Anthropogenic impact on environmental microbiome using nanopore metagenomics
- Published on: May 19 2023
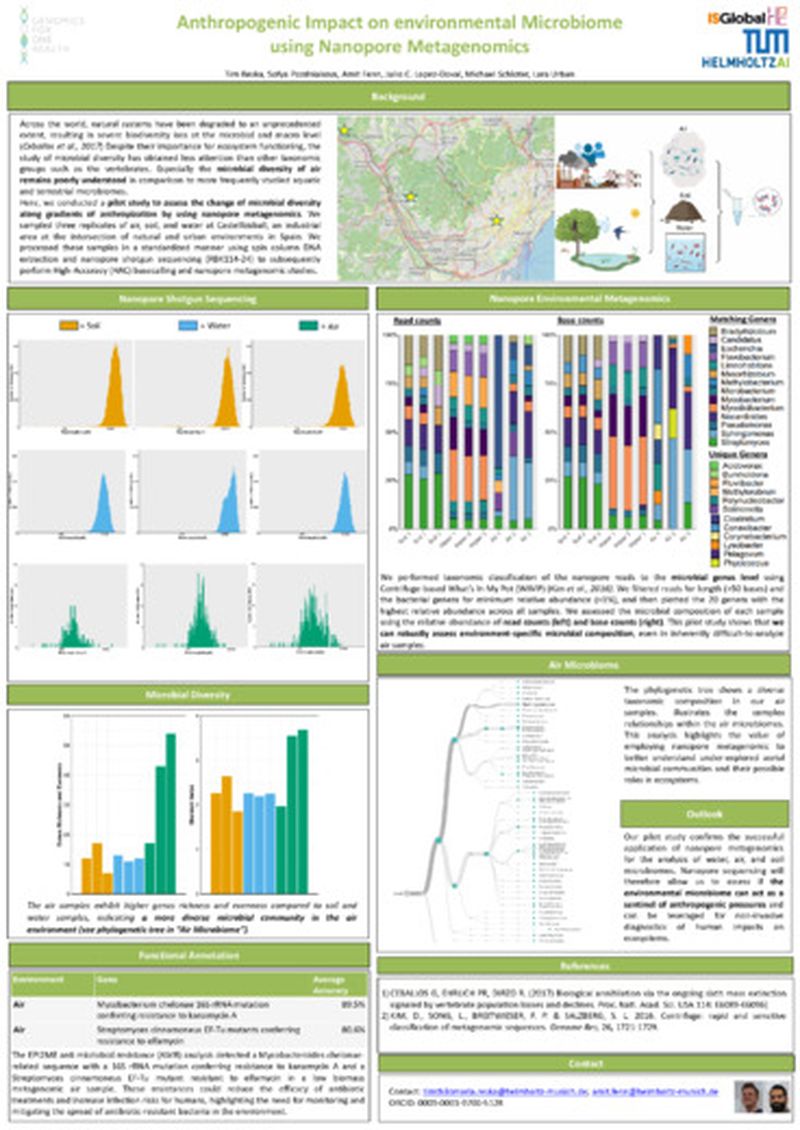
Across the world, natural systems have been degraded to an unprecedented extent, resulting in severe biodiversity loss at the microbial and macro level (Ceballos et al., 2017). Despite their importance for ecosystem functioning, the study of microbial diversity has obtained less attention than other taxonomic groups such as the vertebrates. Especially the microbial diversity of air remains poorly understood in comparison to more frequently studied aquatic and terrestrial microbiomes.
Here, we conducted a pilot study to assess the change of microbial diversity along gradients of anthropization by using nanopore metagenomics. We sampled three replicates of air, soil, and water at Castellbisball, an industrial area at the intersection of natural and urban environments in Spain. We processed these samples in a standardized manner using spin column DNA extraction and nanopore shotgun sequencing (RBK 114-24) to subsequently perform High Accuracy (HAC) basecalling and nanopore metagenomic studies.






