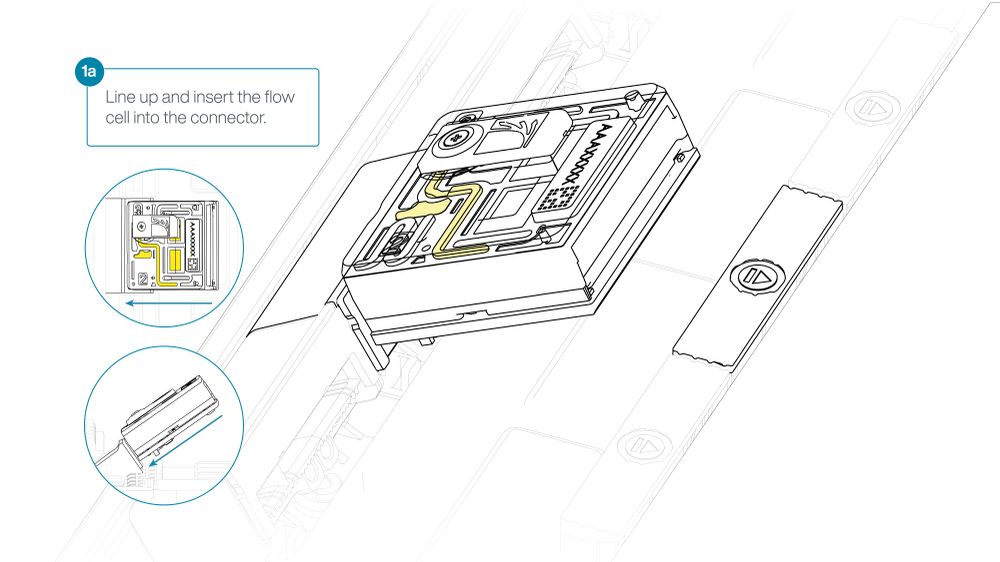
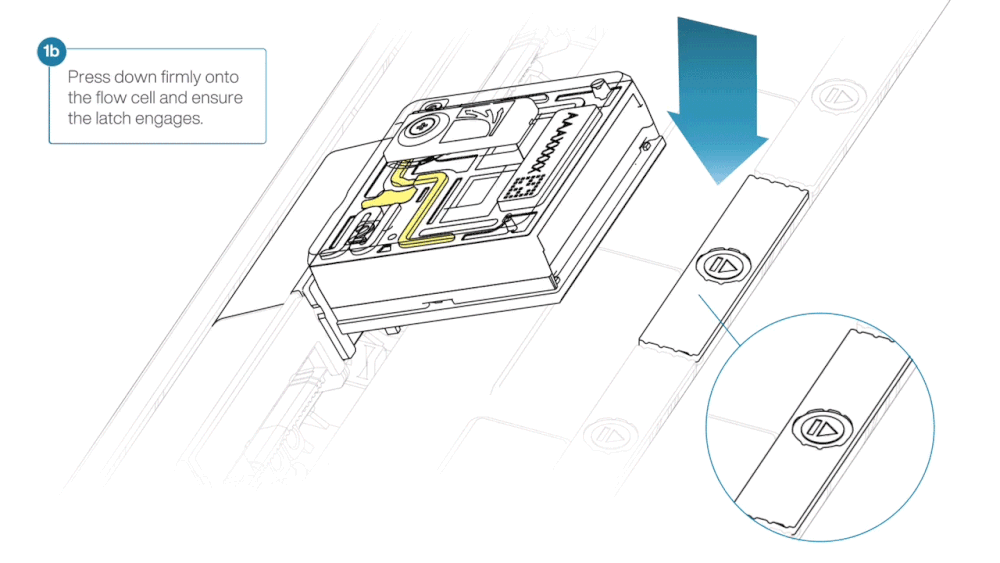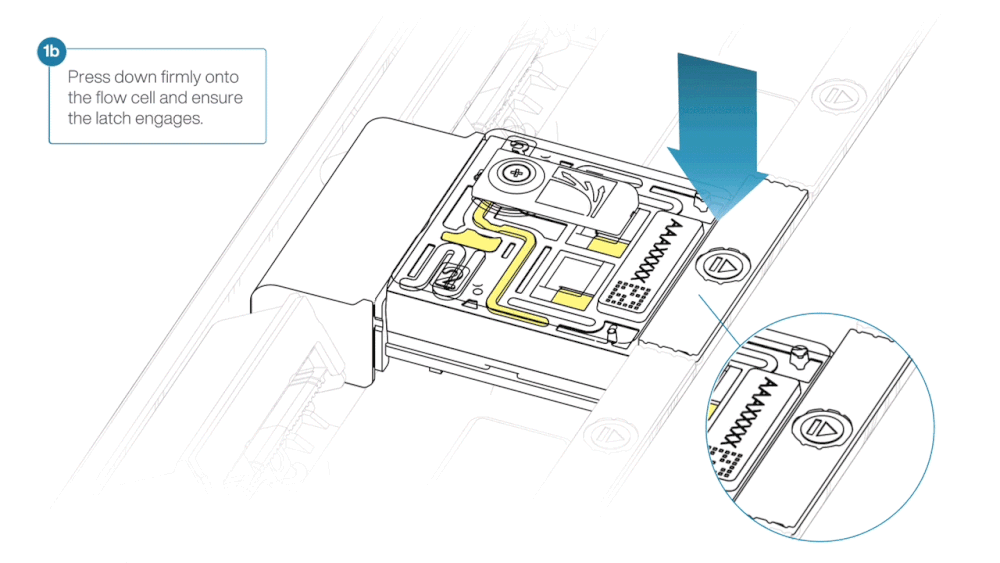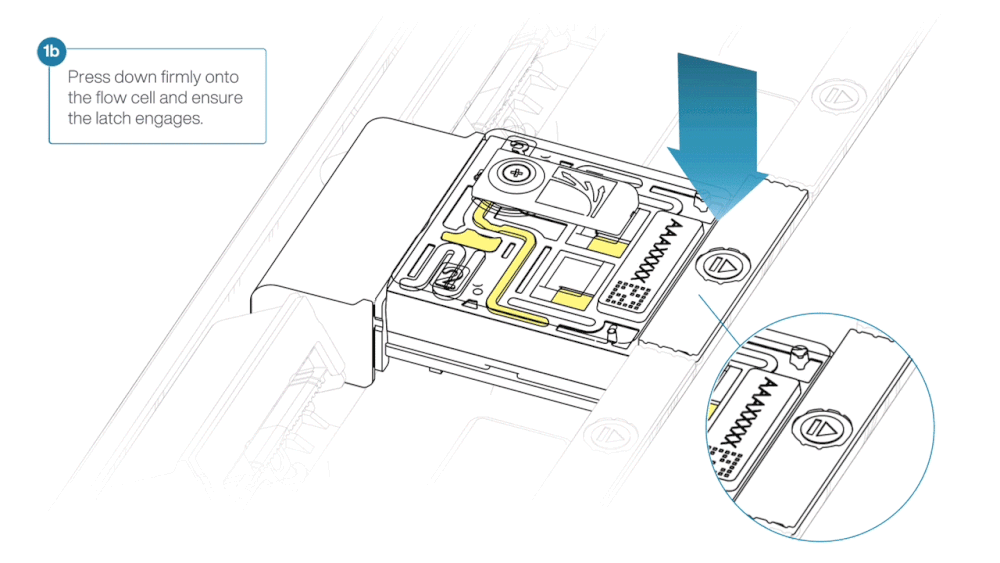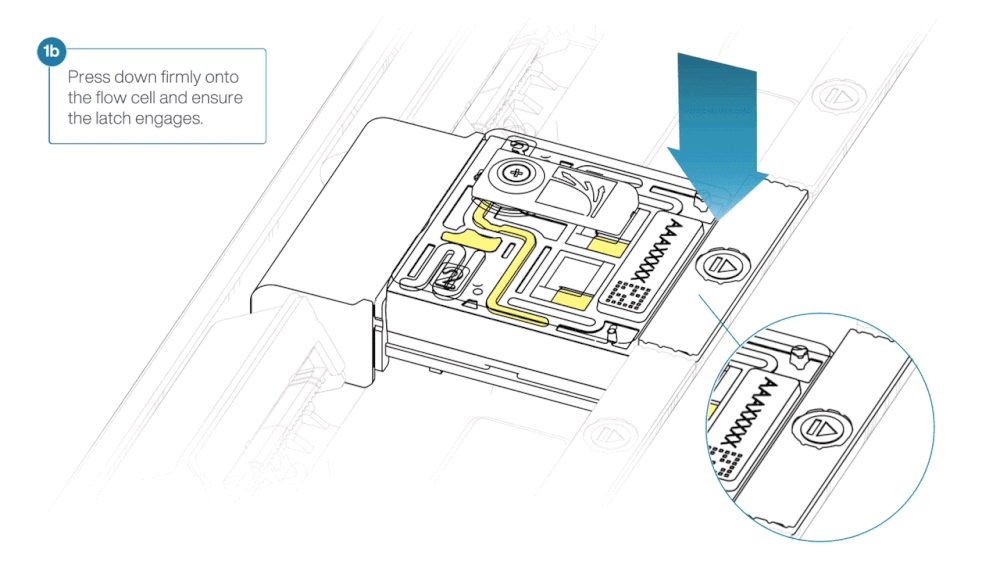
Insertion of the flow cells at the wrong angle can cause damage to the pins on the PromethION and affect your sequencing results. If you find the pins on a PromethION position are damaged, please contact support@nanoporetech.com for assistance.

Complete a flow cell check to assess the number of pores available before loading the library.
This step can be omitted if the flow cell has been checked previously.
See the flow cell check document for more information.
Slide the inlet port cover clockwise to open.
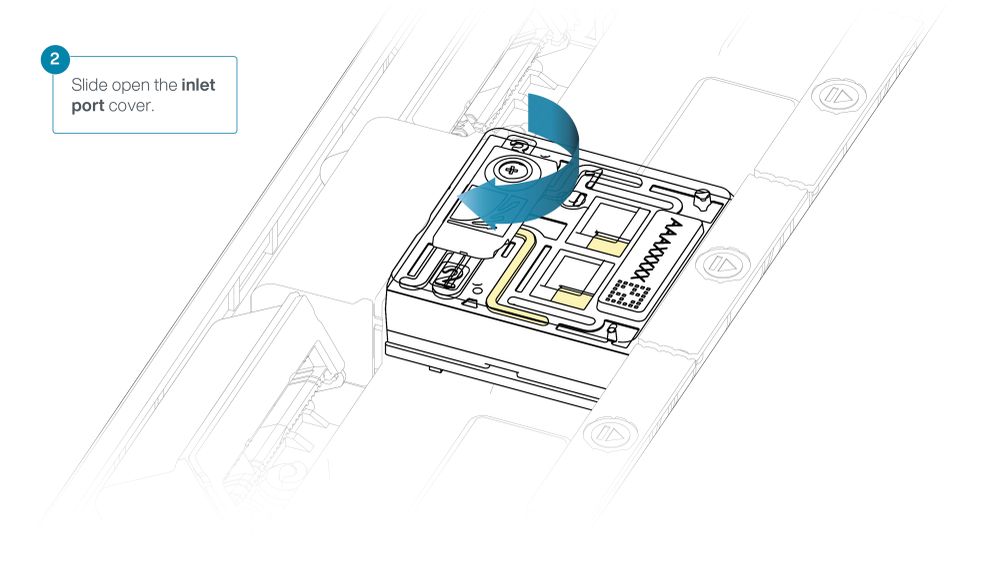
Take care when drawing back buffer from the flow cell. Do not remove more than 20-30 µl, and make sure that the array of pores are covered by buffer at all times. Introducing air bubbles into the array can irreversibly damage pores.
After opening the inlet port, draw back a small volume to remove any air bubbles:
- Set a P1000 pipette tip to 200 µl.
- Insert the tip into the inlet port.
- Turn the wheel until the dial shows 220-230 µl, or until you see a small volume of buffer entering the pipette tip.

Load 500 µl of the priming mix into the flow cell via the inlet port, avoiding the introduction of air bubbles. Wait five minutes. During this time, prepare the library for loading using the next steps in the protocol.

Thoroughly mix the contents of the Library Beads (LIB) by pipetting.
The Library Beads (LIB) tube contains a suspension of beads. These beads settle very quickly. It is vital that they are mixed immediately before use.
We recommend using the Library Beads (LIB) for most sequencing experiments. However, the Library Solution (LIS) is available for more viscous libraries.
In a new 1.5 ml Eppendorf DNA LoBind tube, prepare the library for loading as follows:
| Reagent | Volume per flow cell |
|---|---|
| Sequencing Buffer (SB) | 100 µl |
| Library Beads (LIB) thoroughly mixed before use | 68 µl |
| DNA library | 32 µl |
| Total | 200 µl |
Note: The prepared library is used for loading into the flow cell. Store the library on ice or at 4°C until ready to load.
Complete the flow cell priming by slowly loading 500 µl of the priming mix into the inlet port.

Mix the prepared library gently by pipetting up and down just prior to loading.
Load 200 µl of library into the inlet port using a P1000 pipette.

Close the valve to seal the inlet port.
For optimal sequencing output, install the light shield on your flow cell as soon as the library has been loaded.
We recommend leaving the light shield on the flow cell when library is loaded, including during any washing and reloading steps. The shield can be removed when the library has been removed from the flow cell.
If the light shield has been removed from the flow cell, install the light shield as follows:
- Align the inlet port cut out of the light shield with the inlet port cover on the flow cell. The leading edge of the light shield should sit above the flow cell ID.
- Firmly press the light shield around the inlet port cover. The inlet port clip will click into place underneath the inlet port cover.


Close the PromethION lid when ready to start a sequencing run on MinKNOW.
Wait a minimum of 10 minutes after loading the flow cells onto the PromethION before initiating any experiments. This will help to increase the sequencing output.
8. Data acquisition and basecalling
Ensure you are using the most recent version of MinKNOW.
We recommend updating MinKNOW to the latest version prior to starting a sequencing run for the best sequencing results.
For more information on updating MinKNOW, please refer to our MinKNOW protocol.
How to start sequencing
Once you have loaded your flow cell, the sequencing run can be started on MinKNOW, our sequencing software that controls the device, data acquisition and real-time basecalling.
Please ensure MinKNOW is installed on your computer or device. Further instructions for setting up a sequencing run can be found in the MinKNOW protocol.
Please ensure when setting up a sequencing run you are using the recommendations outlined below. All other parameters can be left to their default settings.
MinKNOW can be used and set up to sequence in multiple ways:
- On a computer either direcly or remotely connected to a sequencing device.
- Directly on a PromethION 24/48 sequencing device.
For more information on using MinKNOW on a sequencing device, please see the device user manuals:
Open the MinKNOW software using the desktop shortcut and log into the MinKNOW software using your Community credentials.
Click on your connected device.
Set up a sequencing run by clicking Start sequencing.
Type in the experiment name, select the flow cell postition and enter sample ID. Choose FLO-PRO114M flow cell type from the drop-down menu.
Click Continue to kit selection.
Select the Native Barcoding Sequencing Kit 24 (SQK-NBD114.24).
An expansion kit does not need to be selected.
Click Continue to run configuration.
Set up basecalling using the following parameters:
- Ensure the basecalling is switched ON.
- Next to "Models", click Edit options and choose High accuracy basecaller (HAC) from the drop-down menu.
- Ensure barcoding is ON.
Click Continue to output.
Keep the output format and filtering options to their default settings.
Click Continue to final review.
Click Start to start sequencing.
You will be automatically navigated to the Sequencing Overview page to monitor the sequencing run.
Data analysis after sequencing
After sequencing has completed on MinKNOW, the flow cell can be reused or returned, as outlined in the Flow cell reuse and returns section.
After sequencing and basecalling, the data can be analysed, as outlined in the Downstream analysis section.
9. Downstream analysis
Bioinformatics analysis
If basecalling is not performed during live sequencing, raw sequencing data (POD5 format) can be processed post-sequencing.
This can be achieved using the tool Dorado, which enables basecalling and subsequent alignment to a reference genome.
Dorado can also detect modified bases by using the modified-bases option (e.g. --modified-bases 5mCG_5hmCG). This will integrate methylation tags directly into the aligned BAM file. We also recommend applying a minimum QScore cutoff (--min-Qscore <min_QScore>), which serves as a quality control measure to ensure only high-quality reads are used in downstream processes.
1. The command below demonstrates how to initiate basecalling with Dorado, followed by sorting, and indexing the output using Samtools. Please see the Dorado documentation here for further details.
Dorado basecaller <model> <input_POD5> --reference <REF> --min-qscore <min_QScore>
| samtools sort -o <OUTPUT_BAM> - && samtools index <OUTPUT_BAM>
For example to SUP basecall with 5mCG and 5hmCG detected in CpG context, and with a QScore filter of 10 we can use:
Dorado basecaller sup --modified-bases 5mCG_5hmCG input.pod5 --reference ref.fasta --min-qscore 10
| samtools sort -o output.bam > - && samtools index output.bam
2. It is also recommended to remove reads that have a poor alignment score i.e. 10. This can be achieved as follows:
samtools view -q <min_map_q> -bh -o <OUTPUT_BAM> <INPUT_BAM> && samtools index -@ <threads> <OUTPUT_BAM>
3. The output from Dorado basecaller can be demultiplexed into per-barcode BAMs using Dorado demux e.g.
Dorado demux --output-dir <output-dir> --no-classify <input-bam>
4. You may optionally omit methylation information from read ends using modkit adjust-mods or modkit tools with --edge-filter option. This may help increase methylation call precision, as the very end of reads, approximately 27 bases, may suffer from loss in methylated bases due to the chemistry used to repair ends in library preparation (see our know-how document for further details).
modkit adjust-mods --edge-filter 0 27 <IN_BAM> <OUTPUT_BAM>
The modified BAM file can be used with external tools that use a BAM file as input for further data analysis and exploration.
Post-basecalling analysis
There are several options for further analysing your basecalled data:
EPI2ME workflows
For in-depth data analysis, Oxford Nanopore Technologies offers a range of bioinformatics tutorials and workflows available in EPI2ME. The platform provides a vehicle where workflows deposited in GitHub by our Research and Applications teams can be showcased with descriptive texts, functional bioinformatics code and example data.
Research analysis tools
Oxford Nanopore Technologies' Research division has created a number of analysis tools, which are available in the Oxford Nanopore GitHub repository. The tools are aimed at advanced users, and contain instructions for how to install and run the software. They are provided as-is, with minimal support.
Community-developed analysis tools
If a data analysis method for your research question is not provided in any of the resources above, please refer to the resource centre and search for bioinformatics tools for your application. Numerous members of the Nanopore Community have developed their own tools and pipelines for analysing nanopore sequencing data, most of which are available on GitHub. Please be aware that these tools are not supported by Oxford Nanopore Technologies, and are not guaranteed to be compatible with the latest chemistry/software configuration.
10. Flow cell reuse and returns
Materials
- Flow Cell Wash Kit (EXP-WSH004)
After your sequencing experiment is complete, if you would like to reuse the flow cell, please follow the Flow Cell Wash Kit protocol and store the washed flow cell at +2°C to +8°C.
The Flow Cell Wash Kit protocol is available on the Nanopore Community.
We recommend you to wash the flow cell as soon as possible after you stop the run. However, if this is not possible, leave the flow cell on the device and wash it the next day.
Alternatively, follow the returns procedure to send the flow cell back to Oxford Nanopore.
Instructions for returning flow cells can be found here.
If you encounter issues or have questions about your sequencing experiment, please refer to the Troubleshooting Guide that can be found in this protocol.
11. Issues during DNA extraction and library preparation
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Low sample quality
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Low DNA purity (Nanodrop reading for DNA OD 260/280 is <1.8 and OD 260/230 is <2.0–2.2) | The DNA extraction method does not provide the required purity | The effects of contaminants are shown in the Contaminants document. Please try an alternative extraction method that does not result in contaminant carryover. Consider performing an additional SPRI clean-up step. |
Low DNA recovery after AMPure bead clean-up
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Low recovery | DNA loss due to a lower than intended AMPure beads-to-sample ratio | 1. AMPure beads settle quickly, so ensure they are well resuspended before adding them to the sample. 2. When the AMPure beads-to-sample ratio is lower than 0.4:1, DNA fragments of any size will be lost during the clean-up. |
| Low recovery | DNA fragments are shorter than expected | The lower the AMPure beads-to-sample ratio, the more stringent the selection against short fragments. Please always determine the input DNA length on an agarose gel (or other gel electrophoresis methods) and then calculate the appropriate amount of AMPure beads to use. 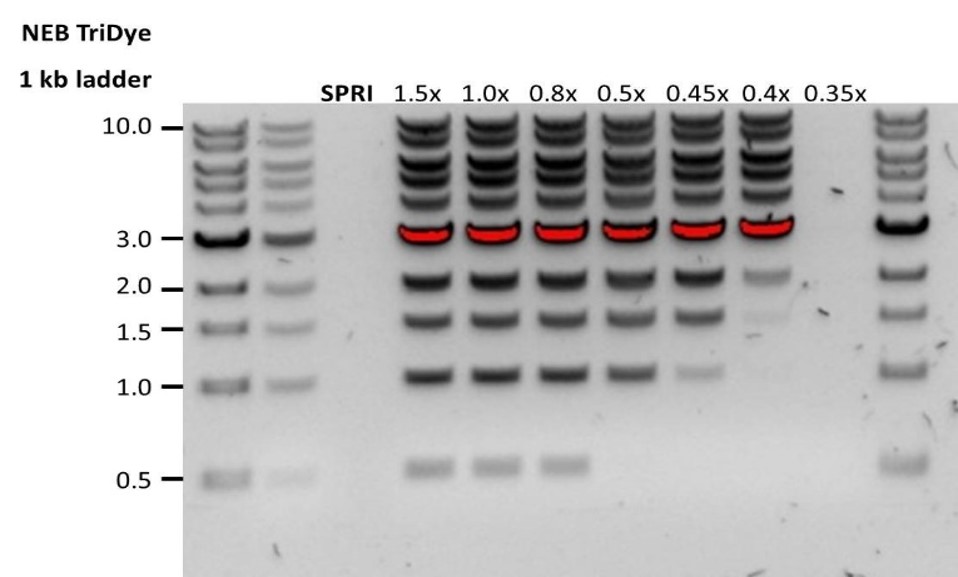 |
| Low recovery after end-prep | The wash step used ethanol <70% | DNA will be eluted from the beads when using ethanol <70%. Make sure to use the correct percentage. |
12. Issues during the sequencing run
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Fewer pores at the start of sequencing than after Flow Cell Check
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | An air bubble was introduced into the nanopore array | After the Flow Cell Check it is essential to remove any air bubbles near the priming port before priming the flow cell. If not removed, the air bubble can travel to the nanopore array and irreversibly damage the nanopores that have been exposed to air. The best practice to prevent this from happening is demonstrated in this video for how to load a PromethION Flow Cell. |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | The flow cell is not correctly inserted into the device | Stop the sequencing run, remove the flow cell from the sequencing device and insert it again, checking that the flow cell is firmly seated in the device and that it has reached the target temperature. If applicable, try a different position on the device (GridION/PromethION). |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | Contaminations in the library damaged or blocked the pores | The pore count during the Flow Cell Check is performed using the QC DNA molecules present in the flow cell storage buffer. At the start of sequencing, the library itself is used to estimate the number of active pores. Because of this, variability of about 10% in the number of pores is expected. A significantly lower pore count reported at the start of sequencing can be due to contaminants in the library that have damaged the membranes or blocked the pores. Alternative DNA/RNA extraction or purification methods may be needed to improve the purity of the input material. The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
MinKNOW script failed
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Script failed" | Restart the computer and then restart MinKNOW. If the issue persists, please collect the MinKNOW log files and contact Technical Support. If you do not have another sequencing device available, we recommend storing the flow cell and the loaded library at 4°C and contact Technical Support for further storage guidance. |
Pore occupancy below 40%
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Pore occupancy <40% | Not enough library was loaded on the flow cell | Ensure the correct volume and concentration as stated on the appropriate protocol for your sequencing library is loaded onto the flow cell. Please quantify the library before loading and calculate fmols using tools like the Promega Biomath Calculator, choosing "dsDNA: µg to fmol" |
| Pore occupancy close to 0 | The Ligation Sequencing Kit was used, and sequencing adapters did not ligate to the DNA | Make sure to use the NEBNext Quick Ligation Module (E6056) and Oxford Nanopore Technologies Ligation Buffer (LNB, provided in the sequencing kit) at the sequencing adapter ligation step, and use the correct amount of each reagent. A Lambda control library can be prepared to test the integrity of the third-party reagents. |
| Pore occupancy close to 0 | The Ligation Sequencing Kit was used, and ethanol was used instead of LFB or SFB at the wash step after sequencing adapter ligation | Ethanol can denature the motor protein on the sequencing adapters. Make sure the LFB or SFB buffer was used after ligation of sequencing adapters. |
| Pore occupancy close to 0 | No tether on the flow cell | Tethers are adding during flow cell priming (FLT tube for Kit 9, 10, 11, FCT for Kit 14, and FTU for ultra-long DNA kits). Make sure FLT/FCT/FTU was added to the buffer (FB for Kit 9, 10, 11, and FCF for Kit 14) before priming. |
Shorter than expected read length
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Shorter than expected read length | Unwanted fragmentation of DNA sample | Read length reflects input DNA fragment length. Input DNA can be fragmented during extraction and library prep. 1. Please review the Extraction Methods in the Nanopore Community for best practice for extraction. 2. Visualise the input DNA fragment length distribution on an agarose gel before proceeding to the library prep. 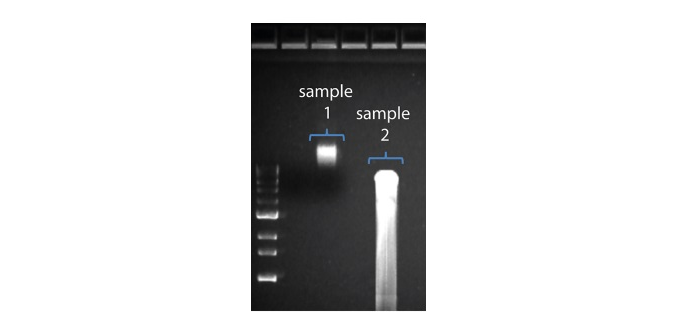 In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented. In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented.3. During library prep, avoid pipetting and vortexing when mixing reagents. Flicking or inverting the tube is sufficient. |
Large proportion of unavailable pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
Large proportion of unavailable pores (shown as blue in the channels panel and pore activity plot) 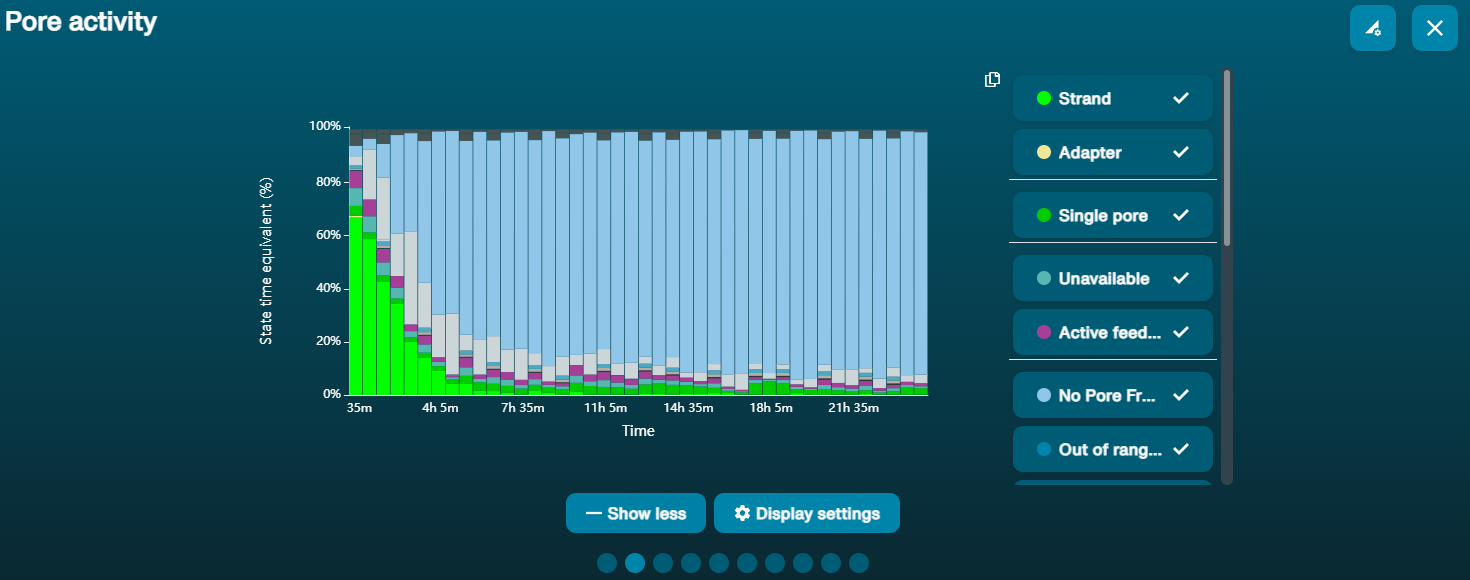 The pore activity plot above shows an increasing proportion of "unavailable" pores over time. The pore activity plot above shows an increasing proportion of "unavailable" pores over time. | Contaminants are present in the sample | Some contaminants can be cleared from the pores by the unblocking function built into MinKNOW. If this is successful, the pore status will change to "sequencing pore". If the portion of unavailable pores stays large or increases: 1. A nuclease flush using the Flow Cell Wash Kit (EXP-WSH004) can be performed, or 2. Run several cycles of PCR to try and dilute any contaminants that may be causing problems. |
Large proportion of inactive pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Large proportion of inactive/unavailable pores (shown as light blue in the channels panel and pore activity plot. Pores or membranes are irreversibly damaged) | Air bubbles have been introduced into the flow cell | Air bubbles introduced through flow cell priming and library loading can irreversibly damage the pores. Watch the how to load a PromethION Flow Cell video for best practice. |
| Large proportion of inactive/unavailable pores | Certain compounds co-purified with DNA | Known compounds include polysaccharides. 1. Clean-up using the QIAGEN PowerClean Pro kit. 2. Perform a whole genome amplification with the original gDNA sample using the QIAGEN REPLI-g kit. |
| Large proportion of inactive/unavailable pores | Contaminants are present in the sample | The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
Temperature fluctuation
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Temperature fluctuation | The flow cell has lost contact with the device | Check that there is a heat pad covering the metal plate on the back of the flow cell. Re-insert the flow cell and press it down to make sure the connector pins are firmly in contact with the device. If the problem persists, please contact Technical Services. |
Failed to reach target temperature
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Failed to reach target temperature" | The instrument was placed in a location that is colder than normal room temperature, or a location with poor ventilation (which leads to the flow cells overheating) | MinKNOW has a default timeframe for the flow cell to reach the target temperature. Once the timeframe is exceeded, an error message will appear and the sequencing experiment will continue. However, sequencing at an incorrect temperature may lead to a decrease in throughput and lower q-scores. Please adjust the location of the sequencing device to ensure that it is placed at room temperature with good ventilation, then re-start the process in MinKNOW. |







