Barcoding-free, high-throughput single-cell sequencing enabled by the 1-read-1-cell paradigm
- Published on: May 28 2022
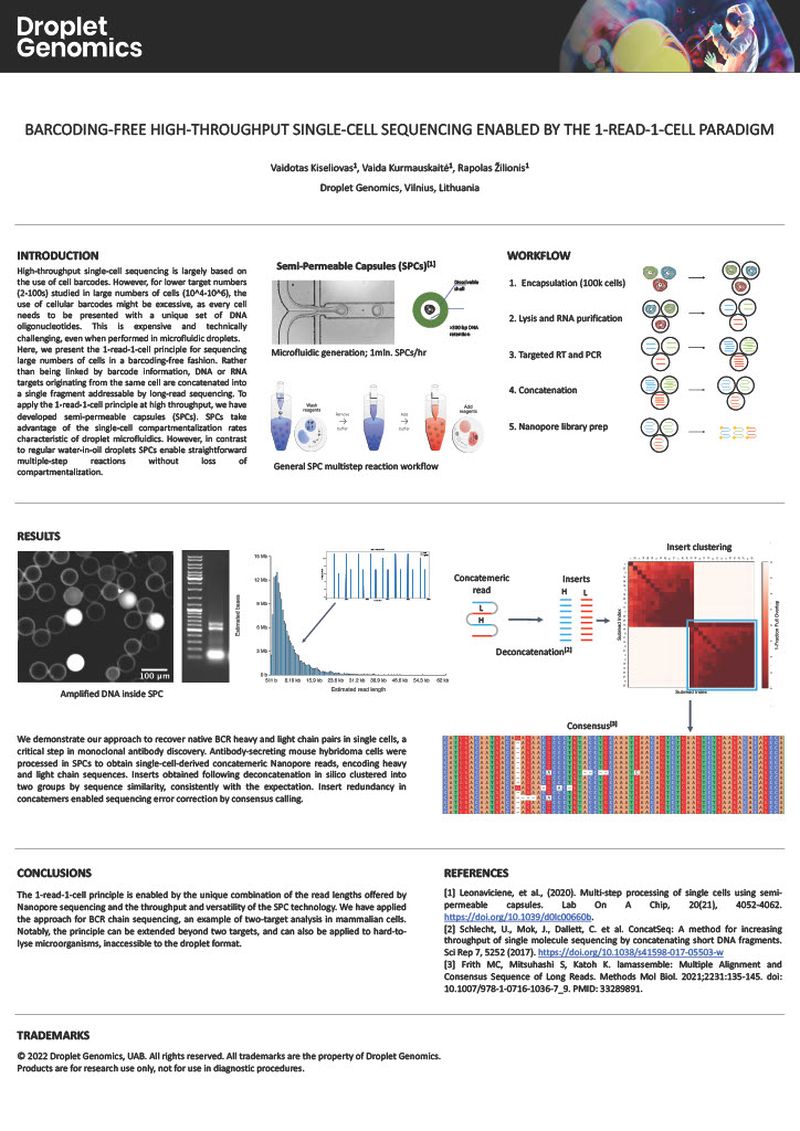
High-throughput single-cell sequencing is largely based on the use of cell barcodes. However, for lower target numbers (2-100s) studied in large numbers of cells (10^4-10^6), the use of cellular barcodes might be excessive, as every cell needs to be presented with a unique set of DNA oligonucleotides. This is expensive and technically challenging, even when performed in microfluidic droplets. Here, we present the 1-read-1-cell principle for sequencing large numbers of cells in a barcoding-free fashion. Rather than being linked by barcode information, DNA or RNA targets originating from the same cell are concatenated into a single fragment addressable by long-read sequencing. To apply the 1-read-1-cell principle at high throughput, we have developed semi-permeable capsules (SPCs). SPCs take advantage of the single-cell compartmentalisation rates characteristic of droplet microfluidics. However, in contrast to regular water-in-oil droplets SPCs enable straightforward multiple-step reactions without loss of compartmentalisation.






