A POretable Metagenomics Pipeline (aPOMP): Fast and Efficient Nanopore sequencing-based metagenomics
- Published on: May 19 2023
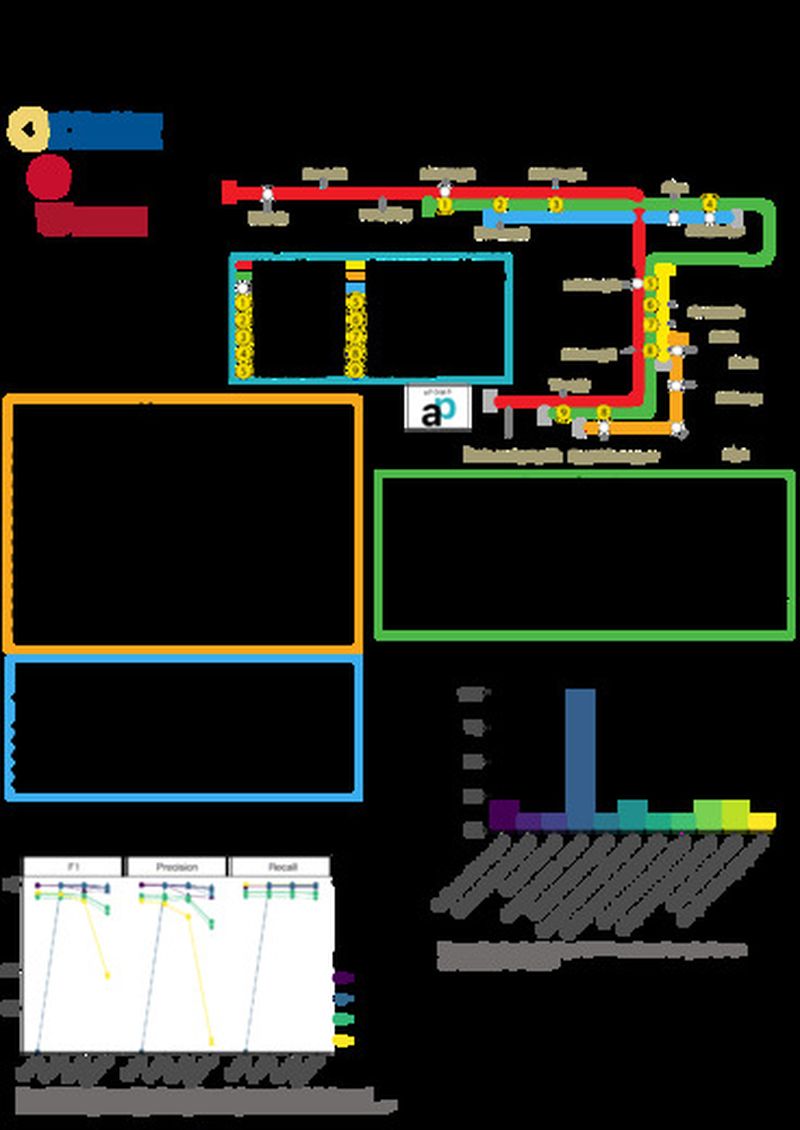
We present aPOMP: a Nextflow based open source metagenomic analysis pipeline for Nanopore sequencing data. Using the full NCBI NT database, aPOMP employs a pre-filtering step to estimate abundant genera, followed by directed alignments to the detected genera and LCA classification to resolve strain level specificity of the organisms in a sample. When requested, antimicrobial resistance genes can also be identified. Compared to existing long read metagenomic pipelines our approach performs well in recall, precision, F1-score, and speed. aPoMP can also be implemented offline in the field on resource limited computers running MinIONs, or with adequate computational resources for basecalling, profile reads in real time. We will present results from aPOMP applied to Nanopore sequencing data from a orthopedic device infection dataset.






