Solving the parent-of-origin effect in retinoblastoma to determine disease severity
- Published on: April 4 2025
Retinoblastoma is a common paediatric eye cancer in children under five years of age1,2. It is most often caused by biallelic mutations of the retinoblastoma gene (RB1), resulting in a loss of function and tumour formation3.
This cancer has high survival rates1, but heritable retinoblastoma — caused by a germline mutation, then a secondary somatic mutation2 — has variable clinical outcomes depending on which parental allele the mutation resides, known as the parent-of-origin effect4 .
What is the parent-of-origin effect?
The parent-of-origin effect is an epigenetic event caused by genomic imprinting that impacts methylation and gene expression4. This causes the phenotypic effects of a gene to vary depending on whether a genetic mutation is inherited maternally or paternally2. This effect is best characterised in imprinting disorders, such as Prader-Willi and Angelman syndromes, where the parent-of-origin effect on chromosome 15 results in different syndromes and phenotypes5.
In the case of heritable retinoblastoma, the parent-of-origin effect alters clinical outcomes2. For example, paternally inherited RB1 variants are associated with a higher likelihood of developing bilateral tumours and secondary cancers in later life than maternally inherited variants6–8. A study by Eloy et al. suggests that this is due to a differentially methylated region (DMR) in intron 2 of RB1 that impacts the tumour-suppressor activity of the gene. They proposed that maternally inherited mutations have a weaker effect on tumour-suppressor activity than paternally inherited mutations, which cause minimal suppressor activity, leading to more severe tumour progression9. Therefore, it is beneficial to identify the parent of origin to inform clinical treatment plans early on.
However, it is estimated that more than 90% of germline cases are caused by de novo mutations, meaning that the parent of origin cannot be assigned because conventional testing requires trio sequencing of familial DNA samples, which would not harbour the mutation2. Yet, despite these difficulties in assigning de novo mutations, they are less likely to predispose to secondary cancers8.
The difficulties of assigning the parent of origin
Studies have determined that the DMR in RB1 is methylated on the maternal allele and unmethylated on the paternal allele6, meaning it can be used to identify the parent of origin of a mutation without the need for trio sequencing2.
To assign the parent of origin of an RB1 mutation by analysing methylation in this region, the variant must, therefore, be phased with the DMR. However, legacy short-read sequencing methods cannot sequence the entire RB1 locus containing the mutation and DMR because short reads cannot provide chromosomal phase information due to their limited read length. Furthermore, secondary assays are required to characterise epigenetic changes, ‘limiting their ability to provide parent-of-origin diagnostics’ in a comprehensive or rapid manner2.
Stacey et al. based at the University of Washington, USA, investigated the potential use of targeted Oxford Nanopore sequencing, as the multiomic method provides epigenetic and genetic data in a single assay. Furthermore, nanopore sequencing generates reads of any length, allowing long fragments of native DNA to be sequenced without losing base modifications. This means that a variant in RB1 can be analysed and phased with the DMR using the same nanopore reads in a single sequencing run, demonstrating the potential to enable confirmation of the parent of origin from just the affected individual.
To target nanopore sequencing, Stacey et al. utilised adaptive sampling, a unique feature of Oxford Nanopore sequencing that provides real-time enrichment of selected regions of interest during sequencing without additional library preparation. The team sequenced seven research samples from individuals diagnosed with hereditary retinoblastoma and nine retinoblastoma research samples with de novo germline mutations. Peripheral blood samples were prepared using the Ligation Sequencing Kit before sequencing on a MinION for 72 hours. Adaptive sampling was employed during sequencing to target a region of the RB1 gene containing both the DMR in intron 2 and the pathogenic variant, with an additional 50 kb flanking region on either side of the targeted locus.
‘This work demonstrates parent-of-origin in disease can be resolved by phasing a disease variant with a differentially methylated signal in the genome, either local or distant, with a targeted long-read sequencing approach’
Stacey et al. 20242
Capturing de novo germline mutations
Using this method, the RB1 variant was sequenced and phased with the DMR, identifying the parent of origin in all research samples from the affected individual for both inherited and de novo variants — without familial DNA samples (Figure 1). Oxford Nanopore sequencing also identified the parent of origin for complex de novo mutations caused by structural variants and somatic mosaicisms, which typically require multiple secondary assays with legacy short-read methods. The authors stated that all ‘previously unknown cases with de novo pathogenic variants in RB1, which account for the majority of heritable cases of [retinoblastoma]’, were resolved with this method.
The results also demonstrated that low-frequency variants can be identified at a variant allele fraction (VAF) of 16% from peripheral blood cells. This was made possible with adaptive sampling, which focused sequencing depth on a targeted region. Stacey et al. found ‘a distinct advantage [with] targeted adaptive sequencing’ because it ‘provides the adequate read depth and ability to phase variants to identify pathogenic variants rapidly’, and provides methylation information.
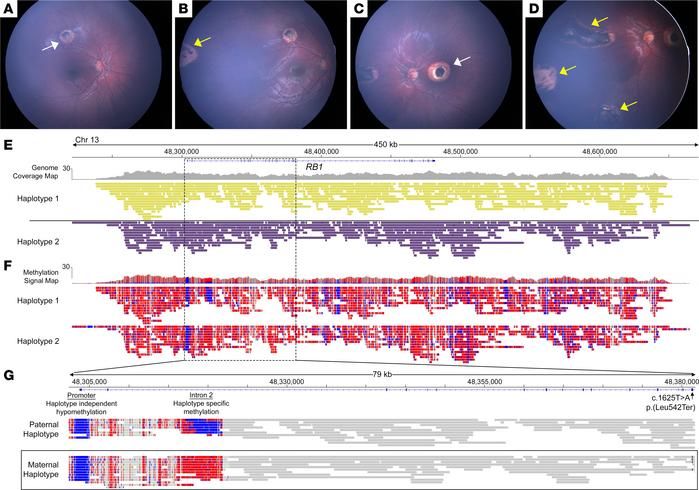
Figure 1. Targeted Oxford Nanopore sequencing generated haplotyped genetic and epigenetic data to assign parent of origin from the proband. A, B, C, and D) Macular (white arrows) and extramacular (yellow arrows) tumours. E) Genomic data, F) epigenomic data, and G), the targeted 79 kb region spanning the promoter region, DMR, and mutation on exon 17 (black lines) of the maternal haplotype, demonstrating maternal inheritance. Figure from Stacey et al.2 and available under Creative Commons license (https://creativecommons.org/licenses/by/4.0).
Accessing novel prognostic biomarkers
The team found that Oxford Nanopore sequencing solves the parent-of-origin question for all cases of retinoblastoma from just the proband, providing access to a potentially important prognostic biomarker for this disease. This is particularly important for identifying variants on the paternal allele, which typically demonstrate worse clinical outcomes with more severe ocular disease than maternally inherited mutations.
In conclusion, targeted Oxford Nanopore sequencing delivers ‘the ability to phase a germline variant with an imprinting signal in a single sequencing run’, potentially providing prognostic information for all retinoblastoma cases. Stacey et al. proposed that implementing this multiomic platform in clinical settings has the potential to provide ‘a more complete diagnostic outlook for children and their families afflicted with [retinoblastoma] to better guide management and therapy in a more personalised manner’.
1. Cancer Research UK. What is retinoblastoma? https://www.cancerresearchuk.org/about-cancer/childrens-cancer/eye-cancer-retinoblastoma/about (2024) [Accessed 20 March 2025]
2. Stacey, A.W. et al. Prognostic importance of direct assignment of parent of origin via long-read genome and epigenome sequencing in retinoblastoma. JCI Insight. 10(4):e188216 (2024). DOI: https://doi.org/10.1172/jci.insight.188216
3. Dimaras, H. et al. Retinoblastoma. Nat. Rev. Dis. Primers. 27:1:15021 (2015). DOI: https://doi.org/10.1038/nrdp.2015.21
4. Lawson, H.A. et al. Genomic imprinting and parent-of-origin effects on complex traits. Nat. Rev. Genet. 14(9):609–617 (2013). DOI: https://doi.org/10.1038/nrg3543
5. Butler, M.G. Imprinting disorders in humans: a review. Curr. Opin. Pediatr. 32(6):719–729 (2020). DOI: https://doi.org/10.1097/MOP.0000000000000965
6. Kanber, D. et al. The human retinoblastoma gene is imprinted. PLoS Genet. 5(12):e1000790 (2009). DOI: https://doi.org/10.1371/journal.pgen.1000790
7. Eiger-Moscovich, M. et al. Familial retinoblastoma: variations in clinical presentation and management based on paternal versus maternal inheritance. J. AAPOS. 28(1):103804 (2024). DOI: https://doi.org/10.1016/j.jaapos.2023.11.007
8. Kleinerman, R. et al. Variation of second cancer risk by family history of retinoblastoma among long-term survivors. J. Clin. Oncol. 30(9):950–957 (2021). DOI: https://doi.org/10.1200/jco.2011.37.0239
9. Eloy P. et al. A parent-of-origin effect impacts the phenotype in low penetrance retinoblastoma families segregating the c.1981C>T/p.Arg661Trp mutation of RB1. PLoS Genet. 12(2):e1005888 (2016). DOI: https://doi.org/10.1371/journal.pgen.1005888







