Plant research
Resolve complex plant genomes
Offering reads of unrestricted length, from short to ultra long, Oxford Nanopore technology enables accurate assembly of large, highly repetitive plant genomes — resolving structural variants (SVs), transposons, and transgene insertions — to deliver new insights into plant biology, evolution, and breeding strategies for biodiversity studies and agrigenomics.
Epigenetic modifications (e.g. methylation) can be identified alongside nucleotide sequences — in one go — with no additional protocol steps or expense, while the facility to sequence full-length transcripts supports enhanced gene annotation and gene expression studies.
Featured content
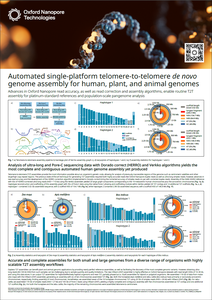
Advances in Oxford Nanopore read accuracy
This poster describes how to achieve accurate and complete assemblies for genomes of any size across a wide variety of organisms, including maize (Zea mays), using highly scalable telomere-to-telomere (T2T) assembly workflows.

Assembling high-quality plant genomes
Oxford Nanopore technology can easily sequence large and complex plant genomes due to nanopore reads of unrestricted length. This end-to-end workflow introduces how to generate a high-quality plant genome assembly from a leaf sample.
Recommended device for plant research

PromethION 24
The flexible, high-throughput PromethION 24 delivers complete genomic and transcriptomic characterisation of large numbers of plant lines.
Generate high sequencing outputs on up to 24 independently addressable PromethION Flow Cells and utilise powerful onboard compute.









