About us
To enable the analysis of anything, by anyone, anywhere
Oxford Nanopore makes a novel generation of DNA/RNA sequencing technology that provides rich data, is fast, accessible and easy to use. Our goal is to disrupt the way that biological analyses are currently performed, and open up new applications that have a profound, positive impact on society.
Who we are
Oxford Nanopore Technologies plc was founded in 2005 as a spin-out from the University of Oxford. The company now employs more than 1000 employees from multiple disciplines including nanopore science, molecular biology and applications, informatics, engineering, electronics, manufacturing and commercialisation. The management team, led by CEO Francis Van Parys, has a track record of delivering disruptive technologies to the market.
Where we are
Oxford Nanopore is a public company, headquartered at the Oxford Science Park outside Oxford, UK, with satellite offices in Cambridge (UK), New York, Cambridge, San Francisco (US), Singapore, Shanghai, Beijing, and a broader commercial presence that includes Japan, Germany, France and India. The company sells to more than 120 countries and is in a period of rapid growth.
In the summer of 2019, we opened a new high-tech manufacturing facility in Oxford.
The technology
Oxford Nanopore has developed a new generation of DNA/RNA sequencing technology. It is the only sequencing technology that offers real-time analysis (for rapid insights), in fully scalable formats from pocket to population scale, that can analyse native DNA or RNA and sequence any length of fragment to achieve short to ultra-long read lengths.
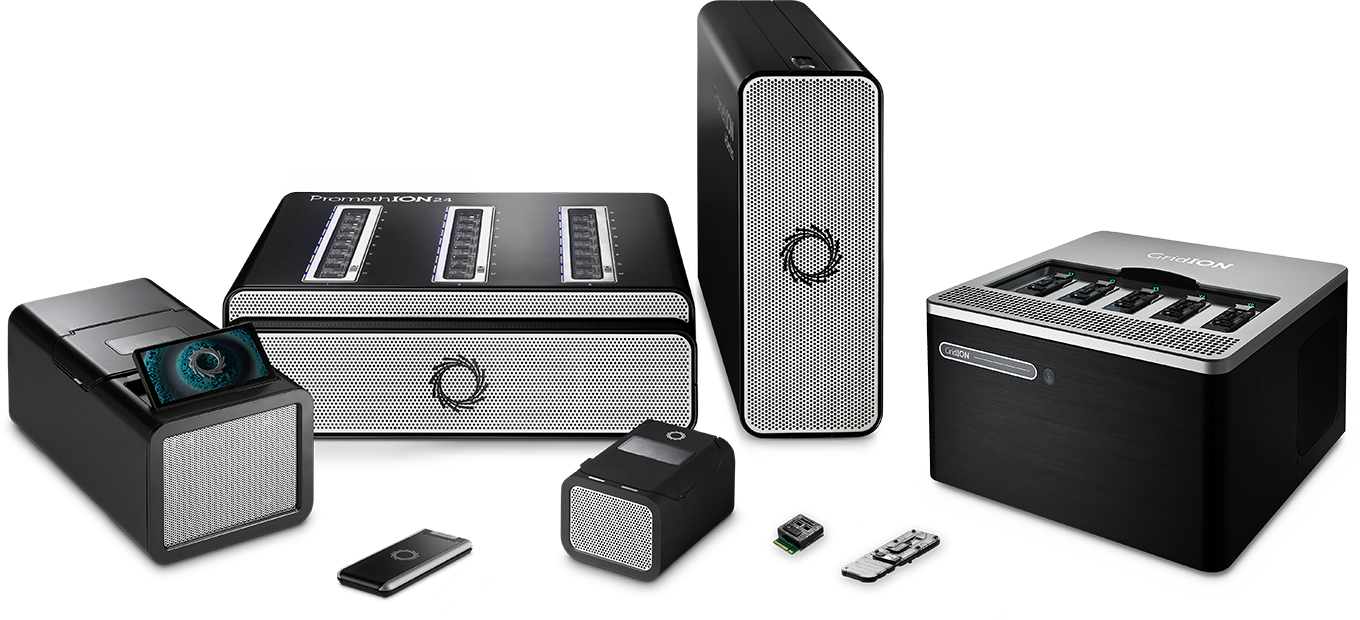
Small formats such as Flongle address the need for on-demand, rapid, smaller tests or experiments, and can be used in labs or in the field. The pocket-sized MinION is a powerful and portable sequencing device that can deliver high volumes of long read sequence data. The benchtop GridION can run up to five MinION Flow Cells at a time, on-demand, for larger genomics projects. PromethION is the largest format for nanopore sequencing, designed to offer on-demand use of up to 48 Flow Cells - capable of delivering more than 10 Tb of sequence data in a full run, and is now being used in population-scale sequencing projects. The palm sized PromethION 2 makes high-output nanopore sequencing broadly accessible
To make the technology suitable for any user, we focus on increasing ease of use and automation. Nanopore sequencing offers easy and rapid preparation, including a ten minute library preparation kit. We also provide a range of analysis workflows.
Who needs DNA / RNA information?

Scientific research
Sequence data is used throughout scientific research, whether in university, government or industrial research groups, to help biologists answer a range of questions.
The majority of users of Oxford Nanopore’s sequencing technology are currently research scientists.

Applied markets
Outside scientific research, DNA/RNA information can be used to support ‘real life’ decision making, whether that is in healthcare, industrial or other environments. Some of these sectors are regulated, such as in healthcare and food safety.
As well as providing devices for lab-based testing, Oxford Nanopore’s sequencing technology is uniquely deployable in distributed, near-s
Applied markets
Healthcare
Oxford Nanopore’s technology is currently being used in multiple external clinical research laboratories, for example to characterise infectious disease samples, to investigate tissue typing in advance of transplants, and to provide insights into reproductive health.
Nanopore technology is also highly suited to the development of diagnostic assays.
A broad range of clinical areas may potentially benefit from rapid genomic insights: What disease does this person have? What is the optimal treatment pathway for this person?
Scientific research
Genome science
What is the structure and function of the human genome?
Human genetics
How does variation between individuals influence their characteristics, disease risk or response to medication?
Pathogens / Microbiology
What is this virus/bacteria/fungus? What makes it pathogenic? Is it resistant to antimicrobial drugs? How could we use this information to prevent or treat the disease that it causes?
Clinical research
How can we use sequence data to personalise medicine? How can we integrate sequence data into clinical decision making? How does a person’s genome influence how they may respond to a disease or an infection?
Environmental
How is the microbial composition of this ocean/glacier/lake changing? What is this species? Is it endangered? What can we understand about the biodiversity within this area?
Plant research
What are the differences between these tomato crops? How can we breed better varieties, that are more productive, long lasting or taste better? How can we apply this knowledge to a variety of plants from cereals to flowers?
Epidemiology
What is this pathogen? How is it changing? How is it being transmitted? At March 2021, approximately a quarter of the COVID-19 sequences in the GISAID platform were sequenced on a nanopore device.
Education
School and university students: Can I use sequencing as a fundamental tool to understand biology, data analysis, experimental design, and become a citizen scientist of the future?
Innovation
With a best-in-class R&D team, Oxford Nanopore's innovation drive has delivered a novel generation of sequencing technology and surrounding, enabling technologies. We continue to invest heavily in innovation, to drive disruption of the scientific markets.
Supported by a broad patent portfolio generated by internal R&D and external collaborations, the Oxford Nanopore pipeline includes more than 2,600 patents and applications. The portfolio includes multiple generations of nanopore-based sensing technologies, including those based on both biological and solid-state nanopores.
Oxford Nanopore’s technology has now been featured in more than 18,000 scientific publications.
You can find out more about the work of the nanopore community at our Resource Centre, and more detail about the research applications in our Applications area.
Anyone anywhere
Accessibility is central to our goal, and it is baked into everything we do, whether that is product design, pricing or how we support our customers.
In 2015, we chose to launch a device to the market that was accessible to any scientist. A MinION starter pack cost $1,000, could be shipped or used anywhere and was easy to use. A thriving community of scientists grew around the device, and these pioneering researchers continue to break boundaries, whether by sequencing in the Antarctic or characterising drug resistant tuberculosis in Madagascar.
Today, as well as providing tools for analysis in any location, we also provide powerful devices that can provide rich, deep analyses of very large datasets – for example for population-scale human genomes. The devices available are built on the same platform and is key to our goal of addressing all needs for biological insights.
Get in touch
Subscribe
Talk to us
If you have any questions about our products or services, chat directly with a member of our sales team.










