Continuous development and improvement
MinION was launched into the MinION Access Programme in Spring 2014 and made commercially available in May 2015. Since that time, Oxford Nanopore has delivered continual improvement in performance, usability and other metrics. The format of the hardware, software and chemistry is continuously improved; typically through upgrades in the consumable and so requiring no device change.
Oxford Nanopore deploys continuous, iterative improvement of its technology so that the performance and user experience are improved as rapidly as possible. Typically, improvements are deployed through consumables, software or firmware so that devices do not need to be changed. Iterative improvement processes will continue throughout the lifetime of all Oxford Nanopore products, however the Q-Line is available for those who wish to lock down their workflows.
Continuous improvement has been achieved through a combination of software updates, changes in the library preparation kits and protocols, changes in the flow cell design and changes in the flow cell chemistries.
Examples of developments to date

Library preparation kits
- Many elements of the library preparation process have been improved over time to deliver enhanced performance. Additional kits and protocols have been introduced to enable new applications, for example cDNA sequencing and barcoding of genomic DNA and amplicons.
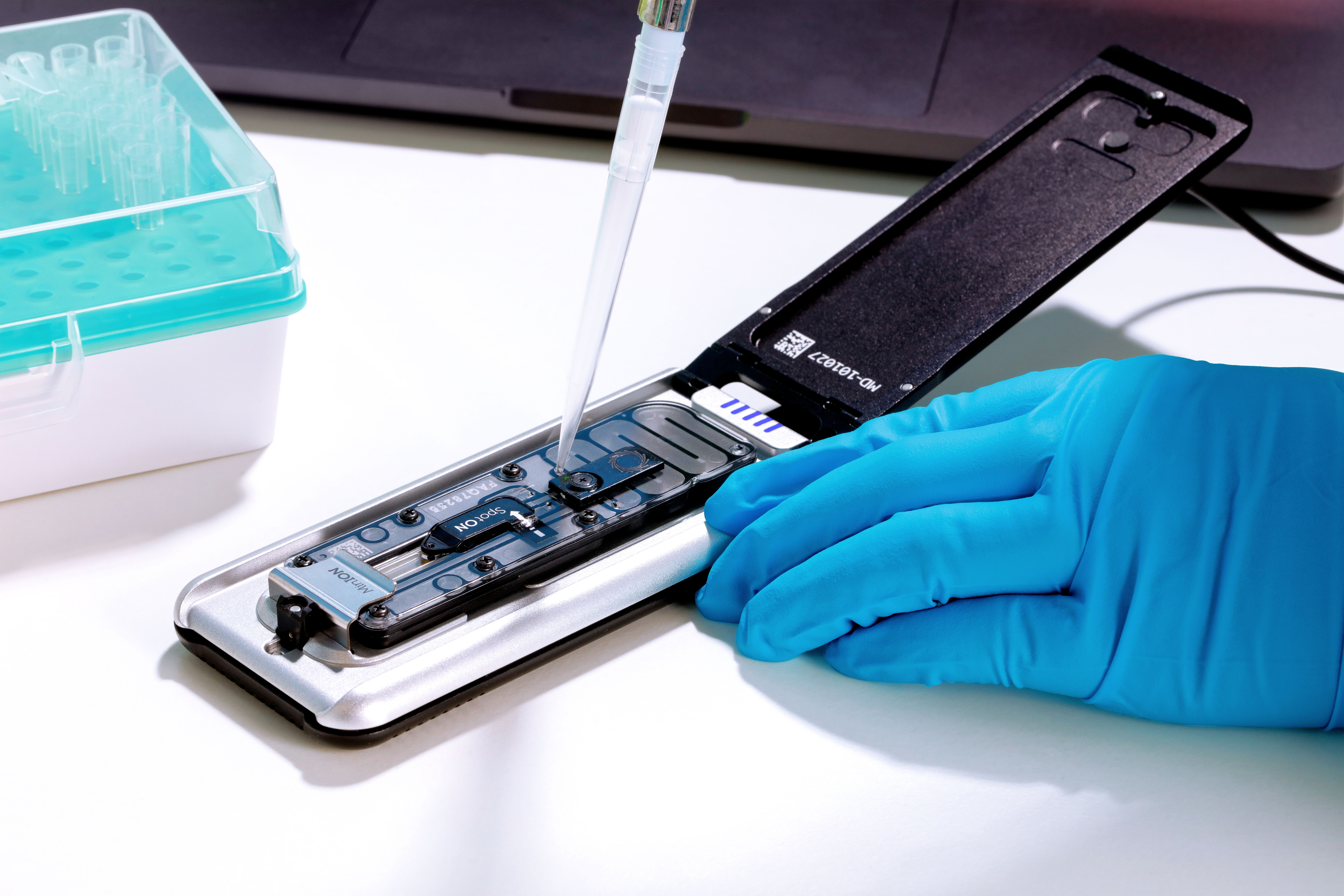
- We have also made several changes to our library preparation kits to improve the user experience. These include reducing the number of steps and consequently the time taken, and improving robustness and performance. The Rapid Sequencing kit prepares a library in 5-10 minutes.
- VolTRAX, an automated library preparation device, is designed for ease of use, anywhere, and to make it easier to prepare high quality libraries for the best sequencing results.
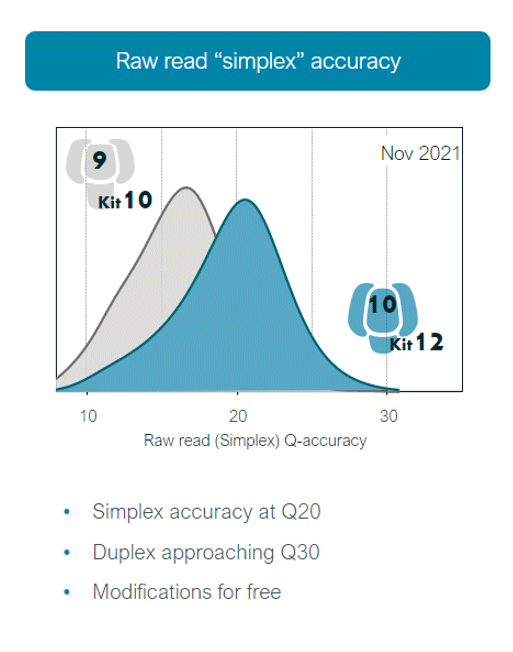
- In 2022, users are offered the latest "kit12", used with R10.4 flow cells to achieve the highest accuracy sequencing data
Sequencing chemistry
Nanopore sequencing takes place in consumables called flow cells, which contain bespoke nanopore sensors, motor proteins and associated chemistries. Multiple upgrades in the sequencing apparatus have been delivered over time, for enhanced yields and accuracy.
Three flow cells: Flongle, MinION/GridION and PromethION, are available. While different scales, the same sequencing chemistry is used across these consumables.
- In the summer of 2016, 'R9' was released to supercede the previous R7 flow cell. This was designed to improve sequencing accuracy.
- In October 2016, new flow cells containing R9.4 were shipped, increasing sequencing speeds to 450 bases per second and enabling 10Gb DNA sequencing data to be obtained from a MinION Flow Cell.
- In May 2017, R9.5 was shipped, to be compatible with the new 1D squared method of sequencing. Oxford Nanopore was at this stage producing more than 20Gb from a single Flow Cell.
- In June 2019, updates to the flow cells were enabling users to achieve tens of Gb of sequencing data from a single flow cell
- In 2019, R10 flow cells were shipped to users in early access. Early results indicate enhanced consensus accuracy and accurate variant calling with this novel nanopore.
- In 2020, R10.3 – an updated version of the R10 nanopore, was introduced, for further enhanced performance.
- In 2021, the Q20+ kit was released, and users were also able to use Duplex chemistry to drive raw read accuracy towards 99.9%
- In 2022, the R10.4 flow cell is now available, offering the latest design of nanopore optimised for accuracy
Device iteration
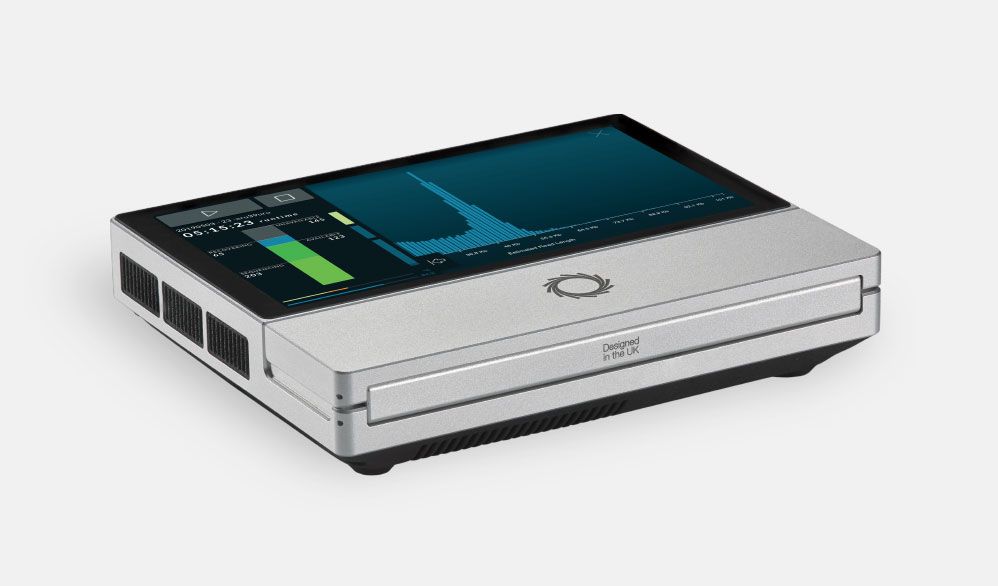

- After early access for MinION in 2014, in May 2015 the second version of the MinION, the MinION Mk1 was introduced. The MinION Mk1 was a full production device featuring improvements of performance and ease of use.
- In May 2016, the MinION Mk 1B was introduced. Preparing for future iterations of nanopore chemistry it included improvements such as greater temperature control of the flow cell.
- In 2019, the GridION X5 transitioned to the matured GridION Mk1. The beta PromethION also transitioned to P24 and P48; these upgrades included improved computational power, temperature control and a variety of other performance-enhancing qualities.
- The MinION Mk1C integrates sequencing, GPU for real time data analysis, screen and real time connectivity.
Driving yields

Speed of individual nanopore processing
DNA can translocate a nanopore at a variety of speeds. The faster the DNA passes through, the faster the data is generated and the higher the yields. Speed is affected by multiple factors including buffers, temperature and the motor enzyme deployed. In combination with other improvements, increasing translocation speed means that flow cells have gone from being able to produce hundreds of megabases of data in 2014, to tens of gigabases in 2020.
- During the first year, Oxford Nanopore recommended a translocation speed of around 30 bases per second per nanopore.
- By 2015 users could run at 70bps.
- With the introduction of R9 in the summer of 2016, DNA was passed through the nanopore at 250+bps.
- From October 2016, processing speeds have been ~400-450 bases per second for DNA sequencing.
- Electronics are specced to have the potential to measure as much as 1000 bases per second
Duration of use
- Nanopore devices do not have a fixed run time; users may run the flow cell for as long as it takes to accumulate sufficient data for their needs. The total available life time of a flow cell does not need to be consumed in a single experiment.
- More recent releases of flow cells and software have enabled flow cells to be run for longer (at a constant price), enabling resulting in increased overall yields.

Performance: the instrument control software (MinKNOW)
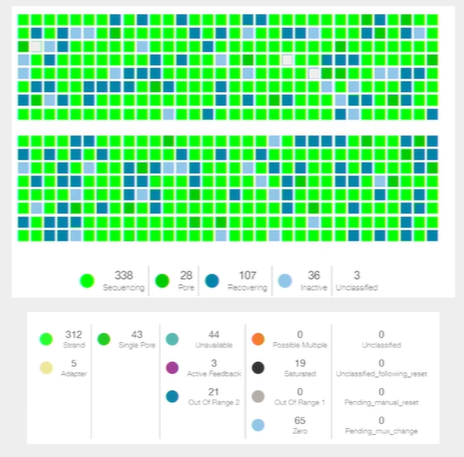

Performance: the instrument control software (MinKNOW)
- New versions of MinKNOW, the software that enables users to run nanopore sequencers, have been released to improve MinION performance for all applications. For example, adjusting the frequency of data sampling can improve yield and accuracy.
- Further upgrades have included features such as progressive unblock, allowing enhanced yields by allowing longer run times, and improvement in 'MUX' processes to select the most productive channels early in an experiment – these have all contributed to higher yields.
The right throughput for the right project
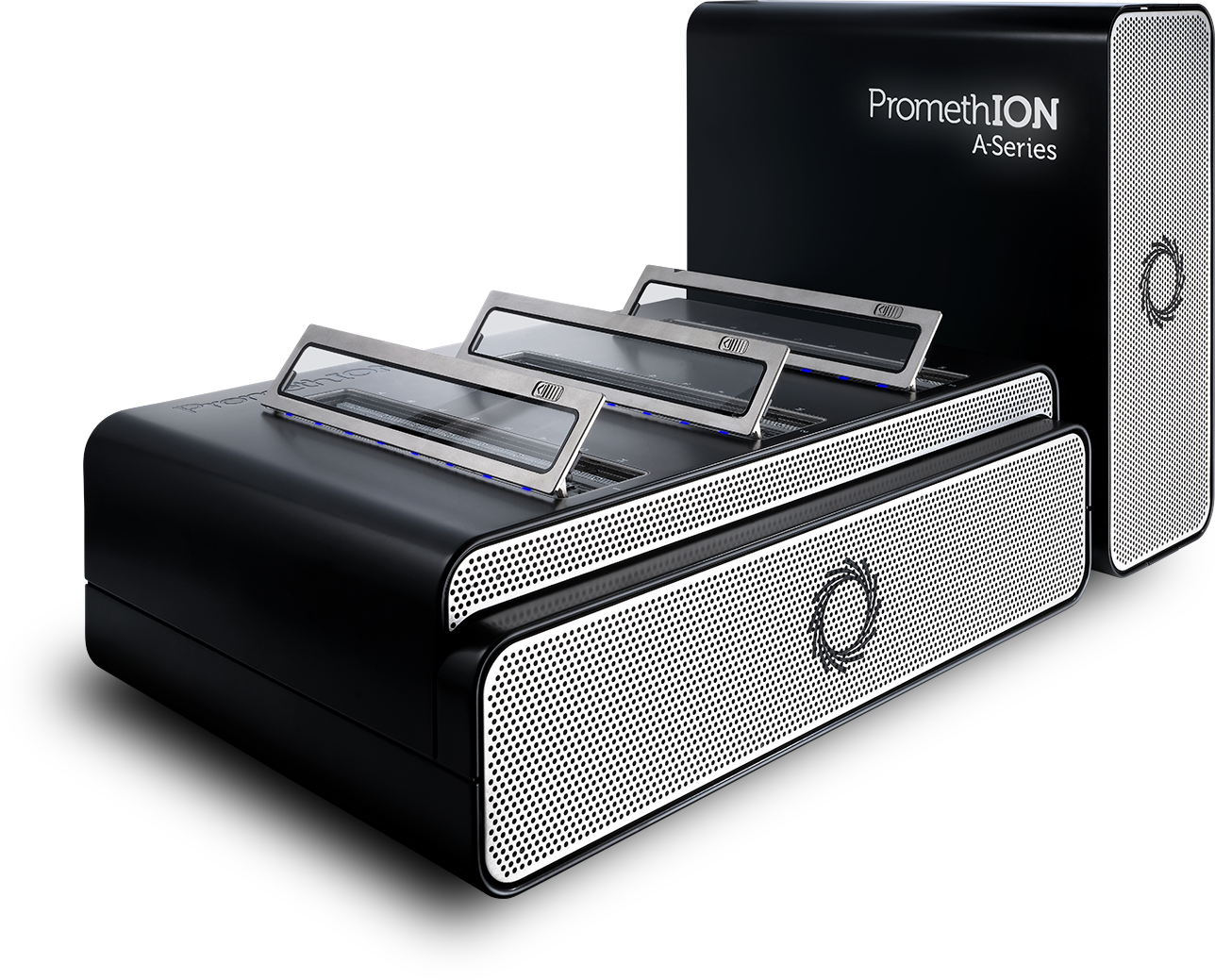

- Following on from the success of the MinION, the GridION X5 was launched in 2017. This can run up to five MinION Flow Cells with onboard compute and has now been upgraded to the GridION Mk1, with enhanced compute power.
- The PromethION 24 and 48 are now available and offers nearly 300 times the power of a MinION, except modular and on-demand. The PromethION is designed to operate up to 24 or 48 Flow Cells individually or together.
- The Flongle was introduced in 2019, for rapid, smaller analyses.
- Oxford Nanopore continues to develop SmidgION, a smartphone sequencer, and other form factors.
Analysis tools
Based on electronics rather than optics, nanopore technology can scale to any size:
- Continuous iteration of basecalling algorithms, device control software and the availability of a broad range of analysis tools from Oxford Nanopore or the community, have contributed to improvements in accuracy of raw and consensus data, and improved variant detection.
- Read more about the latest accuracy performance made possible by continuous iteration of analysis algorithms.
Get in touch
Subscribe
Talk to us
If you have any questions about our products or services, chat directly with a member of our sales team.
Book a sales call
To book a call with one of our sales team, please click below.







