Repeat expansions
Span tandem repeats in a single read
Tandem repeats (TRs) are continuously repeated bases and are a type of structural variant (SV). TR expansions are important genetic aberrations that are associated with over 50 neurodegenerative disorders. The study of such diseases requires accurate size determination of the expanded repeat, which has proven to be challenging with legacy sequencing methods such as short-read sequencing. Oxford Nanopore sequencing generates reads of unrestricted length that can span repeat expansions end to end in single reads without the need for PCR, enabling unambiguous size determination.
Featured content

Resolve challenging medically relevant genes
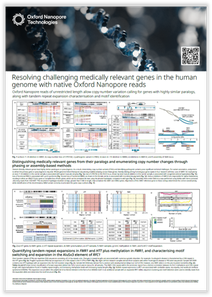
Long nanopore reads allow calling of copy number variants of genes with highly similar paralogs and the enumeration and motif identification of tandem repeat expansions. This poster shows how Oxford Nanopore sequencing can resolve challenging medically relevant genes.
Related content
Resolving repetitive regions with Oxford Nanopore sequencing
Recommended device for repeat expansion detection

PromethION 24
Combine up to 24 independently addressable, high-capacity flow cells with a flexible, large-scale, direct DNA and RNA sequencing device for on-demand access to terabases of data — ideal for high-throughput, whole-genome sequencing with repeat expansion detection and characterisation.