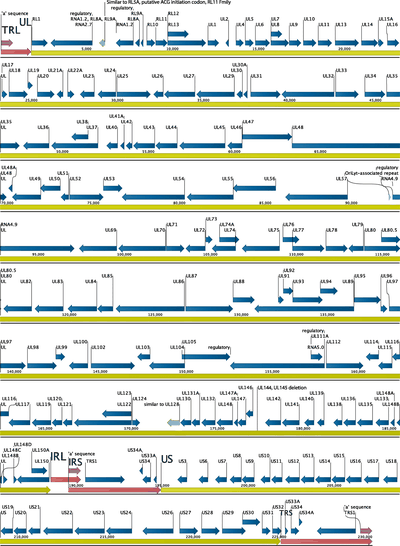
Nanopore sequencing and full genome de novo assembly of human cytomegalovirus TB40/E reveals clonal diversity and structural variations
- Published on: August 2 2018
- Source: BMC Genomics
Human cytomegalovirus (HCMV) has a double-stranded DNA genome of approximately 235 Kbp that is structurally complex including extended GC-rich repeated regions. Genomic recombination events are frequent in HCMV cultures but have also been observed in vivo. Thus, the assembly of HCMV whole genomes from technologies producing shorter than 500 bp sequences is technically challenging. Here we improved the reconstruction of HCMV full genomes by means of a hybrid, de novo genome-assembly bioinformatics pipeline upon data generated from the recently released MinION MkI B sequencer from Oxford Nanopore Technologies.