Is the end to the diagnostic odyssey within reach?
- Published on: August 27 2025
For millions of families with rare diseases, the search for answers begins with uncertainty. Rare diseases affect an estimated 300 million people worldwide, with symptoms appearing in childhood for almost 70% of those affected1. Most rare diseases have a genetic cause — yet despite advancements in genomic sequencing, patients typically wait an average of five to eight years1,2 for a correct diagnosis, with more than half remaining undiagnosed3.
Sinha, Rabea, and Ramaswamy et al.3 set out to address this challenge by exploring whether Oxford Nanopore sequencing, combined with a simplified funnel-down filtration strategy, could enhance the detection of pathogenic variants in rare diseases.
‘Long-read sequencing (LRS) technologies could serve as a unified platform for clinical genetic testing, particularly in rare disease settings’
Sinha, Rabea, and Ramaswamy et al.3
Expanding detection capabilities of genomic sequencing
Current diagnostic methods urgently need improvement — up to 30% of children with a rare disease die before their fifth birthday1. Additionally, many patients remain on a diagnostic odyssey for years, without answers or specific treatment plans, placing a significant burden on families and healthcare systems3. Some never get an answer.
Diagnosis typically relies on multiple costly genomic tests, which utilise a diverse number of methods, including short-read sequencing, which limits the ability to capture complex variants, such as structural variants (SVs) and methylation profiles3. In contrast, the unrestricted read lengths offered by Oxford Nanopore sequencing enable all these variants to be captured in a single run. Plus, methylation data is available alongside genomic data, without additional library prep steps, such as bisulfite treatment or enzymatic conversion.
This led Sinha, Rabea, and Ramaswamy et al. to investigate whether nanopore sequencing could identify additional known pathogenic variants in research samples from previously tested but undiagnosed patients.

The team performed whole-genome sequencing (WGS) using whole blood samples prepared with the Ligation Sequencing Kit on a PromethION. They achieved a minimum coverage of 30x with an average N50 of 12 kb . Data was analysed using the EPI2ME wf-human-variation4 bioinformatics pipeline alongside their in-house long-read data pipeline, resulting in the detection of thousands of variants. To focus on clinically relevant findings, the research team employed a filtration strategy to retain only known SVs and copy number variants (CNVs) associated with disease, as determined by the Online Mendelian Inheritance in Man (OMIM)5 and the Gene Curation Coalition (GenCC)6 databases. Methylation data were also captured, and epigenetic signatures were compared to 36 Mendelian neurodevelopmental disorders' methylation signatures7.
The filtration strategy was first optimised on control DNA samples with known genomic and epigenomic pathogenic variants before being used to analyse research samples from 51 undiagnosed patients who had all previously received inconclusive results from short-read whole-exome sequencing (WES).
Unravelling genetic variants in undiagnosed rare diseases
In one case, the Oxford Nanopore workflow successfully identified a pathogenic CNV deletion event at 2q11.1–q11.2, which was validated by chromosomal microarray (CMA) testing, aligning with the patient phenotype. A second 1.4 Mb CNV was also identified by the nanopore workflow and validated with CMA testing, but was not considered causative, as the only known disease-associated gene in the region, NSMCE3, is linked to a different clinical presentation.
The workflow was also successfully used to analyse a sample where WES previously suggested anterior segment dysgenesis (ASD) due to a heterozygous pathogenic indel in SLC38A8. However, ASD is an autosomal recessive disease, meaning both alleles must be affected to confirm the diagnosis. Oxford Nanopore sequencing detected a second 80 kb heterozygous pathogenic deletion in the same gene, previously missed by WES. Additionally, the team harnessed unrestricted nanopore read lengths to phase the two variants, revealing them to be on distinct haplotypes, confirming the compound heterozygosity and biallelic impairment of SLC38A8. The patient’s phenotype matched other patients with ASD, strongly supporting these findings.
Investigating the epigenetic landscape
The researchers further used Oxford Nanopore sequencing to investigate the methylome generated alongside the WGS data, circumventing laborious wet lab steps required with legacy sequencing methods. During the optimisation of their analysis workflow — ‘Epimarker’ — they observed the methylation profile across the SMN1 gene, where biallelic loss causes spinal muscular atrophy (SMA). SMA is a group of hereditary diseases that impact motor neuron function, with a wide range of severities and symptoms9.
The team explored whether the methylation profile of the SMN1 gene could be used as a marker for the disease. They observed distinct levels of methylation: SMA patients had low levels (<15%), carriers had moderate levels (50–70%), and non-carriers had high levels (98–100%). Further validation using deconvoluted nanopore reads revealed that SMA patients had minimal reads mapping to SMN1, as confirmed by droplet digital PCR (Figure 1). Based on these findings, the researchers propose that Oxford Nanopore sequencing has the potential to diagnose patients with SMA and to identify carriers by detecting reduced read coverage across SMN1.
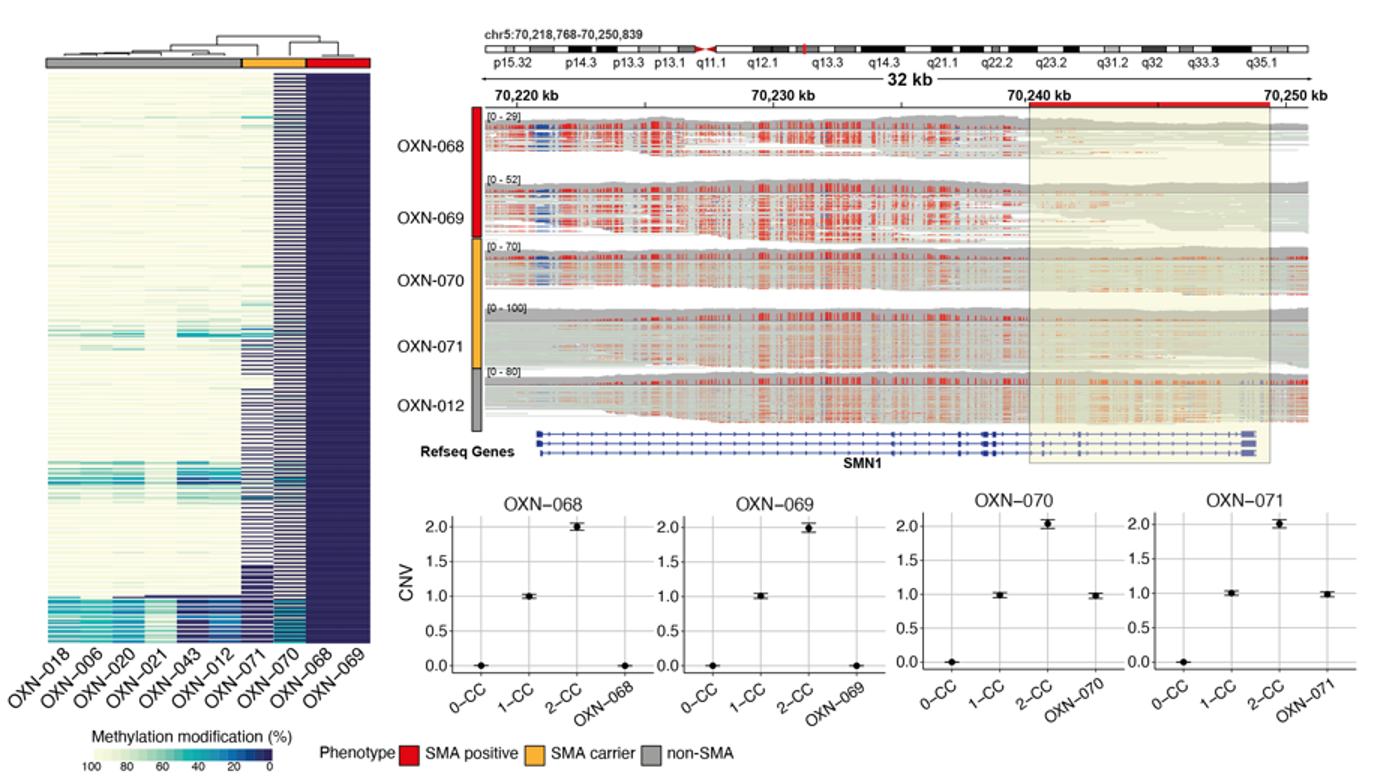
Figure 1. The SMA episignature from the preliminary results. The left panel is the methylation heatmap across positive, carrier, and negative samples. The top-right panel is the IGV methylation of SMA-positive, SMA carriers, and SMA-negative samples. The bottom-right panel shows the droplet digital PCR results with CNVs. Figure from Tayoun et al.10 and available under Creative Commons license (https://creativecommons.org/licenses/by/4.0).
‘… our pipeline was able to correctly identify all the pathogenic variants, including complex rearrangements and aberrant methylation, in the optimisation cohort’
Sinha, Rabea, and Ramaswamy et al.3
Using the WGS data, methylation patterns were also investigated across 51 undiagnosed patient research samples. In one sample, the methylation profile showed hypomethylation across NSD1, which is associated with Hunter McAlpine syndrome (HMA). To confirm this, the team investigated the whole-genome nanopore data and found a duplication in the NSD1 gene. These findings and a subsequent diagnosis of HMA were confirmed by CMA testing.
Ending the diagnostic odyssey with Oxford Nanopore sequencing
Sinha, Rabea, and Ramaswamy et al. demonstrate that native whole-genome Oxford Nanopore sequencing generates comprehensive data to access the genome and epigenome to investigate rare diseases. With access to both genomic and epigenomic layers, a vast array of variants can be identified on a single platform, potentially reducing the number of assays required for accurate disease identification. In the future, this approach could potentially help end the diagnostic odyssey currently faced by patients with rare diseases — providing faster and more comprehensive answers than current methods.
Using this single-platform sequencing workflow alongside a considered variant filtration strategy, the research team were able to focus on known pathogenic variants across a wide genetic spectrum, pinpointing potential genomic causes of rare disease in five currently undiagnosed patients.
Find out how other researchers are using nanopore sequencing to resolve rare diseases in the blog: Reshaping rare disease diagnosis — an end to the diagnostic odyssey?
Oxford Nanopore Technologies products are not intended for use for health assessment or to diagnose, treat, mitigate, cure, or prevent any disease or condition.
- The Lancet Global Health. The landscape for rare diseases in 2024. The Lancet Global Health. 12(3):e341 (2024). DOI: https://doi.org/10.1016/S2214-109X(24)00056-1
- Faye, F. et al. Time to diagnosis and determinants of diagnostic delays of people living with a rare disease: results of a rare barometer retrospective patient survey. Eur. J. Hum. Genet. 32(9):1116–1126 (2024). DOI: https://doi.org/10.1038/s41431-024-01604-z
- Sinha, S., Rabea, F., and Ramaswamy, S. et al. Long-read sequencing enhances pathogenic and novel variation discovery in patients with rare diseases. Nat. Commun. 16(1):2500 (2025). DOI: https://doi.org/10.1038/s41467-025-57695-9
- GitHub. wf-human-variation. Available at: https://github.com/epi2me-labs/wf-human-variation [Accessed 19 August 2025]
- Online Mendelian Inheritance in Man (OMIM). An online catalogue of human genes and genetic disorders. https://www.omim.org/ (2025) [Accessed 19 August 2025]
- The Gene Curation Coalition. A global effort to harmonise gene-level resources. https://thegencc.org/ (2025) [Accessed 19 August 2025]
- Aref-Eshghi, E. et al. Evaluation of DNA methylation episignatures for diagnosis and phenotype correlations in 42 Mendelian neurodevelopmental disorders. Am. J. Hum. Genet. 106(3):356–370 (2020). DOI: https://doi.org/10.1016/j.ajhg.2020.01.019
- Kuht, H.J., Ha, J., and Maconachie, G.D.E. et al. SLC38A8 mutations result in arrested retinal development with loss of cone photoreceptor specialisation. Hum. Mol. Genet. 29(18):2989–3002 (2020). DOI: https://doi.org/10.1093/hmg/ddaa166
- National Institute of Neurological Disorders and Stroke. Spinal muscular atrophy. https://www.ninds.nih.gov/health-information/disorders/spinal-muscular-atrophy (2024) [Accessed 10 July 2025]
- Tayoun, A.A. et al. Long read sequencing enhances pathogenic and novel variation discovery in patients with rare diseases. Research Square 1 (2024). DOI: https://doi.org/10.21203/rs.3.rs-4235049/v1







