Analyse
Data analysis: for all levels of expertise
Oxford Nanopore provides a wide range of tools to support Oxford Nanopore data analysis. From preconfigured analysis workflows in EPI2ME to the latest cutting-edge tools to make the most of your sequencing runs.

Preconfigured analysis workflows with EPI2ME
- Suitable for all experience levels via an intuitive desktop app or command line
- Utilise preconfigured workflows for a wide range of applications
- Analyse your data anywhere, whether on a laptop, cluster, cloud environment, or Oxford Nanopore device
- Access results immediately with real-time analysis workflows
- Build custom workflows and share via the EPI2ME application
For scientists

Data analysis for everyone
- Intuitive interface, seamless analysis experience
- Comprehensive reports for rapid insights
- No bioinformatics experience required
- Keep ownership of your data from sample to answer
For bioinformaticians

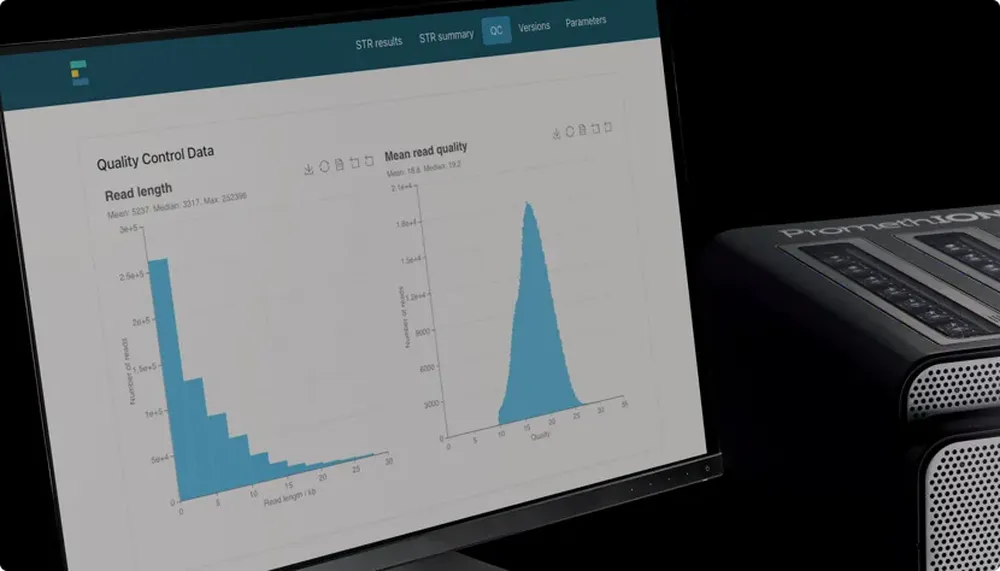
Cutting-edge analytical tools
- Find the latest basecallers and cutting-edge tools in the Oxford Nanopore GitHub repository
- Access EPI2ME workflows on GitHub or integrate your own code
- Command-line experience required
Intuitive analysis with EPI2ME
A full suite of data analysis workflows for a wide range of applications.

Datasets
Oxford Nanopore Technologies Open Data provides researchers open access to nanopore sequencing datasets from Oxford Nanopore sequencing hosted by AWS.
Compatible interpretation solutions
We are working with leading tertiary analysis partners to provide end-to-end solutions for a range of sequencing applications. Find data analysis compatible products here

Online training
Get started with Oxford Nanopore sequencing data analysis. Learn how in this materclass.
MinKNOW: intuitive software native to all platforms
MinKNOW is the operating system for all of our devices. It performs raw data acquisition and basecalling, with built-in methylation detection. MinKNOW further enables in silico targeted sequencing using adaptive sampling and can be connected to upstream and downstream tools using MinKNOW API.