Exploring the use of Nanopore cDNA sequencing for haplotype phasing in F1 hybrids and polyploid plant species
- Published on: January 22 2019
- Source: PAG 2018
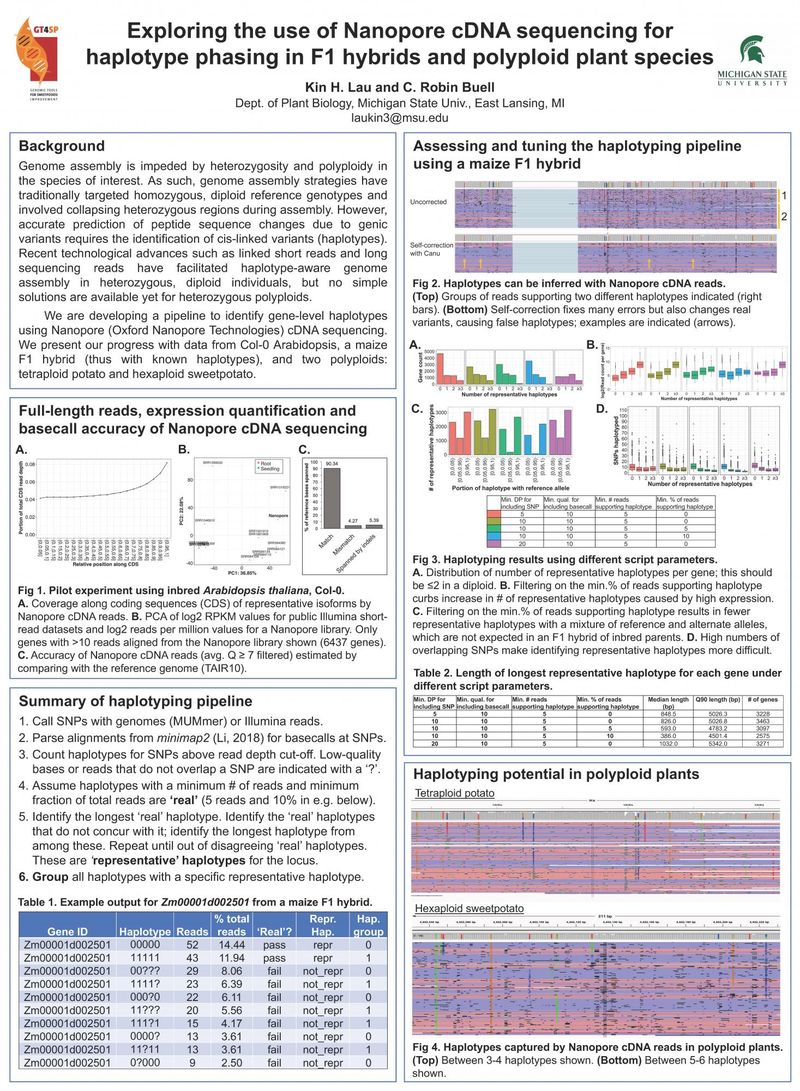
Genome assembly is impeded by heterozygosity and polyploidy in the species of interest. As such, genome assembly strategies have traditionally targeted homozygous, diploid reference genotypes and involved collapsing heterozygous regions during assembly. However, accurate prediction of peptide sequence changes due to genic variants requires the identification of cis-linked variants (haplotypes). Recent technological advances such as linked short reads and long sequencing reads have facilitated haplotype-aware genome assembly in heterozygous, diploid individuals, but no simple solutions are available yet for heterozygous polyploids.
We are developing a pipeline to identify gene-level haplotypes using Nanopore (Oxford Nanopore Technologies) cDNA sequencing. We present our progress with data from Col-0 Arabidopsis, a maize F1 hybrid (thus with known haplotypes), and two polyploids: tetraploid potato and hexaploid sweetpotato.






