De novo sequencing and high-quality assembly of yeast genomes using a MinION device
- Published on: June 4 2018
- Source: London Calling 2018
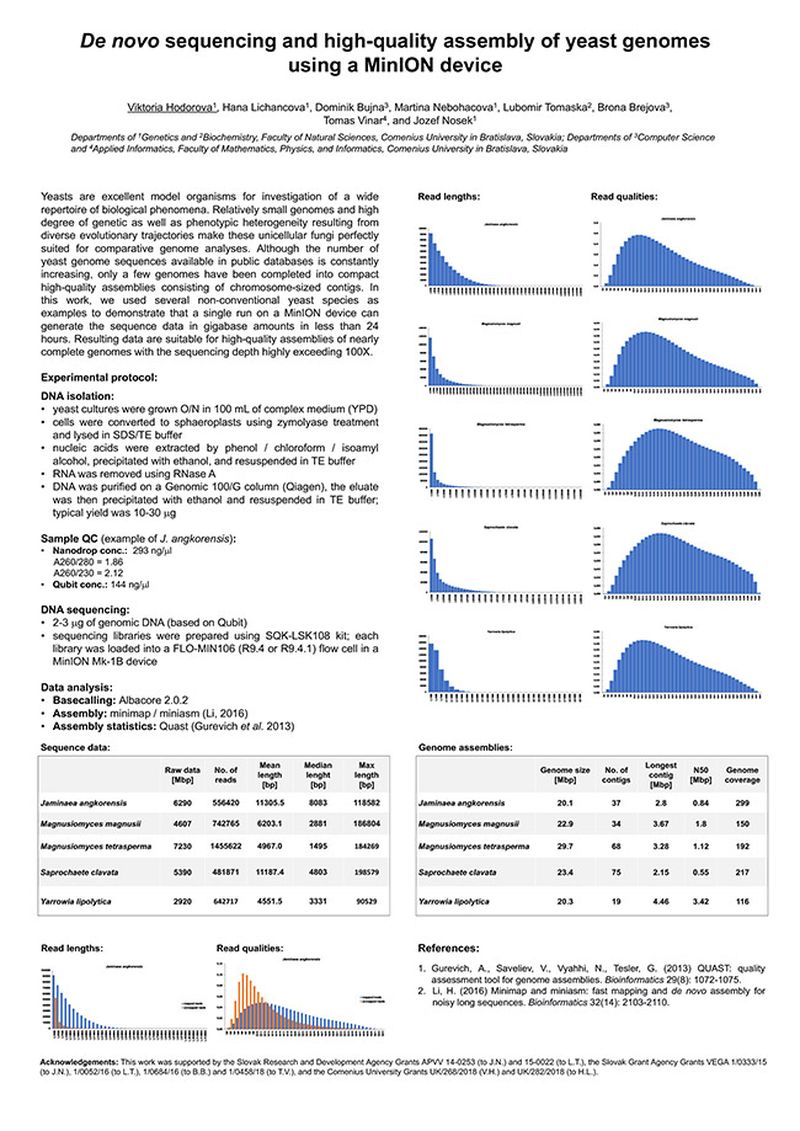
Yeasts are excellent model organisms for investigation of a wide repertoire of biological phenomena. Relatively small genomes and high degree of genetic as well as phenotypic heterogeneity resulting from diverse evolutionary trajectories make these unicellular fungi perfectly suited for comparative genome analyses. Although the number of yeast genome sequences available in public databases is constantly increasing, only a few genomes have been completed into compact high-quality assemblies consisting of chromosome-sized contigs. In this work, we used several non-conventional yeast species as examples to demonstrate that a single run on a MinION device can generate the sequence data in gigabase amounts in less than 24 hours. Resulting data are suitable for high-quality assemblies of nearly complete genomes with the sequencing depth highly exceeding 100X.






