Comprehensive resolution of challenging genomic variants with Oxford Nanopore de novo genome assemblies
- Published on: May 19 2026
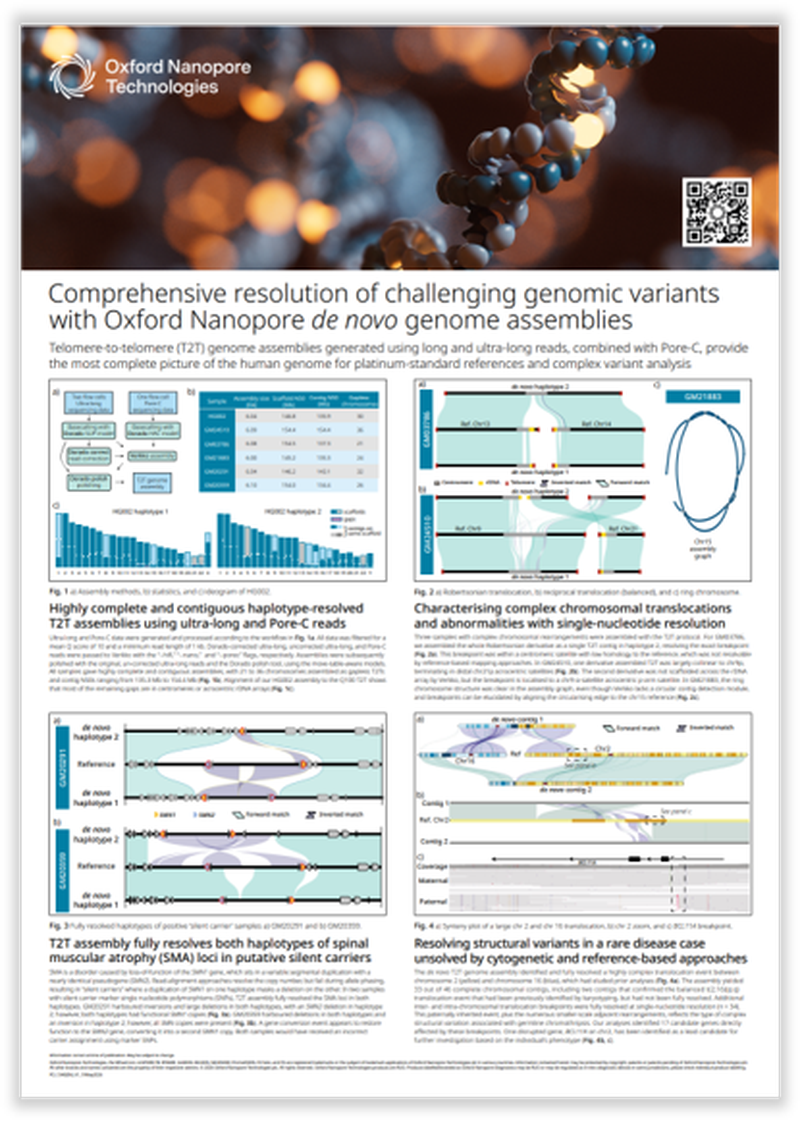
Telomere-to-telomere (T2T) genome assemblies generated using long and ultra-long reads, combined with Pore-C, provide the most complete picture of the human genome for platinum-standard reference and complex variant analysis.
Download the poster to find out:
- How to assemble T2T genomes using nanopore-only data
- About characterising complex chromosomal translocations
- How T2T assemblies can fully resolve haplotypes and previously unresolved structural variants (SVs)







