Advantages of real-time sequencing
Oxford Nanopore sequencing is the only sequencing technology capable of providing insights in real time, which offers several advantages compared to traditional methods.
On-demand and real-time adjustment
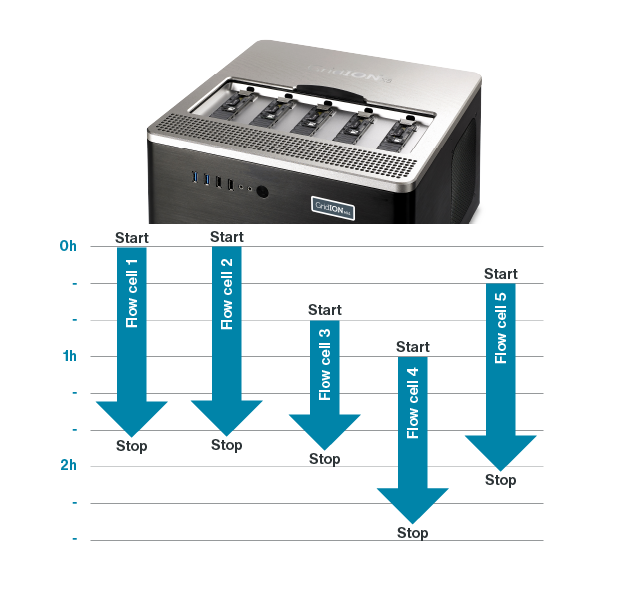
Real-time insights enable the user to adjust their experiment at any stage. For example, during a sequencing run you might identify that you won’t have enough data, or your sample quality is low. Users can stop existing runs or add additional flow cells, without causing disruption.
The ability to add flow cells on demand means that users do not have to wait for sufficient samples to start a run — they can use between one to five GridION, or one to 24 PromethION Flow Cells initially, and then additional flow cells as samples become available.

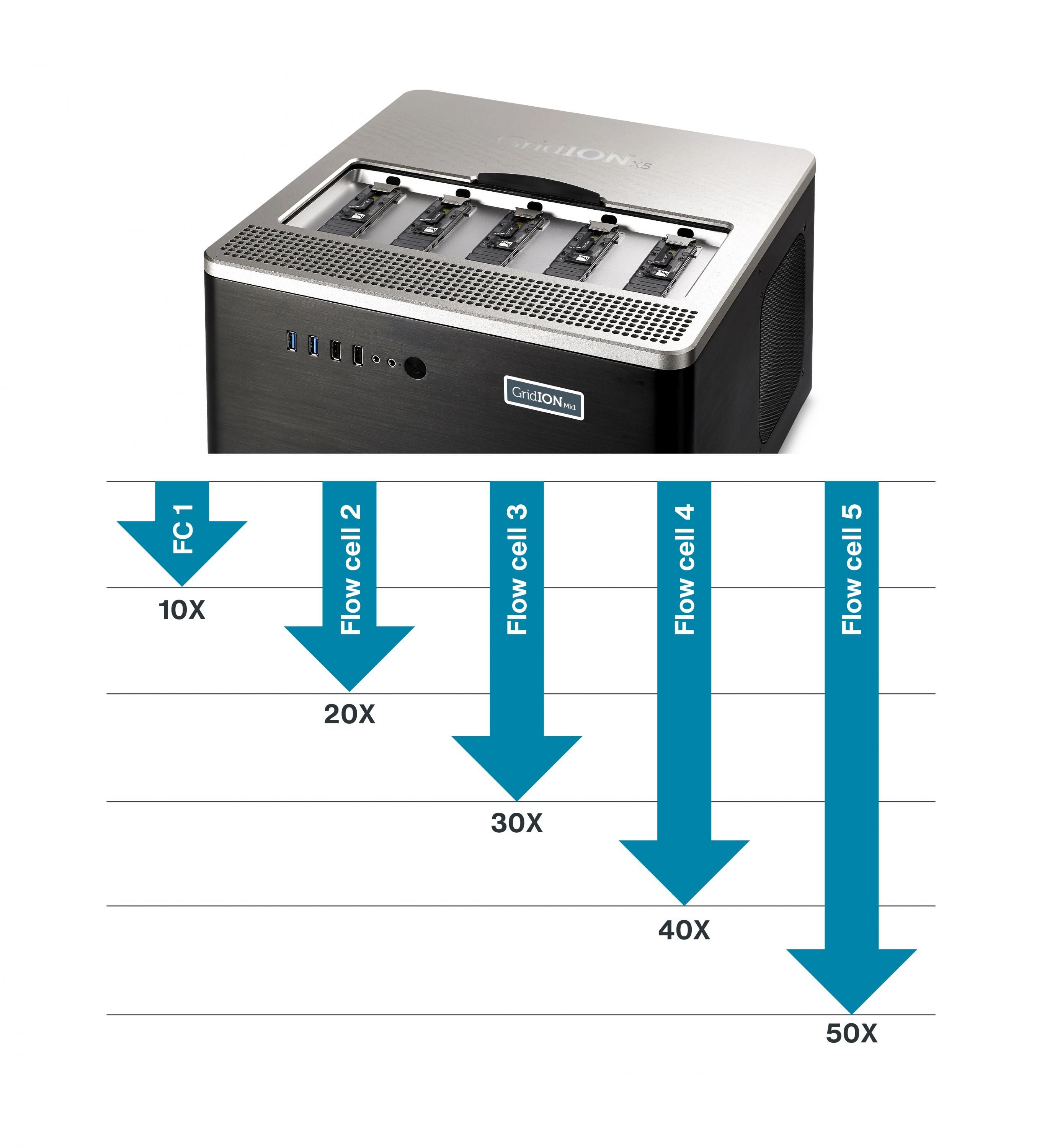
Run until
Run until, as the name suggests, allows users to run their experiment until a predetermined parameter is met. For example: the system may process the sample until a minimum of tenfold read depth over specified regions of interest has been seen, until a specific mutation has been observed in a sample, or until enough sequence data has been collected to reliably assemble a sample against a reference. As there is no fixed run time, the experiment is defined by the user — not by the machine.

Adaptive sampling pushes the boundaries of target enrichment with a completely bioinformatics-driven strategy, allowing target enrichment or depletion of specific genomic regions (or whole genomes) during sequencing itself, with no requirement for target enrichment during sample prep.
As molecules of DNA pass through a nanopore, its sequence is decoded in real time. If the sequence is not of interest it is ejected from the nanopore, leaving the pore free and ready for the next strand. This enables only selected sequences of interest to be analysed, resulting in high coverage of the region of interest.
With adaptive sampling, you can achieve high coverage of native DNA and RNA molecules, enabling DNA and RNA modifications to be preserved and directly sequenced — with no additional library prep steps. Long-range epigenetic changes, structural variants, and single nucleotide polymorphisms can therefore be identified and phased in a single dataset.
Ultimately, as adaptive sampling is a feature in the software, it enables targeted sequencing applications to be carried out without complex targeted sample preparation techniques.
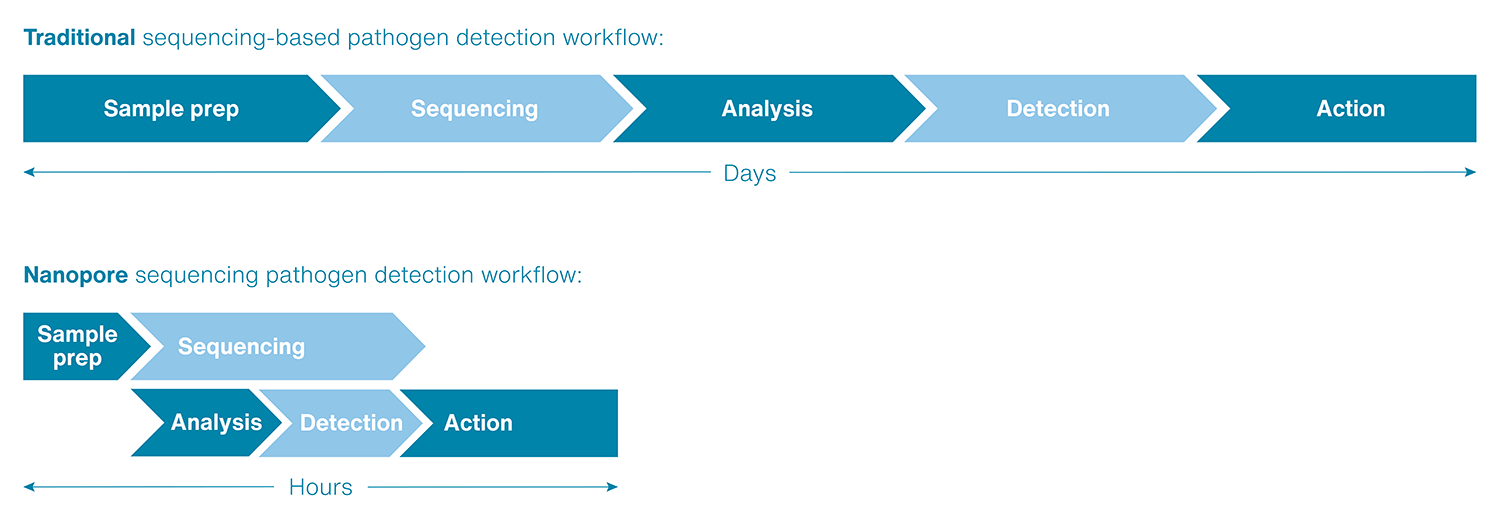
Increase speed of result, such as pathogen detection
Real-time data streaming means rapid access to time-critical results. Unlike traditional methods where analysis can only be done once sequencing is complete, Oxford Nanopore sequencing enables initial insights to be gathered as soon as sequencing starts. This is ideal in situations such as pathogen identification or looking for a microbial resistance gene. The sequencing run can be left to continue to provide greater confidence in your results.
Longer run, greater confidence
Nanopore sequencing is dependent on single molecules of DNA/RNA passing through thousands of nanopores in parallel. As soon as DNA/RNA passes through the pore the data becomes available. A longer sequencing run enables more molecules of DNA/RNA to be sequenced, increasing the likelihood of identifying low abundance sequences within a sample. This results in greater confidence in the result and a greater range of analysis. This is particularly important for applications such as detection of low abundance fusion genes and transcripts, identification of low frequency variants, alleles and biomarkers, as well as differentiating metagenomic communities.
Get in touch
Subscribe
Talk to us
If you have any questions about our products or services, chat directly with a member of our sales team.
Book a sales call
To book a call with one of our sales team, please click below.