使用 SQK-RBK114(.24 或 .96)进行质粒 DNA 测序 (PRB_9188_v114_revK_07Apr2026)
MinION: Protocol
使用 SQK-RBK114(.24 或 .96)进行质粒 DNA 测序 V PRB_9188_v114_revK_07Apr2026
本方案提供了最便捷高效的质粒 DNA 测序流程。
此实验指南:
- 支持多达96个不同样本的混合测序
- 文库制备用时约60分钟
- 产量高
- 包含片段化步骤
- 兼容 R10.4.1 测序芯片
仅供研究使用
FOR RESEARCH USE ONLY.
概览
本方案提供了最便捷高效的质粒 DNA 测序流程。
此实验指南:
- 支持多达96个不同样本的混合测序
- 文库制备用时约60分钟
- 产量高
- 包含片段化步骤
- 兼容 R10.4.1 测序芯片
仅供研究使用
1. 实验方案概览
快速条形码测序试剂盒的特点
我们推荐有以下需求的用户使用该试剂盒:
- 希望混样测序以降低单个样本的测序成本
- 希望通过免扩增的方式混样建库,以保留碱基修饰等信息
- 希望在短时间内制备文库
- 当前实验器材有限
使用 V14 快速条形码测序试剂盒-24或96 为质粒测序
本实验指南详细描述了使用V14 快速条形码测序试剂盒-24或96(SQK-RBK114.24 或 SQK-RBK114.96)为多达96个质粒样本快速添加条形码并建库测序的操作流程及常见问题的解决方法。该方法可用于构建质粒的验证、质粒DNA样本质控,及基因工程质粒的分析。在文库构建过程中,质粒 DNA 样本通过快速连接条形码在片段化的同时完成条形码连接,随后将样本混合并纯化。接着将快速测序接头连接至 DNA 末端,最后将制备好的文库上样至芯片进行测序。
我们建议新用户将测序时间定为12个小时,尽管更短时间可能也能获得足够的数据。测序后,您可使用EPI2ME 中的 Clone Validation(克隆验证)工作流程(wf-clone-validation)进行下游数据分析,报告每个质粒的共有序列。后文中将提供有关 MinKNOW 和 EPI2ME 工作流程的详细说明。
测序工作流程:
实验准备
您将需要:
- 提取DNA*,并评估DNA的长度、浓度和纯度。 质量评估步骤对确保实验成功至关重要。
- 确保您已准备好测序试剂盒、正确的仪器以及第三方试剂。
- 下载数据收集和分析软件。
- 检查您的测序芯片上有足够的活性纳米孔,以确保测序良好运行。
请注意: *有关提取方法的建议,请参阅我们的质粒 DNA 提取方法。
文库制备
您将需要:
- 通过快速连接条形码对 DNA 进行片段化,并同时连接一对条形码。
- 合并带条形码的样本
- 将快速测序文库接头连接到 DNA 末端
- 对测序芯片进行预处理,将 DNA 文库加至芯片。

测序和分析
您将需要:
- 使用 MinKNOW 软件运行测序,该软件将收集由测序仪产出的原始数据,并将其识别成碱基序列并进行条码拆分;
- 使用 EPI2ME 软件中的 Clone Validation(克隆验证)工作流程进行分析。
实验方案适用性
本实验方案只适用于与以下产品搭配使用:
- 快速条形码测序试剂盒-24 V14(SQK-RBK114.24)
- 快速条形码测序试剂盒-96 V14(SQK-RBK114.96)
- R10.4.1 测序芯片(FLO-MIN114)
- 测序芯片清洗试剂盒(EXP-WSH004)
- 测序芯片预处理试剂盒 V14 (EXP-FLP004)
- 测序辅助扩展包 V14(EXP-AUX003)
- 快速测序文库接头辅助扩展包 V14 (EXP-RAA114)
- MinION Mk1D - MinION Mk1D IT 配置要求文档
- GridION - GridION IT 配置要求文档
2. 仪器及耗材
材料
- 每个样本 50ng 高分子量质粒DNA
- 快速条形码测序试剂盒-24 V14 (SQK-RBK114.24)或 快速条形码测序试剂盒-96 V14 (SQK-RBK114.96)
耗材
- 1.5 ml Eppendorf DNA LoBind 离心管
- 2 ml Eppendorf DNA LoBind 离心管
- 0.2 ml 薄壁 PCR 离心管 或 0.2 ml 96 孔 PCR 板
- 无核酸酶水(如 Thermo Scientific,AM9937)
- 新制备的 80% 乙醇(用无核酸酶水配制)
- 牛血清白蛋白(BSA)(50 mg/mL)(例如 Invitrogen™ UltraPure™ BSA 50 mg/mL, AM2616)
- Qubit™ 分析管(Invitrogen, Q32856)
- Qubit dsDNA HS Assay(双链DNA高灵敏度检测)试剂盒(Invitrogen, Q32851)
仪器
- MinION 或 GridION 测序仪
- 盛有冰的冰桶
- 微孔板离心机
- 计时器
- 热循环仪或恒温加热仪
- 磁力架
- Hula混匀仪(低速旋转式混匀仪)
- P1000 移液枪和枪头
- P200 移液枪和枪头
- P100 移液枪和枪头
- P20 移液枪和枪头
- P2 移液枪和枪头
- 多通道移液枪和枪头
- Qubit™ 荧光计(或用于质控检测的等效仪器)
可选仪器
- 标准凝胶电泳设备
- Agilent 生物分析仪(或等效仪器)
根据本实验指南,您将需要为每个样本准备 50 ng 高分子量质粒DNA。
起始DNA
DNA质控
选择符合质量和浓度要求的起始DNA至关重要。使用过少或过多的DNA,或者质量较差的DNA(如,高度碎片化、含有RNA或化学污染物的DNA)都会影响文库制备。
有关如何对DNA样品进行质控,请参考起始DNA/RNA质控实验指南。
化学污染物
从原始样本中提取DNA的方法不同,可能会导致经纯化的DNA中所残留的化学污染物不同。这会影响文库的制备效率和测序质量。请参阅牛津纳米孔社区的污染物 页面了解更多信息。
测序芯片质检
我们强烈建议您在开始测序实验前,对测序芯片的活性纳米孔数进行质检。质检需在您收到 MinION/GridION 测序芯片12周内进行。Oxford Nanopore Technologies 会对活性孔数量少于以下标准,且尚未投入测序使用的芯片进行替换**:
| 测序芯片 | 芯片上的活性孔数确保不少于 |
|---|---|
| MinION/GridION 测序芯片 | 800 |
** 请注意:自收到之日起,芯片须一直贮存于 Oxford Nanopore Technologies 推荐的条件下。且质检结果须在质检后的两天内递交给我们。请您按照测序芯片质检文档中的说明进行芯片质检。
本试剂盒及其实验指南中使用的快速测序文库接头(RA)不可与其他测序接头互换使用。
快速条形码测序试剂盒-24 V14 (SQK-RBK114.24)内容物
我们正在将试剂盒中的条形码包装更改为96孔板形式。 这一改动将减少塑料浪费,并支持自动化应用。
孔板装形式
| 名称 | 缩写 | 管盖颜色 | 管数 | 每管溶液体积 (μl) |
|---|---|---|---|---|
| 快速测序文库接头 | RA | 绿色 | 1 | 15 |
| 接头缓冲液 | ADB | 透明 | 1 | 100 |
| AMPure XP 磁珠 | AXP | 琥珀色 | 2 | 1200 |
| 洗脱缓冲液 | EB | 黑色 | 1 | 500 |
| 测序缓冲液 | SB | 红色 | 1 | 700 |
| 文库颗粒 | LIB | 粉色 | 1 | 600 |
| 文库溶液 | LIS | 白色管盖,粉色标签 | 1 | 600 |
| 测序芯片冲洗液 | FCF | 透明管盖,浅蓝色标签 | 1 | 8000 |
| 测序芯片系绳 | FCT | 紫色 | 1 | 200 |
| 快速连接条形码孔板 | RB01-24 | - | 两板,每板三套条形码组合 | 每孔5µl |
本产品包含由贝克曼库尔特公司(Beckman Coulter, Inc)生产的 AMPure XP 试剂,并可与试剂盒一起于 -20°C 下储存(试剂稳定性将不受损害)。
管装形式
| 名称 | 缩写 | 管盖颜色 | 管数 | 溶液体积 (μl) |
|---|---|---|---|---|
| 快速测序文库接头 | RA | 绿色 | 1 | 15 |
| 接头缓冲液 | ADB | 透明 | 1 | 100 |
| AMPure XP 磁珠 | AXP | 琥珀色 | 2 | 1200 |
| 洗脱缓冲液 | EB | 黑色 | 1 | 500 |
| 测序缓冲液 | SB | 红色 | 1 | 700 |
| 文库颗粒 | LIB | 粉色 | 1 | 600 |
| 文库溶液 | LIS | 白色管盖,粉色标签 | 1 | 600 |
| 测序芯片冲洗液 | FCF | 蓝色 | 6 | 1170 |
| 测序芯片系绳 | FCT | 紫色 | 1 | 200 |
| 快速连接条形码 | RB01-24 | 透明 | 24 | 15 |
本产品包含由贝克曼库尔特公司(Beckman Coulter, Inc)生产的 AMPure XP 试剂,并可与试剂盒一起于-20°C 下储存(试剂稳定性将不受损害)。
快速条形码测序试剂盒-96 V14 (SQK-RBK114.96)内容物
| 名称 | 缩写 | 管盖颜色 | 管数 | 溶液体积 (μl) |
|---|---|---|---|---|
| 快速测序文库接头 | RA | 绿色 | 2 | 15 |
| 接头缓冲液 | ADB | 透明 | 1 | 100 |
| AMPure XP 磁珠 | AXP | 琥珀色 | 3 | 1,200 |
| 洗脱缓冲液 | EB | 黑色 | 1 | 1,500 |
| 测序缓冲液 | SB | 红色 | 1 | 1,700 |
| 文库颗粒 | LIB | 粉色 | 1 | 1,800 |
| 文库溶液 | LIS | 白色管盖,粉色标签 | 1 | 1,800 |
| 测序芯片冲洗液 | FCF | 透明 | 1 | 15,500 |
| 测序芯片系绳 | FCT | 紫色 | 2 | 200 |
| 快速连接条形码 | RB01-96 | - | 3 盘 | 每孔 8 µl |
本产品包含由贝克曼库尔特公司(Beckman Coulter, Inc)生产的 AMPure XP 试剂,并可与试剂盒一起于 -20°C 下储存(试剂稳定性将不受损害)。
快速连接条形码序列
| 条码名称 | 序列 |
|---|---|
| RB01 | AAGAAAGTTGTCGGTGTCTTTGTG |
| RB02 | TCGATTCCGTTTGTAGTCGTCTGT |
| RB03 | GAGTCTTGTGTCCCAGTTACCAGG |
| RB04 | TTCGGATTCTATCGTGTTTCCCTA |
| RB05 | CTTGTCCAGGGTTTGTGTAACCTT |
| RB06 | TTCTCGCAAAGGCAGAAAGTAGTC |
| RB07 | GTGTTACCGTGGGAATGAATCCTT |
| RB08 | TTCAGGGAACAAACCAAGTTACGT |
| RB09 | AACTAGGCACAGCGAGTCTTGGTT |
| RB10 | AAGCGTTGAAACCTTTGTCCTCTC |
| RB11 | GTTTCATCTATCGGAGGGAATGGA |
| RB12 | CAGGTAGAAAGAAGCAGAATCGGA |
| RB13 | AGAACGACTTCCATACTCGTGTGA |
| RB14 | AACGAGTCTCTTGGGACCCATAGA |
| RB15 | AGGTCTACCTCGCTAACACCACTG |
| RB16 | CGTCAACTGACAGTGGTTCGTACT |
| RB17 | ACCCTCCAGGAAAGTACCTCTGAT |
| RB18 | CCAAACCCAACAACCTAGATAGGC |
| RB19 | GTTCCTCGTGCAGTGTCAAGAGAT |
| RB20 | TTGCGTCCTGTTACGAGAACTCAT |
| RB21 | GAGCCTCTCATTGTCCGTTCTCTA |
| RB22 | ACCACTGCCATGTATCAAAGTACG |
| RB23 | CTTACTACCCAGTGAACCTCCTCG |
| RB24 | GCATAGTTCTGCATGATGGGTTAG |
| RB25 | GTAAGTTGGGTATGCAACGCAATG |
| RB26 | CATACAGCGACTACGCATTCTCAT |
| RB27 | CGACGGTTAGATTCACCTCTTACA |
| RB28 | TGAAACCTAAGAAGGCACCGTATC |
| RB29 | CTAGACACCTTGGGTTGACAGACC |
| RB30 | TCAGTGAGGATCTACTTCGACCCA |
| RB31 | TGCGTACAGCAATCAGTTACATTG |
| RB32 | CCAGTAGAAGTCCGACAACGTCAT |
| RB33 | CAGACTTGGTACGGTTGGGTAACT |
| RB34 | GGACGAAGAACTCAAGTCAAAGGC |
| RB35 | CTACTTACGAAGCTGAGGGACTGC |
| RB36 | ATGTCCCAGTTAGAGGAGGAAACA |
| RB37 | GCTTGCGATTGATGCTTAGTATCA |
| RB38 | ACCACAGGAGGACGATACAGAGAA |
| RB39 | CCACAGTGTCAACTAGAGCCTCTC |
| RB40 | TAGTTTGGATGACCAAGGATAGCC |
| RB41 | GGAGTTCGTCCAGAGAAGTACACG |
| RB42 | CTACGTGTAAGGCATACCTGCCAG |
| RB43 | CTTTCGTTGTTGACTCGACGGTAG |
| RB44 | AGTAGAAAGGGTTCCTTCCCACTC |
| RB45 | GATCCAACAGAGATGCCTTCAGTG |
| RB46 | GCTGTGTTCCACTTCATTCTCCTG |
| RB47 | GTGCAACTTTCCCACAGGTAGTTC |
| RB48 | CATCTGGAACGTGGTACACCTGTA |
| RB49 | ACTGGTGCAGCTTTGAACATCTAG |
| RB50 | ATGGACTTTGGTAACTTCCTGCGT |
| RB51 | GTTGAATGAGCCTACTGGGTCCTC |
| RB52 | TGAGAGACAAGATTGTTCGTGGAC |
| RB53 | AGATTCAGACCGTCTCATGCAAAG |
| RB54 | CAAGAGCTTTGACTAAGGAGCATG |
| RB55 | TGGAAGATGAGACCCTGATCTACG |
| RB56 | TCACTACTCAACAGGTGGCATGAA |
| RB57 | GCTAGGTCAATCTCCTTCGGAAGT |
| RB58 | CAGGTTACTCCTCCGTGAGTCTGA |
| RB59 | TCAATCAAGAAGGGAAAGCAAGGT |
| RB60 | CATGTTCAACCAAGGCTTCTATGG |
| RB61 | AGAGGGTACTATGTGCCTCAGCAC |
| RB62 | CACCCACACTTACTTCAGGACGTA |
| RB63 | TTCTGAAGTTCCTGGGTCTTGAAC |
| RB64 | GACAGACACCGTTCATCGACTTTC |
| RB65 | TTCTCAGTCTTCCTCCAGACAAGG |
| RB66 | CCGATCCTTGTGGCTTCTAACTTC |
| RB67 | GTTTGTCATACTCGTGTGCTCACC |
| RB68 | GAATCTAAGCAAACACGAAGGTGG |
| RB69 | TACAGTCCGAGCCTCATGTGATCT |
| RB70 | ACCGAGATCCTACGAATGGAGTGT |
| RB71 | CCTGGGAGCATCAGGTAGTAACAG |
| RB72 | TAGCTGACTGTCTTCCATACCGAC |
| RB73 | AAGAAACAGGATGACAGAACCCTC |
| RB74 | TACAAGCATCCCAACACTTCCACT |
| RB75 | GACCATTGTGATGAACCCTGTTGT |
| RB76 | ATGCTTGTTACATCAACCCTGGAC |
| RB77 | CGACCTGTTTCTCAGGGATACAAC |
| RB78 | AACAACCGAACCTTTGAATCAGAA |
| RB79 | TCTCGGAGATAGTTCTCACTGCTG |
| RB80 | CGGATGAACATAGGATAGCGATTC |
| RB81 | CCTCATCTTGTGAAGTTGTTTCGG |
| RB82 | ACGGTATGTCGAGTTCCAGGACTA |
| RB83 | TGGCTTGATCTAGGTAAGGTCGAA |
| RB84 | GTAGTGGACCTAGAACCTGTGCCA |
| RB85 | AACGGAGGAGTTAGTTGGATGATC |
| RB86 | AGGTGATCCCAACAAGCGTAAGTA |
| RB87 | TACATGCTCCTGTTGTTAGGGAGG |
| RB88 | TCTTCTACTACCGATCCGAAGCAG |
| RB89 | ACAGCATCAATGTTTGGCTAGTTG |
| RB90 | GATGTAGAGGGTACGGTTTGAGGC |
| RB91 | GGCTCCATAGGAACTCACGCTACT |
| RB92 | TTGTGAGTGGAAAGATACAGGACC |
| RB93 | AGTTTCCATCACTTCAGACTTGGG |
| RB94 | GATTGTCCTCAAACTGCCACCTAC |
| RB95 | CCTGTCTGGAAGAAGAATGGACTT |
| RB96 | CTGAACGGTCATAGAGTCCACCAT |
3. 文库制备
材料
- 每个样本:50 ng 高分子量质粒 DNA
- 快速连接条形码(RB01-24 或 RB01-96)
- 快速测序文库接头(RA)
- 接头缓冲液(ADB)
- AMPure XP 磁珠(AXP)
- 洗脱缓冲液(EB)
耗材
- 0.2 ml 薄壁 PCR 离心管 或 0.2 ml 96 孔 PCR 板
- 1.5 ml Eppendorf DNA LoBind 离心管
- 2 ml Eppendorf DNA LoBind 离心管
- 无核酸酶水(如 Thermo Scientific,AM9937)
- 新制备的 80% 乙醇(用无核酸酶水配制)
- Qubit™ 分析管(Invitrogen, Q32856)
- Qubit dsDNA HS Assay(双链DNA高灵敏度检测)试剂盒(Invitrogen, Q32851)
仪器
- 盛有冰的冰桶
- 计时器
- 热循环仪
- 微孔板离心机
- 磁力架
- Hula混匀仪(低速旋转式混匀仪)
- P1000 移液枪和枪头
- P200 移液枪和枪头
- P100 移液枪和枪头
- P20 移液枪和枪头
- P10 移液枪和枪头
- P2 移液枪和枪头
- 多通道移液枪和枪头
设定热循环仪的程序:30℃两分钟,后接80℃两分钟。
测序芯片质检
我们强烈建议在开始文库制备之前,对测序芯片的活性纳米孔数量进行质检,以确保其足够支持实验的顺利进行。
详情请参阅 MinKNOW 实验指南中的 测序芯片质检说明。
将下表所列试剂置于室温解冻,经迷你离心机瞬时离心后吹打混匀:
| 试剂 | 1.于室温下解冻 | 2.瞬时离心 | 3.吹打混匀 |
|---|---|---|---|
| 快速连接条形码(RB01-24 或 RB01-96) | 未冻结 | ✓ | ✓ |
| 快速测序文库接头(RA) | 未冻结 | ✓ | ✓ |
| AMPure XP 磁珠(AXP) | ✓ | ✓ | 临使用前,吹打或涡旋混匀 |
| 洗脱缓冲液(EB) | ✓ | ✓ | ✓ |
| 接头缓冲液(ADB) | ✓ | ✓ | 涡旋混匀 |
条形码板孔仅限一次性使用。使用前请确认所选孔密封完好;一旦刺穿或开启,不得再次使用。
使用无核酸酶水稀释质粒 DNA 样本。在添加条形码之前,应将每个样本配制为含约 50 ng 质粒 DNA 的 9μl 溶液。
- 请参照下表,使用无核酸酶水将质粒 DNA 样本稀释至约 50 ng:
| 起始浓度 | DNA 体积 | 无核酸酶水体积 | 总体积 |
|---|---|---|---|
| 100 ng/µl | 2 µl | 34 µl | 36 µl |
| 90 ng/µl | 2 µl | 31 µl | 33 µl |
| 80 ng/µl | 2 µl | 27 µl | 29 µl |
| 70 ng/µl | 3 µl | 35 µl | 38 µl |
| 60 ng/µl | 2 µl | 20 µl | 22 µl |
| 50 ng/µl | 2 µl | 16 µl | 18 µl |
| 40 ng/µl | 5 µl | 31 µl | 36 µl |
| 30 ng/µl | 5 µl | 22 µl | 27 µl |
| 20 ng/µl | 5 µl | 13 µl | 18 µl |
| 10 ng/µl | 10 µl | 8 µl | 18 µl |
| <5.56 ng/µl | 9 µl | 0 µl | 9 µl |
- 吹打混匀稀释后样本,然后瞬时离心。
- 向每支 0.2 ml PCR 管或 96 孔板的孔中加入一个 9 μl 样本。
为计划上样于同一测序芯片的样本选择不同的条形码。一个实验中最多可包含96个样本。
请注意: 每个样本应仅对应一种条形码。
在一支 0.2ml 的薄壁 PCR 管 或 96 孔板中,混合以下试剂:您可使用多通道移液枪转移快速连接条形码。
| 试剂 | 体积 |
|---|---|
| 50 ng 模板 DNA | 9 μl |
| 快速连接条形码(RB01-96,每个样本使用一种条形码) | 1 μl |
| 总体积 | 10 μl |
充分吹打混匀,瞬时离心。
将离心管或96孔板在30℃下孵育两分钟,然后在80℃下孵育两分钟。将离心管或96孔板短暂置于冰上冷却。
将离心管或96孔板瞬时离心,收集管/板底的液体。
将所有带条码样本合并至一支洁净的1.5 或 2ml Eppendorf DNA LoBind 离心管中,记下管内液体总体积。
| . | 每个样本的体积 | 12个样本 | 24个样本 | 48个样本 | 96个样本 |
|---|---|---|---|---|---|
| 总体积 | 10 μl | 120 μl | 240 μl | 480 μl | 960 μl |
涡旋振荡以重悬 AMPure XP 磁珠(AXP)。
向全部合并后的带条码样本中,加入等体积的经重悬的AMPure XP磁珠,轻弹离心管以混匀。
| . | 每个样本的体积 | 12个样本 | 24个样本 | 48个样本 | 96个样本 |
|---|---|---|---|---|---|
| AMPure XP 磁珠体积 | 10 μl | 120 μl | 240 μl | 480 μl | 960 μl |
将离心管置于Hula混匀仪(低速旋转式混匀仪)上室温孵育5分钟。
准备3 ml 新制备的 80% 乙醇(用无核酸酶水配制)。
将样品瞬时离心后置于磁力架上,待磁珠与液相完全分离,且液相澄清无色。保持离心管在磁力架上不动,用移液枪吸去清液。
保持试管在磁力架上不动,以 1.5 ml 新鲜制备的 80% 乙醇洗涤磁珠。小心不要扰动磁珠。用移液枪将乙醇吸走并弃掉。
重复上述步骤。
将离心管瞬时离心后置于磁力架上。用移液枪吸走残留的乙醇。将磁珠在空气中干燥约 30 秒,避免干至表面开裂。
将离心管从磁力架上移开。将磁珠重悬于 15µl 洗脱缓冲液中(EB)。室温下孵育10分钟。
将离心管静置于磁力架上至少1分钟,直到磁珠和液相分离,且洗脱液澄清无色。
将15µl洗脱液转移至一支新的1.5ml Eppendorf DNA LoBind管中。
- 将含有 DNA 文库 的洗脱液转移至新的 1.5 ml Eppendorf DNA LoBind 管中
- 将磁珠丢弃
取1µl洗脱样品,用Qubit定量。
将 11 µl 样品转至干净的1.5 ml Eppendorf DNA LoBind 离心管中。
请注意: 我们建议所转移样本中的 DNA 文库量不超过 800 ng。 如体积不足 11 μl,请使用洗脱缓冲液(EB)补足。
在一支新的 1.5 ml Eppendorf DNA LoBind 管中,按以下比例稀释快速测序文库接头(RA),并吹打混匀:
| 试剂 | 体积 |
|---|---|
| 快速测序文库接头(RA) | 1.5 μl |
| 接头缓冲液(ADB) | 3.5 μl |
| 总体积 | 5 μl |
向带有条形码的 DNA 中加入 1 µl 稀释后的快速测序文库接头(RA)。
轻弹离心管以充分混合,并瞬时离心。
室温下孵育5分钟。
构建好的文库即可用于测序芯片上样。在上样前,请将文库置于冰上保存。
4. MinION 及 GridION 测序芯片的预处理及上样
材料
- 测序芯片冲洗液(FCF)
- 测序芯片系绳(FCT)
- 文库溶液(LIS)
- 文库颗粒(LIB)
- 测序缓冲液(SB)
耗材
- 1.5 ml Eppendorf DNA LoBind 离心管
- MinION/GridION 测序芯片
- 无核酸酶水(如 Thermo Scientific,AM9937)
- 牛血清白蛋白(BSA)(50 mg/mL)(例如 Invitrogen™ UltraPure™ BSA 50 mg/mL, AM2616)
仪器
- MinION 或 GridION 测序仪
- MinION/GridION 测序芯片遮光片
- P1000 移液枪和枪头
- P100 移液枪和枪头
- P20 移液枪和枪头
- P10 移液枪和枪头
请注意:本试剂盒仅兼容 R10.4.1 测序芯片(FLO-MIN114)。
从冰箱中取出测序芯片,在室温下放置 20 分钟,以便在预处理和上样时更清晰地观察到传感器阵列。
测序芯片的预处理及上样
我们建议所有新用户在首次运行测序芯片前,观看视频测序芯片的预处理及上样。
使用文库溶液
对大多数测序实验,我们建议用户使用文库颗粒(LIB)为测序芯片上样。但对于黏稠度较高的文库,借助文库颗粒进行上样可能较为困难,建议使用文库缓冲液(LIS)替代。
于室温下解冻测序缓冲液(SB)、文库颗粒(LIB)或文库溶液(LIS)、测序芯片系绳(FCT)和测序芯片冲洗液(FCF)。完全解冻后,涡旋振荡混匀。然后瞬时离心,置于冰上。
为在 MinION R10.4.1 测序芯片(FLO-MIN114)上获得最优测序表现并提高测序产出,请向测序芯片预处理液中加入终浓度为 0.2 mg/ml 的牛血清白蛋白(BSA)。
注意: 我们不建议使用其他类型的白蛋白(如重组人血清白蛋白)。
在室温条件下,将下表所列试剂加入合适的离心管中,吹打混匀,以制备含 BSA 的测序芯片预处理液:
| 试剂 | 体积(每张芯片) |
|---|---|
| 测序芯片冲洗液 (FCF) | 1170 µl |
| 浓度为 50 mg/mL 的牛血清白蛋白 (BSA) | 5 µl |
| 测序芯片系绳(FCT) | 30 µl |
| 总体积 | 1205 µl |
打开 MinION 或 GridION 测序仪的盖子,将测序芯片插入金属固定夹的下方。用力向下按压预处理孔孔盖处,以确保正确的热、电接触。

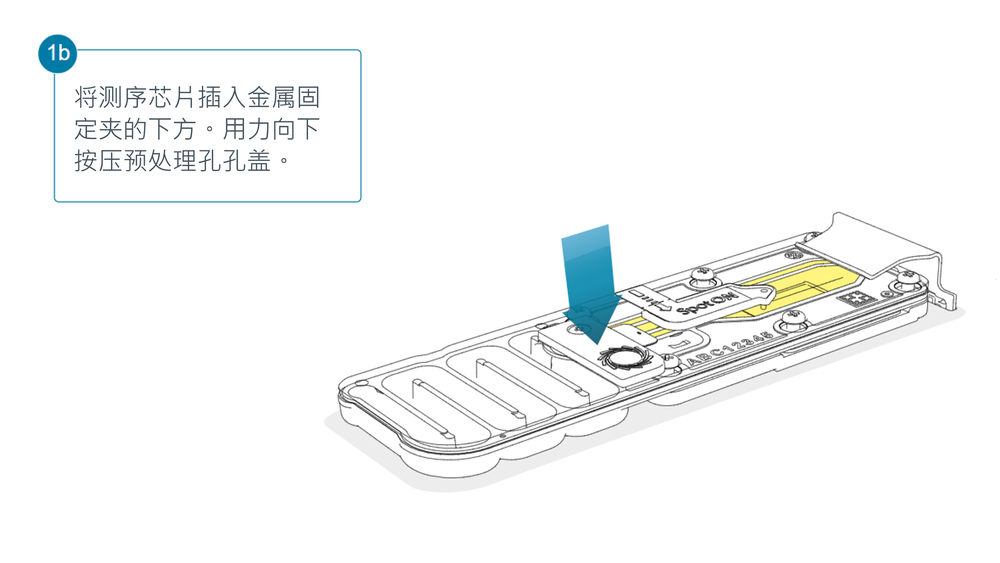
为文库上样前,完成测序芯片质检,查看可用孔数目。
如此前已对测序芯片进行过质检,则此步骤可省略。
详情请参阅 MinKNOW 实验指南中的测序芯片质检说明 。
顺时针转动测序芯片的预处理孔孔盖,使预处理孔显露出来。

小心地从测序芯片中反旋吸出缓冲液。请勿吸出超过 20-30 µl的缓冲液,并确保芯片上的纳米孔阵列一直有缓冲液覆盖。将气泡引入阵列会对纳米孔造成不可逆转地损害。
将预处理孔打开后,检查孔周围是否有小气泡。请按照以下方法,从孔中排出少量液体以清除气泡:
1.将 P1000 移液枪转至 200µl 刻度。
2.将枪头垂直插入预处理孔中。
3.反向转动移液枪量程调节转纽,直至移液枪刻度在 220-230 µl之间,或直至您看到有少量缓冲液进入移液枪枪头。
注意: 肉眼检查,确保从预处理孔到传感器阵列的缓冲液连续且无气泡。

通过预处理孔向芯片中加入 800µl 预处理液,避免引入气泡。等待5分钟。在此期间,请按照以下步骤准备用于上样的文库。

将含有文库颗粒的LIB管用移液枪吹打混匀。
LIB管内的文库颗粒分散于悬浮液中。由于颗粒沉降速度非常快,因此请在混匀颗粒后立即使用。
对于大多数测序实验,我们建议您使用文库颗粒(LIB)。但如文库较为粘稠,您可考虑使用文库溶液(LIS)。
在一支新的1.5ml Eppendorf DNA LoBind离心管内,将所有试剂按以下顺序混合:
| 试剂 | 每张测序芯片的上样体积 |
|---|---|
| 测序缓冲液(SB) | 37.5 µl |
| 文库颗粒 (LIB),临用前混匀;或文库溶液 (LIS) | 25.5 µl |
| DNA 文库 | 12 µl |
| 总体积 | 75 µl |
完成测序芯片的预处理:
- 轻轻地翻起SpotON上样孔盖,使SpotON上样孔显露出来。

- 通过预处理孔(而 非 SpotON加样孔)向芯片中加入200µl预处理液,避免引入气泡。

临上样前,用移液枪轻轻吹打混匀制备好的文库。
通过 SpotON 加样孔向芯片中逐滴加入 75µl 制备好的文库。确保液滴流入孔内后,再加下一滴。

轻轻合上 SpotON 加样孔孔盖,确保塞头塞入加样孔内。合上预处理孔盖。


为获得最佳测序产出,在文库样本上样后,请立即在测序芯片上安装遮光片。
我们建议在清洗芯片并重新上样时,将遮光片保留在测序芯片上。一旦文库从测序芯片中吸出,即可取下遮光片。
按下述步骤安装测序芯片遮光片:
1.小心将遮光片的前沿(平端)与金属固定夹的边沿对齐。 注意: 请勿将遮光片强行压到固定夹下方。
2.将遮光片轻轻盖在测序芯片上。遮光片的 SpotON 加样孔孔盖缺口应与芯片上的 SpotON 加样孔孔盖接合,遮盖住整个测序芯片的前部。

MinION测序芯片的遮光片并非固定在测序芯片上,因此当为芯片加装遮光片后,请小心操作。
合上测序设备上盖,在 MinKNOW 上设置测序实验。
将测序芯片插入 MinION Mk1D 测序仪后,仪器上盖会覆盖于芯片上方,芯片四周可能留有一条小缝隙。此为正常现象,不影响设备性能。
请参阅此 常见问题解答 ,了解有关测序仪上盖的更多信息。

5. 数据采集和碱基识别
如何开始测序
MinKNOW软件负责仪器控制,数据采集和实时碱基识别。请确保已在计算机或设备上安装 MinKNOW。有关测序实验的详细设置说明,请参阅 MinKNOW 实验指南。
我们建议您根据下述碱基识别和条码拆分建议,在MinION或GridION设备上设置测序实验。对于其他选项,您可以保持默认设置。
如需了解更多有关在测序设备上使用 MinKNOW 的信息,请参阅相应的设备用户手册:
通过桌面快捷方式打开 MinKNOW 软件,并使用纳米孔账号登录。
点击已连接的设备。
点击“开始测序”(Start sequencing),为您的实验设定运行参数。
输入实验名称、测序芯片位置及样本ID。在下拉菜单中选择 FLO-MIN114 作为测序芯片类型。
点击 继续至试剂盒选择 (Continue to kit selection)。
选择快速条形码测序试剂盒-24(SQK-RBK114.24)或 快速条形码测序试剂-96(SQK-RBK114.96)。
点击 继续至运行选项 (Continue to Run Options)。
点击“选项”(Options),将运行限制条件(run limit)设为12小时。 其它运行条件可保持默认设置。
点击 继续至碱基识别 (Continue to basecalling)。
根据下列参数配置碱基识别和条码拆分功能:
1.请确保碱基识别功能为开(ON)。
2.点击“模型”(Models)旁的 编辑选项 (Edit options),从下拉菜单中选择高精准碱基识别(HAC)。
3.请确保条码拆分功能为开(ON)。
4.其它选项可保持默认设置。
点击 继续至输出 (Continue to output)。
请根据以下步骤设置输出格式和过滤条件:
1.选择 .POD5 作为输出格式。
2.请确保选择 .FASTQ 作为经碱基识别序列片段的输出格式。
3.请确保过滤功能为开(ON)并启用序列分割(Read splitting)。其它选项可保持默认设置。
点击 继续至参数确认 (Continue to final review)。
点击“开始”(Start)启动测序。
系统将自动跳转至“测序概览”(Sequencing Overview)页面,用于实时监控测序运行状态。
6. 使用 EPI2ME 进行下游分析
下游分析
我们建议使用 EPI2ME Desktop 应用程序 进行下游分析。EPI2ME Desktop 为一款桌面软件,可运行 Nextflow 工作流程,帮助用户高效开展生物信息学分析。EPI2ME 的生物信息学分析流程由长读长测序领域的专家精心开发维护。
点击相应链接,获取有关 EPI2ME 工作流程的更多信息和快速入门指南。
我们推荐您使用 wf-clone-validation workflow 组装小质粒序列。在运行此工作流程之前,请确保已安装 Nextflow和Docker。
通过桌面快捷方式打开 EPI2ME 应用程序,并在侧边栏中点击“Launch(启动)”按钮。
在“Workflows(工作流)”中选择 Clone Validation,进行下载并安装。
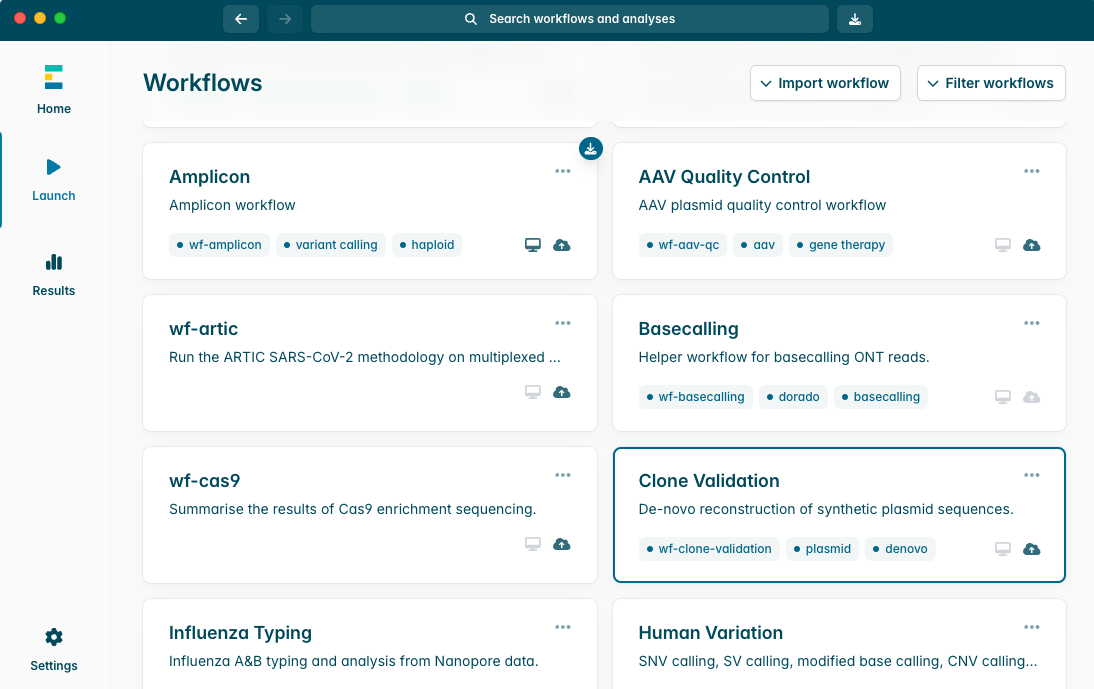
安装 Clone Validation 工作流程。
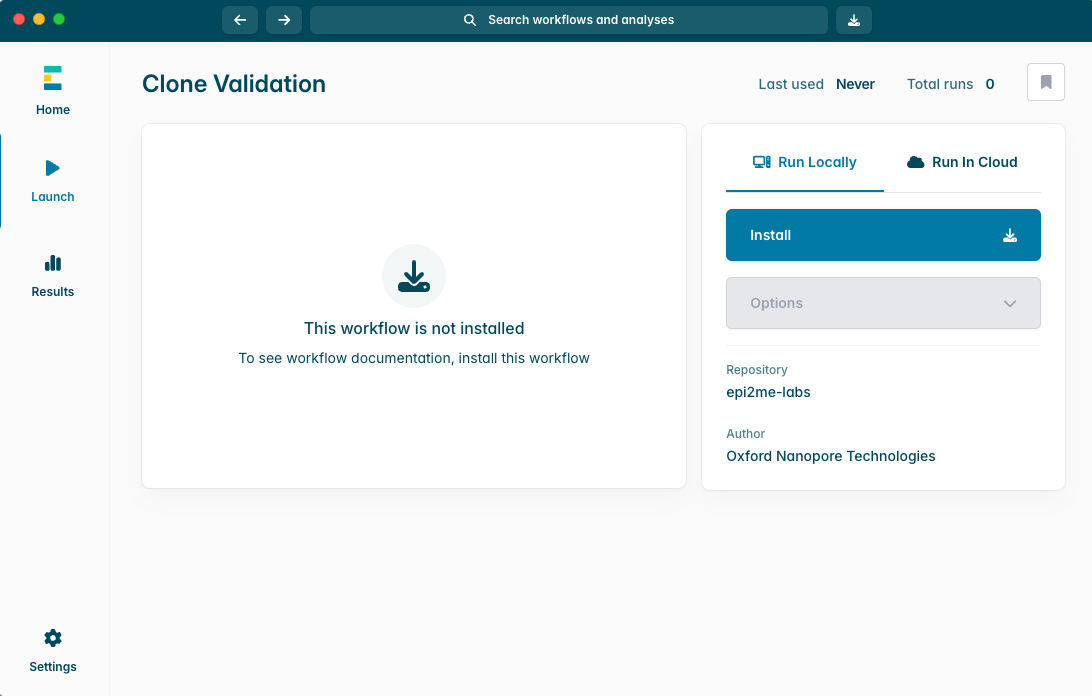
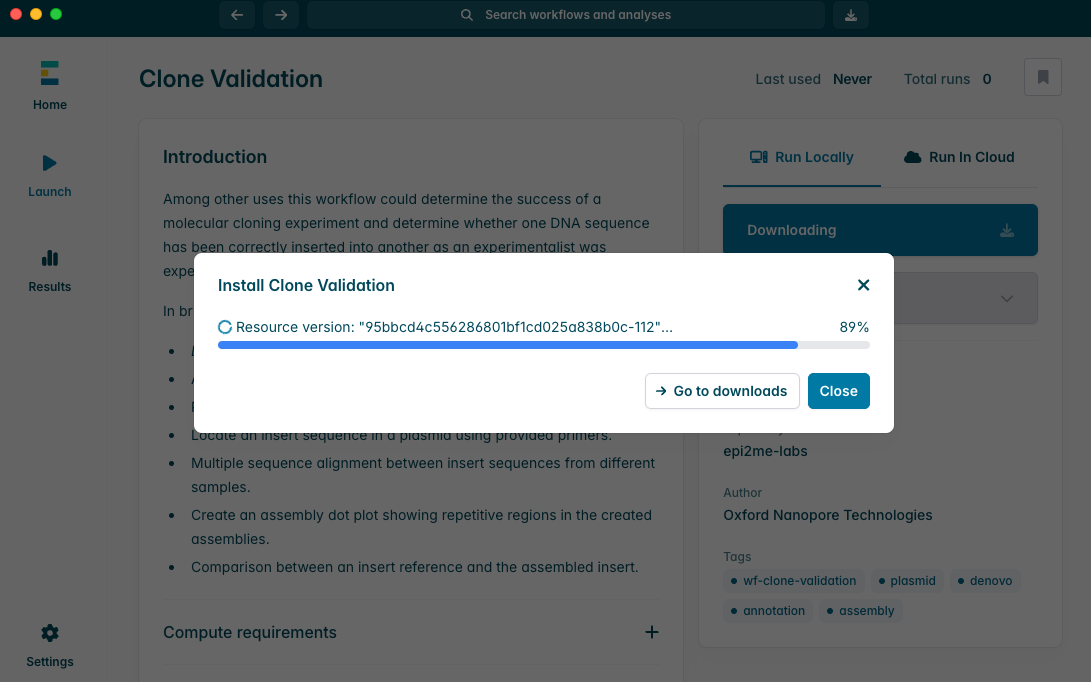
当 Clone Validation 工作流安装完成后,点击“Launch(启动)”按钮以启动分析流程。
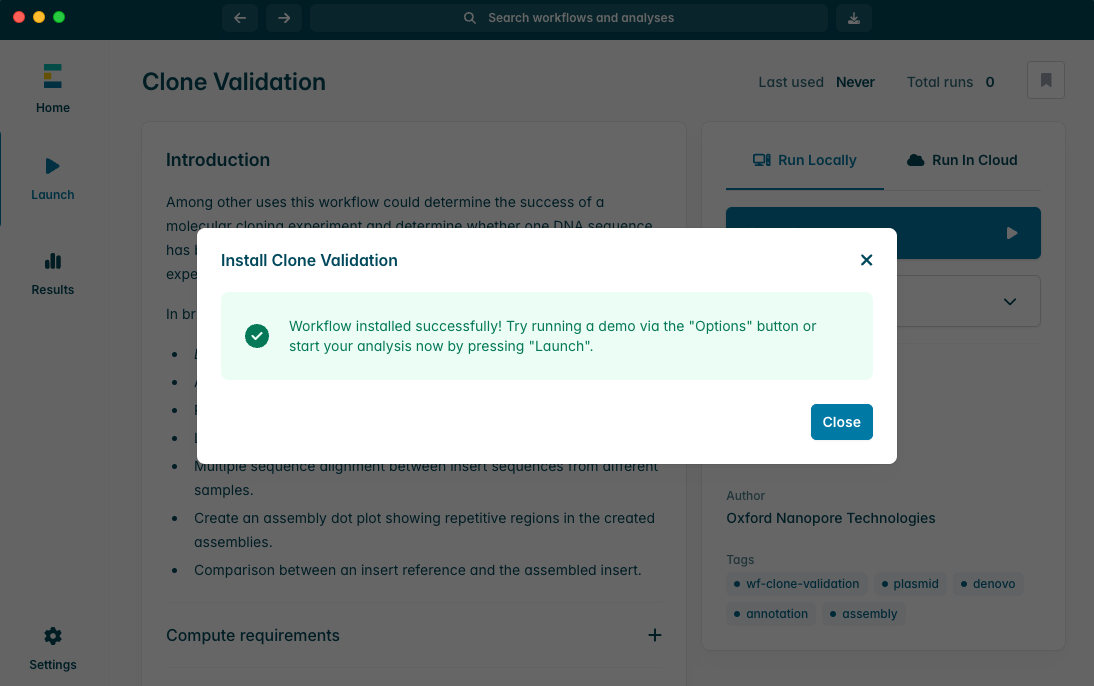
在“Input Options”(输入选项)下上传 FASTQ 文件。请根据需要填写其余参数选项。
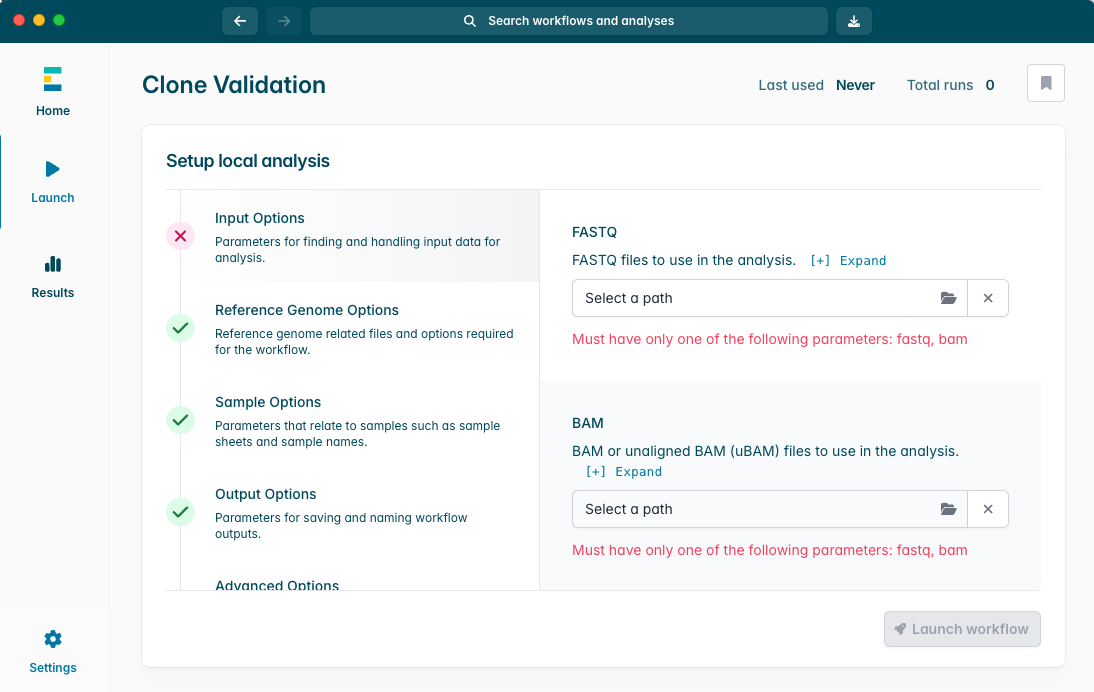
点击“Launch workflow”(启动工作流程)。
确保所有参数选项前均显示绿色对勾。
工作流程运行结束后,将生成报告。
克隆验证工作流程报告
工作流程报告将提供关于质粒序列组装结果的信息,主要包括:
- .fasta文件:显示每个样本的共有序列
- .csv文件:显示每个样本是否通过质控
- 特征列表:包含每个样本的注释信息
- HTML报告文档:详细说明工作流程主要结果
您可在此处查看示例报告。
 报告中的摘要图显示了每种条形码对应的序列数量,并在下方表格中列出了每种条形码对应的共有序列长度。上述数据有助于识别因序列片段数量不足而被排除在序列分析之外的样本。
报告中的摘要图显示了每种条形码对应的序列数量,并在下方表格中列出了每种条形码对应的共有序列长度。上述数据有助于识别因序列片段数量不足而被排除在序列分析之外的样本。
 报告根据条码类型给出读长统计数据和 pLannotate 图,展示了经过校正的质粒共有序列信息。上图示例报告中提供了'barcode01'样本DNA序列的组装结果:共有序列的长度为5385 bp,未填充的区域表示该部分序列信息不完整。
报告根据条码类型给出读长统计数据和 pLannotate 图,展示了经过校正的质粒共有序列信息。上图示例报告中提供了'barcode01'样本DNA序列的组装结果:共有序列的长度为5385 bp,未填充的区域表示该部分序列信息不完整。
 报告还为每种条形码提供了一张特征列表,用于描述注释序列,以精准定位注释特征。
报告还为每种条形码提供了一张特征列表,用于描述注释序列,以精准定位注释特征。
7. 测序芯片的再利用及返还
材料
- 测序芯片清洗剂盒(EXP-WSH004)
完成测序实验后,如您希望再次使用测序芯片,请按照“测序芯片清洗试剂盒实验指南”进行操作,并将清洗后的芯片置于 +2 至 +8℃ 保存。
您可在纳米孔社区获取测序芯片清洗试剂盒实验指南。
或者,请按照回收程序将测序芯片返还至Oxford Nanopore。
您可在 此处找到回收测序芯片的说明。
如果您遇到问题或对测序实验有疑问,请参阅本实验指南“疑难解答指南”一节。
8. DNA 提取和文库制备过程中可能出现的问题
以下表格列出了提取和文库制备过程中的常见问题,以及可能的原因和解决方法。
我们还在 Nanopore 社区的Support板块提供了常见问题解答(FAQ)。
如果以下方案仍无法解决您的问题,请通过电邮(support@nanoporetech.com)或 纳米孔社区的在线支持(LiveChat)联系我们。
低质量样本
经AMPure磁珠纯化后的DNA回收率低
| 现象 | 可能原因 | 措施及备注 |
|---|---|---|
| 低回收率 | AMPure磁珠量与样品量的比例低于预期,导致DNA因未被捕获而丢失 | 1.AMPure 磁珠沉降速度较快,因此在将磁珠加入样品前,请务必充分重悬混匀。 2.当AMPure磁珠量与样品量的比值低于0.4:1时,所有的DNA片段都会在纯化过程中丢失。 |
| 低回收率 | DNA片段短于预期 | AMPure磁珠量与样品量的比值越低,针对短片段的筛选就越严格。每次实验时,请先使用琼脂糖凝胶(或其他凝胶电泳方法)确定起始DNA的长度,据此计算出合适的AMPure磁珠用量。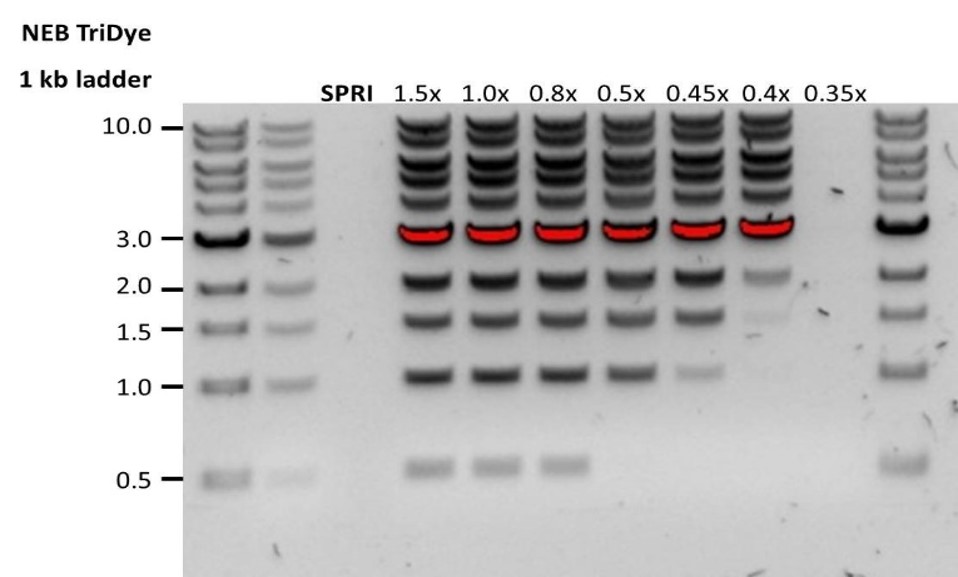 |
| 末端修复后的DNA回收率低 | 清洗步骤所用乙醇的浓度低于70% | 当乙醇浓度低于70%时,DNA会从磁珠上洗脱下来。请确保使用正确浓度的乙醇。 |
9. 测序过程中可能出现的问题
以下表格列出了提取和文库制备过程中的常见问题,以及可能的原因和解决方法。
我们还在 Nanopore 社区的Support板块提供了常见问题解答(FAQ)。
如果以下方案仍无法解决您的问题,请通过电邮(support@nanoporetech.com)或 纳米孔社区的在线支持(LiveChat)联系我们。
MinKNOW Mux 扫描在测序起始时报告的活性孔数少于芯片质检时报告的活性孔数
| 现象 | 可能原因 | 措施及备注 |
|---|---|---|
| MinKNOW Mux 扫描在测序起始时报告的活性孔数少于芯片质检时报告的活性孔数 | 纳米孔阵列中引入了气泡 | 在对通过质控的芯片进行预处理之前,请务必排出预处理孔附近的气泡。否则,气泡会进入纳米孔阵列对其造成不可逆转地损害。视频中演示了避免引入气泡的最佳操作方法。 |
| MinKNOW Mux 扫描在测序起始时报告的活性孔数少于芯片质检时报告的活性孔数 | 测序芯片没有正确插入测序仪 | 停止测序,将芯片从测序仪中取出,再重新插入测序仪内。请确保测序芯片牢固嵌入测序仪中,并已到达目标温度。如用户使用的是GridION/PromethION测序仪,也可尝试将芯片插入仪器的其它芯片槽进行测序。 |
| MinKNOW Mux 扫描在测序起始时报告的活性孔数少于芯片质检时报告的活性孔数 | 文库中残留的污染物对纳米孔造成损害或堵塞 | 在测序芯片质检阶段,我们用芯片储存缓冲液中的质控DNA分子来评估活性纳米孔的数量。而在测序开始时,我们使用DNA文库本身来评估活性纳米孔的数量。因此,活性纳米孔的数量在这两次评估中会有约10%的浮动。如测序开始时报告的孔数明显降低,则可能是由于文库中的污染物对膜结构造成了损坏或将纳米孔堵塞。用户可能需要使用其它的DNA/RNA提取或纯化方法,以提高起始核酸的纯度。您可在 污染物专题技术文档中查看污染物对测序实验的影响。请尝试其它不会导致污染物残留的 提取方法。 |
MinKNOW脚本失败
| 现象 | 可能原因 | 措施及备注 |
|---|---|---|
| MinKNOW显示 "Script failed”(脚本失败) | 重启计算机及MinKNOW。如问题仍未得到解决,请收集 MinKNOW日志文件并联系我们的技术支持。如您没有其他可用的测序设备,我们建议您先将装有文库的测序芯片置于4°C 储存,并联系我们的技术支持团队获取进一步储存上的建议。 |
纳米孔利用率低于40%
| 现象 | 可能原因 | 措施及备注 |
|---|---|---|
| 纳米孔利用率<40% | 测序芯片中的文库量不够 | 请确保您按照相应实验指南,向测序芯片中加入正确浓度和体积的测序文库。请在上样前对文库进行定量,并使用 Promega Biomath Calculator 等工具中的“dsDNA:µg to pmol”功能来计算DNA分子的摩尔量。 |
| 纳米孔利用率接近0 | 使用连接测序试剂盒,但接头并未与DNA成功连接 | 请确保您在“测序接头连接”步骤中使用的是NEBNext快速连接模块(E6056),以及SQK-LSK114试剂盒中的连接缓冲液(LNB)。同时,请确保每种试剂的用量正确。您可通过制备Lambda对照文库来检验第三方试剂的可用性。 |
| 纳米孔利用率接近0 | 使用连接测序试剂盒;但在接头连接后的纯化步骤中并未使用LFB 或SFB洗涤,而是使用了酒精 | 酒精可导致测序接头上的马达蛋白变性。请确保在测序接头连接后使用洗涤缓冲液(LFB或SFB)。 |
| 纳米孔利用率接近0 | 测序芯片中无系绳 | 系绳(FLT或FCT)随预处理液加至芯片。请确保在制备预处理液时,根据需求将 FLT 或 FCT 添加到相应的冲洗缓冲液 (FB) 或 测序芯片冲洗液 (FCF) 中。 |
读长短于预期
| 现象 | 可能原因 | 措施及备注 |
|---|---|---|
| 读长短于预期 | DNA样本降解 | 读长反映了起始DNA片段的长度。起始DNA在提取和文库制备过程中均有可能被打断。 1.请查阅纳米孔社区中的 提取方法 以获得最佳DNA提取方案. 2.在进行文库制备之前,请先跑电泳,查看起始DNA片段的长度分布。 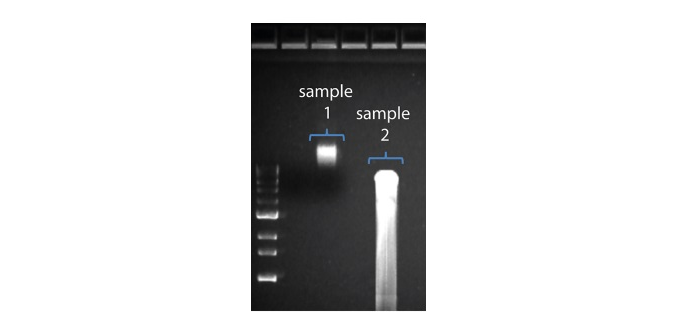 在上图中,样本1为高分子量DNA,而样本2为降解样本。 在上图中,样本1为高分子量DNA,而样本2为降解样本。3.在制备文库的过程中,请避免使用吹打或/和涡旋振荡的方式来混合试剂。轻弹或上下颠倒离心管即可。 |
大量纳米孔处于不可用状态
| 现象 | 可能原因 | 措施及备注 |
|---|---|---|
大量纳米孔处于不可用状态 (在通道面板和纳米孔活动状态图上以蓝色表示) 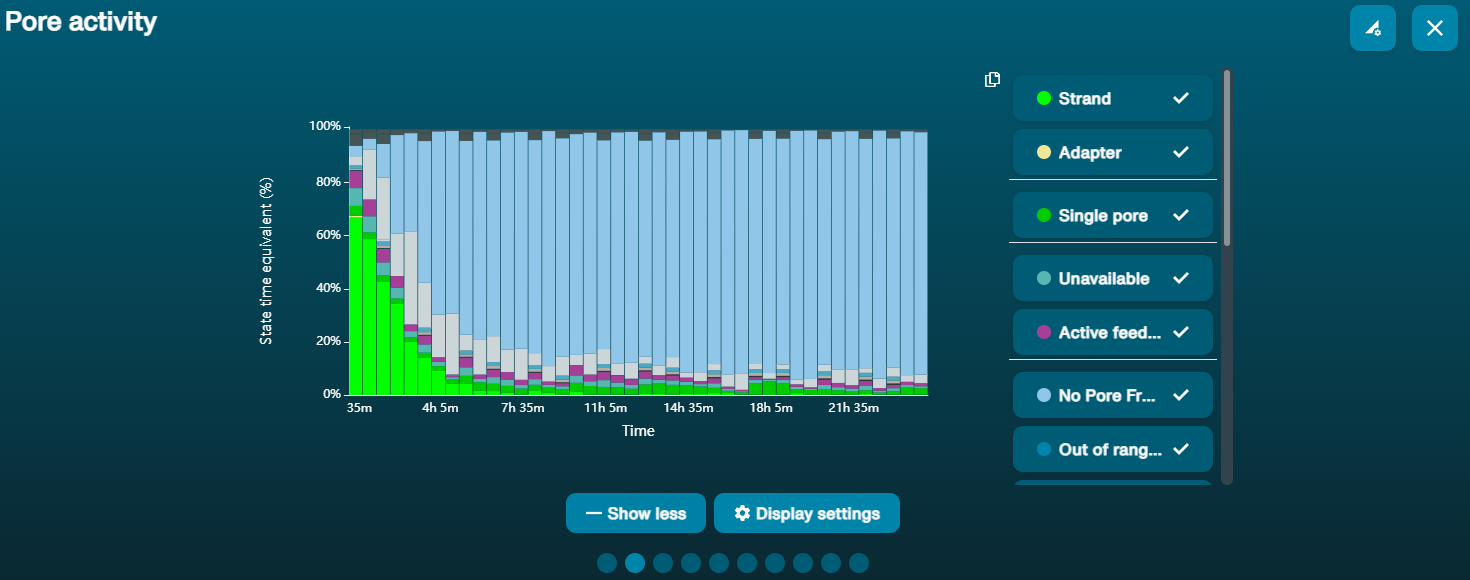 上方的纳米孔活动状态图显示:状态为不可用的纳米孔的比例随着测序进程而不断增加。 上方的纳米孔活动状态图显示:状态为不可用的纳米孔的比例随着测序进程而不断增加。 | 样本中含有污染物 | 使用MinKNOW中的“Unblocking”(疏通)功能,可对一些污染物进行清除。如疏通成功,纳米孔的状态会变为"测序孔"(sequencing pore)。若疏通后,状态为不可用的纳米孔的比例仍然很高甚至增加: 1.用户可使用 测序芯片冲洗试剂盒 (EXP-WSH004)进行核酸酶冲洗 操作,或 2.使用PCR扩增目标片段,以稀释可能导致问题的污染物。 |
大量纳米孔处于“失活”(Inactive)状态
| 现象 | 可能原因 | 措施及备注 |
|---|---|---|
| 大量纳米孔处于失活状态(在通道面板和纳米孔活动状态图上以浅蓝色表示。膜结构或纳米孔遭受不可逆转地损伤 | 测序芯片中引入了气泡 | 芯片预处理和文库上样过程中引入的气泡会对纳米孔带来不可逆转地损害。请观看 测序芯片的预处理及上样 视频了解最佳操作方法 |
| 大量纳米孔处于失活/不可用状态 | 存在与DNA共纯化的化合物 | 已知的化合物包括多糖等。 使用QIAGEN PowerClean Pro试剂盒进行纯化。 |
| 大量纳米孔处于失活/不可用状态 | 样本中含有污染物 | 您可在Contaminants 中查看污染物对测序实验的影响。请尝试其它不会导致污染物残留的提取方法。 |
温度波动
| 现象 | 可能原因 | 措施及备注 |
|---|---|---|
| 温度波动 | 测序芯片和仪器接触不良 | 检查芯片背面的金属板是否有热垫覆盖。重新插入测序芯片,用力向下按压,以确保芯片的连接器引脚与测序仪牢固接触。如问题仍未得到解决,请联系我们的技术支持。 |
未能达到目标温度
| 现象 | 可能原因 | 措施及备注 |
|---|---|---|
| MinKNOW显示“未能达到目标温度” | 测序仪所处环境低于标准室温,或通风不良(以致芯片过热) | MinKNOW会限定测序芯片达到目标温度的时间。当超过限定时间后,系统会显示出错信息,但测序实验仍会继续。值得注意的是,在错误温度下测序可能会导致通量和数据质量(Q值)的降低。请调整测序仪的摆放位置,确保将其置于室温下、通风良好的环境中,再在MinKNOW中重启进程。有关 MinION 温度控制的更多信息,请点击 此链接查看。 |







