Resource Centre
Bioinformatics tool 
Yanocomp: robust prediction of m6A modifications in individual nanopore direct RNA reads
Bioinformatics tool 
yacrd and fpa: upstream tools for long-read genome assembly
Workflow 
Workflow overview: tumour-normal sequencing
Workflow 
Workflow overview: 24-hour human whole-genome sequencing
Workflow 
Workflow overview: pharmacogenomics with adaptive sampling
Workflow Workflow overview: large cohort sequencing
Workflow 
Workflow overview: human variant calling
Bioinformatics tool 
Whole Human Genome Sequencing Project
Video 
Whole-genome sequencing in PulseNet foodborne molecular surveillance systems
Bioinformatics tool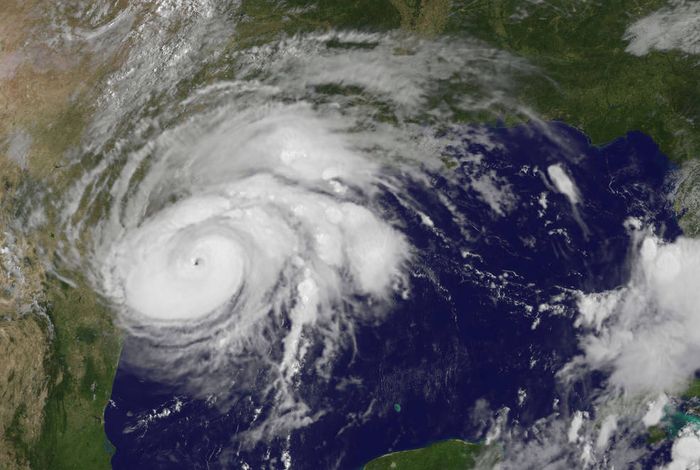 
Whole-Genome Sequencing of a Human Clinical Isolate of emm28 Streptococcus pyogenes Causing Necrotizing Fasciitis Acquired Contemporaneously with Hurricane Harvey
Poster 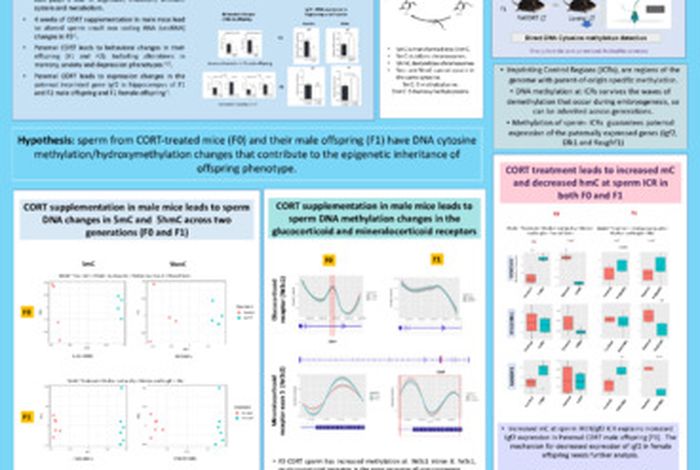
Whole genome Nanopore DNA analysis shows that chronic corticosterone supplementation results in altered sperm DNA methylation.
Poster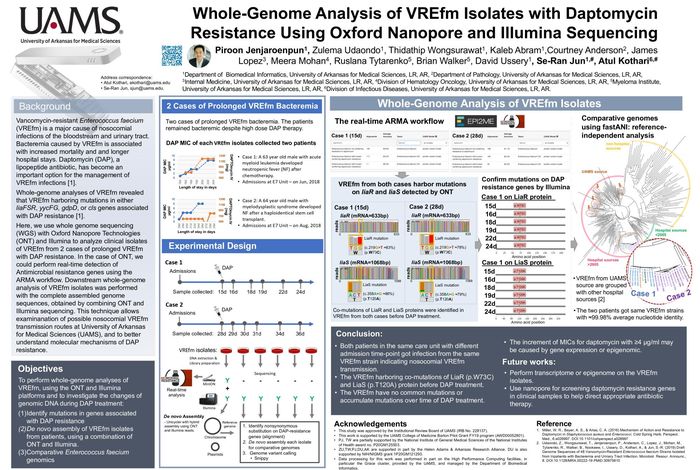 
Whole-genome analysis of VREfm isolates with daptomycin resistance using Oxford Nanopore and Illumina sequencing
Bioinformatics tool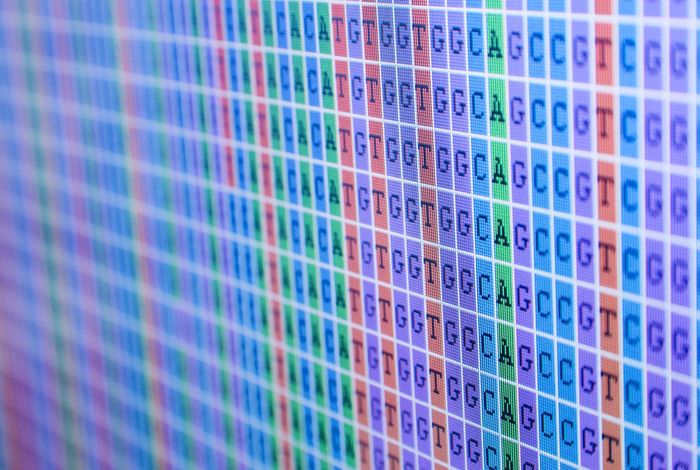 
Wengan: Efficient and high quality hybrid de novo assembly of human genomes
Bioinformatics tool 
WeFaceNano: a user-friendly pipeline for complete ONT sequence assembly and detection of antibiotic resistance in multi-plasmid bacterial isolates
Case study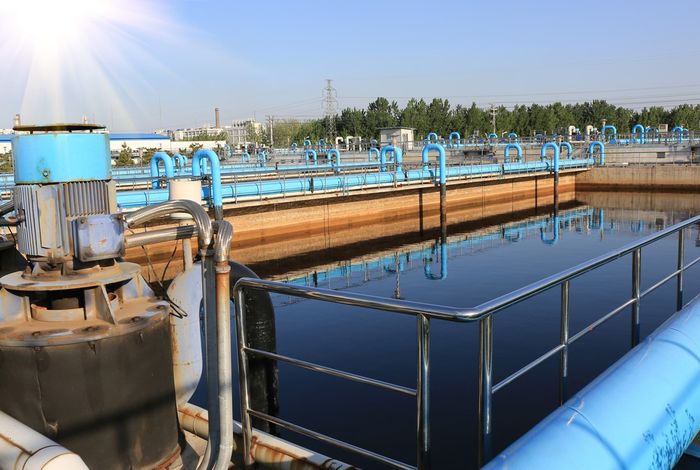 
Wastewater sequencing — an early warning system for infectious disease outbreaks
Publication 
Wakhan: reconstruction of chromosome-scale copy number profiles of tumour genomes with long-read sequencing
Bioinformatics tool 
Vulcan: Improved long-read mapping and structural variant calling via dual-mode alignment
Bioinformatics tool 
VIRUSBreakend: viral integration recognition using single breakends
Publication 
Verkko2: integrating proximity ligation data with long-read De Bruijn graphs for efficient telomere-to-telomere genome assembly, phasing, and scaffold
Bioinformatics tool 
Verkko: telomere-to-telomere assembly of diploid chromosomes






