Ligation sequencing amplicons - dual barcoding (SQK-LSK109 with EXP-NBD104, EXP-NBD114, and EXP-PBC096) (DBC_9098_v109_revI_16Apr2020)
MinION: Protocol
V DBC_9098_v109_revI_16Apr2020
FOR RESEARCH USE ONLY
Contents
Introduction to the protocol
Library preparation
- 4. End-prep
- 5. Ligation of Barcode Adapter
- 6. Barcoding PCR
- 7. End-prep
- 8. Native barcode ligation
- 9. Adapter ligation and clean-up
- 10. Priming and loading the SpotON flow cell
Sequencing and data analysis
Troubleshooting
1. Overview of the protocol
This is a Legacy product.
The kit and expansions have been discontinued. Should you require any assistance or have questions, please contact our Customer Support support@nanoporetech.com.
Introduction to the dual barcoding protocol
This protocol allows massively parallel sequencing of up to 2,304 samples (gDNA or amplicons) on a single flow cell. It uses both the PCR Barcoding Expansion 1-96 (EXP-PBC096) and the Native Barcoding Expansions 1-12 and 13-24 (EXP-NBD104 and EXP-NBD114). First, up to 96 barcodes are added to the DNA ends with PCR. Up to 24 pools of 96 samples can then have a secondary barcode added through the ligation of one of 24 native barcodes. All primary pools can then be combined into a single library and sequenced.
Steps in the sequencing workflow:
Prepare for your experiment
You will need to:
- Extract your DNA, and check its length, quantity and purity. The quality checks performed during the protocol are essential in ensuring experimental success.
- Ensure you have your sequencing kit, the correct equipment and third-party reagents
- Download the software for acquiring and analysing your data
- Check your flow cell to ensure it has enough pores for a good sequencing run
Library preparation
You will need to:
- Prepare the DNA ends for adapter attachment
- Attach barcoding adapters supplied in the 96 PCR Barcoding kit to the DNA ends
- Amplify each barcoded sample by PCR, then pool the samples together
- Prepare the DNA ends for native barcode attachment
- Ligate native barcodes to the DNA ends
- Attach sequencing adapters supplied in the kit to the DNA ends
- Prime the flow cell, and load your DNA library into the flow cell
Sequencing and analysis
You will need to:
- Start a sequencing run using the MinKNOW software, which will collect raw data from the device and convert it into basecalled reads
- Use the Guppy software to split the reads by barcode
96 barcode sequences
| Component | Sequence |
|---|---|
| BC01 / RB01 | AAGAAAGTTGTCGGTGTCTTTGTG |
| BC02 / RB02 | TCGATTCCGTTTGTAGTCGTCTGT |
| BC03 / RB03 | GAGTCTTGTGTCCCAGTTACCAGG |
| BC04 / RB04 | TTCGGATTCTATCGTGTTTCCCTA |
| BC05 / RB05 | CTTGTCCAGGGTTTGTGTAACCTT |
| BC06 / RB06 | TTCTCGCAAAGGCAGAAAGTAGTC |
| BC07 / RB07 | GTGTTACCGTGGGAATGAATCCTT |
| BC08 / RB08 | TTCAGGGAACAAACCAAGTTACGT |
| BC09 / RB09 | AACTAGGCACAGCGAGTCTTGGTT |
| BC10 / RB10 | AAGCGTTGAAACCTTTGTCCTCTC |
| BC11 / RB11 | GTTTCATCTATCGGAGGGAATGGA |
| BC12 / RB12 | CAGGTAGAAAGAAGCAGAATCGGA |
| BC13 / 16S13 / RB13 | AGAACGACTTCCATACTCGTGTGA |
| BC14 / 16S14 / RB14 | AACGAGTCTCTTGGGACCCATAGA |
| BC15 / 16S15 / RB15 | AGGTCTACCTCGCTAACACCACTG |
| BC16 / 16S16 / RB16 | CGTCAACTGACAGTGGTTCGTACT |
| BC17 / 16S17 / RB17 | ACCCTCCAGGAAAGTACCTCTGAT |
| BC18 / 16S18 / RB18 | CCAAACCCAACAACCTAGATAGGC |
| BC19 / 16S19 / RB19 | GTTCCTCGTGCAGTGTCAAGAGAT |
| BC20 / 16S20 / RB20 | TTGCGTCCTGTTACGAGAACTCAT |
| BC21 / 16S21 / RB21 | GAGCCTCTCATTGTCCGTTCTCTA |
| BC22 / 16S22 / RB22 | ACCACTGCCATGTATCAAAGTACG |
| BC23 / 16S23 / RB23 | CTTACTACCCAGTGAACCTCCTCG |
| BC24 / 16S24 / RB24 | GCATAGTTCTGCATGATGGGTTAG |
| BC25 / RB25 | GTAAGTTGGGTATGCAACGCAATG |
| BC26 / RB26 | CATACAGCGACTACGCATTCTCAT |
| BC27 / RB27 | CGACGGTTAGATTCACCTCTTACA |
| BC28 / RB28 | TGAAACCTAAGAAGGCACCGTATC |
| BC29 / RB29 | CTAGACACCTTGGGTTGACAGACC |
| BC30 / RB30 | TCAGTGAGGATCTACTTCGACCCA |
| BC31 / RB31 | TGCGTACAGCAATCAGTTACATTG |
| BC32 / RB32 | CCAGTAGAAGTCCGACAACGTCAT |
| BC33 / RB33 | CAGACTTGGTACGGTTGGGTAACT |
| BC34 / RB34 | GGACGAAGAACTCAAGTCAAAGGC |
| BC35 / RB35 | CTACTTACGAAGCTGAGGGACTGC |
| BC36 / RB36 | ATGTCCCAGTTAGAGGAGGAAACA |
| BC37 / RB37 | GCTTGCGATTGATGCTTAGTATCA |
| BC38 / RB38 | ACCACAGGAGGACGATACAGAGAA |
| BC39 / RB39 | CCACAGTGTCAACTAGAGCCTCTC |
| BC40 / RB40 | TAGTTTGGATGACCAAGGATAGCC |
| BC41 / RB41 | GGAGTTCGTCCAGAGAAGTACACG |
| BC42 / RB42 | CTACGTGTAAGGCATACCTGCCAG |
| BC43 / RB43 | CTTTCGTTGTTGACTCGACGGTAG |
| BC44 / RB44 | AGTAGAAAGGGTTCCTTCCCACTC |
| BC45 / RB45 | GATCCAACAGAGATGCCTTCAGTG |
| BC46 / RB46 | GCTGTGTTCCACTTCATTCTCCTG |
| BC47 / RB47 | GTGCAACTTTCCCACAGGTAGTTC |
| BC48 / RB48 | CATCTGGAACGTGGTACACCTGTA |
| BC49 / RB49 | ACTGGTGCAGCTTTGAACATCTAG |
| BC50 / RB50 | ATGGACTTTGGTAACTTCCTGCGT |
| BC51 / RB51 | GTTGAATGAGCCTACTGGGTCCTC |
| BC52 / RB52 | TGAGAGACAAGATTGTTCGTGGAC |
| BC53 / RB53 | AGATTCAGACCGTCTCATGCAAAG |
| BC54 / RB54 | CAAGAGCTTTGACTAAGGAGCATG |
| BC55 / RB55 | TGGAAGATGAGACCCTGATCTACG |
| BC56 / RB56 | TCACTACTCAACAGGTGGCATGAA |
| BC57 / RB57 | GCTAGGTCAATCTCCTTCGGAAGT |
| BC58 / RB58 | CAGGTTACTCCTCCGTGAGTCTGA |
| BC59 / RB59 | TCAATCAAGAAGGGAAAGCAAGGT |
| BC60 / RB60 | CATGTTCAACCAAGGCTTCTATGG |
| BC61 / RB61 | AGAGGGTACTATGTGCCTCAGCAC |
| BC62 / RB62 | CACCCACACTTACTTCAGGACGTA |
| BC63 / RB63 | TTCTGAAGTTCCTGGGTCTTGAAC |
| BC64 / RB64 | GACAGACACCGTTCATCGACTTTC |
| BC65 / RB65 | TTCTCAGTCTTCCTCCAGACAAGG |
| BC66 / RB66 | CCGATCCTTGTGGCTTCTAACTTC |
| BC67 / RB67 | GTTTGTCATACTCGTGTGCTCACC |
| BC68 / RB68 | GAATCTAAGCAAACACGAAGGTGG |
| BC69 / RB69 | TACAGTCCGAGCCTCATGTGATCT |
| BC70 / RB70 | ACCGAGATCCTACGAATGGAGTGT |
| BC71 / RB71 | CCTGGGAGCATCAGGTAGTAACAG |
| BC72 / RB72 | TAGCTGACTGTCTTCCATACCGAC |
| BC73 / RB73 | AAGAAACAGGATGACAGAACCCTC |
| BC74 / RB74 | TACAAGCATCCCAACACTTCCACT |
| BC75 / RB75 | GACCATTGTGATGAACCCTGTTGT |
| BC76 / RB76 | ATGCTTGTTACATCAACCCTGGAC |
| BC77 / RB77 | CGACCTGTTTCTCAGGGATACAAC |
| BC78 / RB78 | AACAACCGAACCTTTGAATCAGAA |
| BC79 / RB79 | TCTCGGAGATAGTTCTCACTGCTG |
| BC80 / RB80 | CGGATGAACATAGGATAGCGATTC |
| BC81 / RB81 | CCTCATCTTGTGAAGTTGTTTCGG |
| BC82 / RB82 | ACGGTATGTCGAGTTCCAGGACTA |
| BC83 / RB83 | TGGCTTGATCTAGGTAAGGTCGAA |
| BC84 / RB84 | GTAGTGGACCTAGAACCTGTGCCA |
| BC85 / RB85 | AACGGAGGAGTTAGTTGGATGATC |
| BC86 / RB86 | AGGTGATCCCAACAAGCGTAAGTA |
| BC87 / RB87 | TACATGCTCCTGTTGTTAGGGAGG |
| BC88 / RB88 | TCTTCTACTACCGATCCGAAGCAG |
| BC89 / RB89 | ACAGCATCAATGTTTGGCTAGTTG |
| BC90 / RB90 | GATGTAGAGGGTACGGTTTGAGGC |
| BC91 / RB91 | GGCTCCATAGGAACTCACGCTACT |
| BC92 / RB92 | TTGTGAGTGGAAAGATACAGGACC |
| BC93 / RB93 | AGTTTCCATCACTTCAGACTTGGG |
| BC94 / RB94 | GATTGTCCTCAAACTGCCACCTAC |
| BC95 / RB95 | CCTGTCTGGAAGAAGAATGGACTT |
| BC96 / RB96 | CTGAACGGTCATAGAGTCCACCAT |
Native barcode sequences
Below is the full list of our native barcode (NB01-96) sequences. The first 24 unique barcodes are available in the Native Barcoding Kit 24 V14 (SQK-NBD114.24). The Native Barcoding Kit 96 V14 (SQK-NBD114.96) include the first 24 native barcodes, with the additional 72 unique barcodes. The native barcodes are shipped at 640 nM.
In addition to the barcodes, there are also flanking sequences which add an extra level of context during analysis.
Barcode flanking sequences:
Forward sequence: 5' - AAGGTTAA - barcode - CAGCACCT - 3' Reverse sequence: 5' - GGTGCTG - barcode - TTAACCTTAGCAAT - 3'
Native barcode sequences
| Component | Forward sequence | Reverse sequence |
|---|---|---|
| NB01 | CACAAAGACACCGACAACTTTCTT | AAGAAAGTTGTCGGTGTCTTTGTG |
| NB02 | ACAGACGACTACAAACGGAATCGA | TCGATTCCGTTTGTAGTCGTCTGT |
| NB03 | CCTGGTAACTGGGACACAAGACTC | GAGTCTTGTGTCCCAGTTACCAGG |
| NB04 | TAGGGAAACACGATAGAATCCGAA | TTCGGATTCTATCGTGTTTCCCTA |
| NB05 | AAGGTTACACAAACCCTGGACAAG | CTTGTCCAGGGTTTGTGTAACCTT |
| NB06 | GACTACTTTCTGCCTTTGCGAGAA | TTCTCGCAAAGGCAGAAAGTAGTC |
| NB07 | AAGGATTCATTCCCACGGTAACAC | GTGTTACCGTGGGAATGAATCCTT |
| NB08 | ACGTAACTTGGTTTGTTCCCTGAA | TTCAGGGAACAAACCAAGTTACGT |
| NB09 | AACCAAGACTCGCTGTGCCTAGTT | AACTAGGCACAGCGAGTCTTGGTT |
| NB10 | GAGAGGACAAAGGTTTCAACGCTT | AAGCGTTGAAACCTTTGTCCTCTC |
| NB11 | TCCATTCCCTCCGATAGATGAAAC | GTTTCATCTATCGGAGGGAATGGA |
| NB12 | TCCGATTCTGCTTCTTTCTACCTG | CAGGTAGAAAGAAGCAGAATCGGA |
| NB13 | AGAACGACTTCCATACTCGTGTGA | TCACACGAGTATGGAAGTCGTTCT |
| NB14 | AACGAGTCTCTTGGGACCCATAGA | TCTATGGGTCCCAAGAGACTCGTT |
| NB15 | AGGTCTACCTCGCTAACACCACTG | CAGTGGTGTTAGCGAGGTAGACCT |
| NB16 | CGTCAACTGACAGTGGTTCGTACT | AGTACGAACCACTGTCAGTTGACG |
| NB17 | ACCCTCCAGGAAAGTACCTCTGAT | ATCAGAGGTACTTTCCTGGAGGGT |
| NB18 | CCAAACCCAACAACCTAGATAGGC | GCCTATCTAGGTTGTTGGGTTTGG |
| NB19 | GTTCCTCGTGCAGTGTCAAGAGAT | ATCTCTTGACACTGCACGAGGAAC |
| NB20 | TTGCGTCCTGTTACGAGAACTCAT | ATGAGTTCTCGTAACAGGACGCAA |
| NB21 | GAGCCTCTCATTGTCCGTTCTCTA | TAGAGAACGGACAATGAGAGGCTC |
| NB22 | ACCACTGCCATGTATCAAAGTACG | CGTACTTTGATACATGGCAGTGGT |
| NB23 | CTTACTACCCAGTGAACCTCCTCG | CGAGGAGGTTCACTGGGTAGTAAG |
| NB24 | GCATAGTTCTGCATGATGGGTTAG | CTAACCCATCATGCAGAACTATGC |
| NB25 | GTAAGTTGGGTATGCAACGCAATG | CATTGCGTTGCATACCCAACTTAC |
| NB26 | CATACAGCGACTACGCATTCTCAT | ATGAGAATGCGTAGTCGCTGTATG |
| NB27 | CGACGGTTAGATTCACCTCTTACA | TGTAAGAGGTGAATCTAACCGTCG |
| NB28 | TGAAACCTAAGAAGGCACCGTATC | GATACGGTGCCTTCTTAGGTTTCA |
| NB29 | CTAGACACCTTGGGTTGACAGACC | GGTCTGTCAACCCAAGGTGTCTAG |
| NB30 | TCAGTGAGGATCTACTTCGACCCA | TGGGTCGAAGTAGATCCTCACTGA |
| NB31 | TGCGTACAGCAATCAGTTACATTG | CAATGTAACTGATTGCTGTACGCA |
| NB32 | CCAGTAGAAGTCCGACAACGTCAT | ATGACGTTGTCGGACTTCTACTGG |
| NB33 | CAGACTTGGTACGGTTGGGTAACT | AGTTACCCAACCGTACCAAGTCTG |
| NB34 | GGACGAAGAACTCAAGTCAAAGGC | GCCTTTGACTTGAGTTCTTCGTCC |
| NB35 | CTACTTACGAAGCTGAGGGACTGC | GCAGTCCCTCAGCTTCGTAAGTAG |
| NB36 | ATGTCCCAGTTAGAGGAGGAAACA | TGTTTCCTCCTCTAACTGGGACAT |
| NB37 | GCTTGCGATTGATGCTTAGTATCA | TGATACTAAGCATCAATCGCAAGC |
| NB38 | ACCACAGGAGGACGATACAGAGAA | TTCTCTGTATCGTCCTCCTGTGGT |
| NB39 | CCACAGTGTCAACTAGAGCCTCTC | GAGAGGCTCTAGTTGACACTGTGG |
| NB40 | TAGTTTGGATGACCAAGGATAGCC | GGCTATCCTTGGTCATCCAAACTA |
| NB41 | GGAGTTCGTCCAGAGAAGTACACG | CGTGTACTTCTCTGGACGAACTCC |
| NB42 | CTACGTGTAAGGCATACCTGCCAG | CTGGCAGGTATGCCTTACACGTAG |
| NB43 | CTTTCGTTGTTGACTCGACGGTAG | CTACCGTCGAGTCAACAACGAAAG |
| NB44 | AGTAGAAAGGGTTCCTTCCCACTC | GAGTGGGAAGGAACCCTTTCTACT |
| NB45 | GATCCAACAGAGATGCCTTCAGTG | CACTGAAGGCATCTCTGTTGGATC |
| NB46 | GCTGTGTTCCACTTCATTCTCCTG | CAGGAGAATGAAGTGGAACACAGC |
| NB47 | GTGCAACTTTCCCACAGGTAGTTC | GAACTACCTGTGGGAAAGTTGCAC |
| NB48 | CATCTGGAACGTGGTACACCTGTA | TACAGGTGTACCACGTTCCAGATG |
| NB49 | ACTGGTGCAGCTTTGAACATCTAG | CTAGATGTTCAAAGCTGCACCAGT |
| NB50 | ATGGACTTTGGTAACTTCCTGCGT | ACGCAGGAAGTTACCAAAGTCCAT |
| NB51 | GTTGAATGAGCCTACTGGGTCCTC | GAGGACCCAGTAGGCTCATTCAAC |
| NB52 | TGAGAGACAAGATTGTTCGTGGAC | GTCCACGAACAATCTTGTCTCTCA |
| NB53 | AGATTCAGACCGTCTCATGCAAAG | CTTTGCATGAGACGGTCTGAATCT |
| NB54 | CAAGAGCTTTGACTAAGGAGCATG | CATGCTCCTTAGTCAAAGCTCTTG |
| NB55 | TGGAAGATGAGACCCTGATCTACG | CGTAGATCAGGGTCTCATCTTCCA |
| NB56 | TCACTACTCAACAGGTGGCATGAA | TTCATGCCACCTGTTGAGTAGTGA |
| NB57 | GCTAGGTCAATCTCCTTCGGAAGT | ACTTCCGAAGGAGATTGACCTAGC |
| NB58 | CAGGTTACTCCTCCGTGAGTCTGA | TCAGACTCACGGAGGAGTAACCTG |
| NB59 | TCAATCAAGAAGGGAAAGCAAGGT | ACCTTGCTTTCCCTTCTTGATTGA |
| NB60 | CATGTTCAACCAAGGCTTCTATGG | CCATAGAAGCCTTGGTTGAACATG |
| NB61 | AGAGGGTACTATGTGCCTCAGCAC | GTGCTGAGGCACATAGTACCCTCT |
| NB62 | CACCCACACTTACTTCAGGACGTA | TACGTCCTGAAGTAAGTGTGGGTG |
| NB63 | TTCTGAAGTTCCTGGGTCTTGAAC | GTTCAAGACCCAGGAACTTCAGAA |
| NB64 | GACAGACACCGTTCATCGACTTTC | GAAAGTCGATGAACGGTGTCTGTC |
| NB65 | TTCTCAGTCTTCCTCCAGACAAGG | CCTTGTCTGGAGGAAGACTGAGAA |
| NB66 | CCGATCCTTGTGGCTTCTAACTTC | GAAGTTAGAAGCCACAAGGATCGG |
| NB67 | GTTTGTCATACTCGTGTGCTCACC | GGTGAGCACACGAGTATGACAAAC |
| NB68 | GAATCTAAGCAAACACGAAGGTGG | CCACCTTCGTGTTTGCTTAGATTC |
| NB69 | TACAGTCCGAGCCTCATGTGATCT | AGATCACATGAGGCTCGGACTGTA |
| NB70 | ACCGAGATCCTACGAATGGAGTGT | ACACTCCATTCGTAGGATCTCGGT |
| NB71 | CCTGGGAGCATCAGGTAGTAACAG | CTGTTACTACCTGATGCTCCCAGG |
| NB72 | TAGCTGACTGTCTTCCATACCGAC | GTCGGTATGGAAGACAGTCAGCTA |
| NB73 | AAGAAACAGGATGACAGAACCCTC | GAGGGTTCTGTCATCCTGTTTCTT |
| NB74 | TACAAGCATCCCAACACTTCCACT | AGTGGAAGTGTTGGGATGCTTGTA |
| NB75 | GACCATTGTGATGAACCCTGTTGT | ACAACAGGGTTCATCACAATGGTC |
| NB76 | ATGCTTGTTACATCAACCCTGGAC | GTCCAGGGTTGATGTAACAAGCAT |
| NB77 | CGACCTGTTTCTCAGGGATACAAC | GTTGTATCCCTGAGAAACAGGTCG |
| NB78 | AACAACCGAACCTTTGAATCAGAA | TTCTGATTCAAAGGTTCGGTTGTT |
| NB79 | TCTCGGAGATAGTTCTCACTGCTG | CAGCAGTGAGAACTATCTCCGAGA |
| NB80 | CGGATGAACATAGGATAGCGATTC | GAATCGCTATCCTATGTTCATCCG |
| NB81 | CCTCATCTTGTGAAGTTGTTTCGG | CCGAAACAACTTCACAAGATGAGG |
| NB82 | ACGGTATGTCGAGTTCCAGGACTA | TAGTCCTGGAACTCGACATACCGT |
| NB83 | TGGCTTGATCTAGGTAAGGTCGAA | TTCGACCTTACCTAGATCAAGCCA |
| NB84 | GTAGTGGACCTAGAACCTGTGCCA | TGGCACAGGTTCTAGGTCCACTAC |
| NB85 | AACGGAGGAGTTAGTTGGATGATC | GATCATCCAACTAACTCCTCCGTT |
| NB86 | AGGTGATCCCAACAAGCGTAAGTA | TACTTACGCTTGTTGGGATCACCT |
| NB87 | TACATGCTCCTGTTGTTAGGGAGG | CCTCCCTAACAACAGGAGCATGTA |
| NB88 | TCTTCTACTACCGATCCGAAGCAG | CTGCTTCGGATCGGTAGTAGAAGA |
| NB89 | ACAGCATCAATGTTTGGCTAGTTG | CAACTAGCCAAACATTGATGCTGT |
| NB90 | GATGTAGAGGGTACGGTTTGAGGC | GCCTCAAACCGTACCCTCTACATC |
| NB91 | GGCTCCATAGGAACTCACGCTACT | AGTAGCGTGAGTTCCTATGGAGCC |
| NB92 | TTGTGAGTGGAAAGATACAGGACC | GGTCCTGTATCTTTCCACTCACAA |
| NB93 | AGTTTCCATCACTTCAGACTTGGG | CCCAAGTCTGAAGTGATGGAAACT |
| NB94 | GATTGTCCTCAAACTGCCACCTAC | GTAGGTGGCAGTTTGAGGACAATC |
| NB95 | CCTGTCTGGAAGAAGAATGGACTT | AAGTCCATTCTTCTTCCAGACAGG |
| NB96 | CTGAACGGTCATAGAGTCCACCAT | ATGGTGGACTCTATGACCGTTCAG |
We do not recommend mixing barcoded libraries with non-barcoded libraries prior to sequencing.
Compatibility of this protocol
This protocol should only be used in combination with:
- PCR Barcoding Expansion Pack 1-96 (EXP-PBC096)
- Native Barcoding Expansion Pack 1-12 and/or 13-24 (EXP-NBD104/EXP-NBD114)
- Ligation Sequencing Kit 1D (SQK-LSK109)
- FLO-MIN106 (R9.4.1) flow cells
- Flow Cell Wash Kit (EXP-WSH004)
2. Equipment and consumables
Material
- 100–200 fmol of each DNA sample to be barcoded in 45 µl
- PCR Barcoding Expansion 1-96 (EXP-PBC096)
- Native Barcoding Expansion 1-12 (EXP-NBD104) and 13-24 (EXP-NBD114) if multiplexing more than 12 samples
- Ligation Sequencing Kit (SQK-LSK109)
- Flow Cell Priming Kit (EXP-FLP002)
- Adapter Mix II Expansion (EXP-AMII001)
Instrumental
- Hula mixer (mezclador de rotación suave)
- Magnetic rack suitable for 96-well PCR plates, e.g. DynaMag™-96 Side Skirted Magnet (Thermo Fisher, cat # 12027)
- Gradilla magnética
- Centrifugadora de microplacas
- Microcentrífuga
- Mezclador vórtex
- Termociclador
- Pipeta y puntas P1000
- Pipeta y puntas P200
- Pipeta y puntas P100
- Pipeta y puntas P20
- Pipeta y puntas P10
- Pipeta y puntas P2
- Multichannel pipette and tips
- Cubeta con hielo
- Temporizador
Equipo opcional
- Bioanalizador Agilent (o equivalente)
- Fluorímetro Qubit™ (o equivalente para el control de calidad)
- Centrifugadora Eppendorf 5424 (o equivalente)
For this protocol, you will need the following amounts of each DNA sample to be barcoded in 45 µl:
- 1 µg (or 100-200 fmol) gDNA is required for R9.4.1 flow cells
- 1.5-3 µg (or 150-300 fmol) gDNA is required for R10.3 flow cells
Input DNA
How to QC your input DNA
It is important that the input DNA meets the quantity and quality requirements. Using too little or too much DNA, or DNA of poor quality (e.g. highly fragmented or containing RNA or chemical contaminants) can affect your library preparation.
For instructions on how to perform quality control of your DNA sample, please read the Input DNA/RNA QC protocol.
Chemical contaminants
Depending on how the DNA is extracted from the raw sample, certain chemical contaminants may remain in the purified DNA, which can affect library preparation efficiency and sequencing quality. Read more about contaminants on the Contaminants page of the Community.
PCR Barcoding Expansion Pack 1-96 (EXP-PBC096)
The kit allows up to 96 different libraries to be combined; each of these libraries will be barcoded with one of the 96 PCR barcodes (BC1-BC96). These libraries can then be barcoded for the second time with a native barcode from the Native Barcoding Expansions.
The reagents are divided between two 96 tube plates. One plate (blue caps) contains barcode adapters that are ligated to the DNA. All tubes in this plate have identical content. The other plate (white caps) contains the barcodes, one barcode per tube.

| Name | No. of plates | Fill volume per well (µl) | | --- | --- | --- | --- | | PCR Primer mix | 1 | 24 | | Barcode adapter plate | 1 | 240 |
Layout of barcodes in the 96 tube plate
The wells of the 96 tube plate correspond to the barcodes in the following way. All barcodes are supplied at 10 µM concentration and to be used at a final concentration of 0.2 µM.
Native Barcoding Expansion 1-12 (EXP-NBD104) and 13-24 (EXP-NBD114) contents
EXP-NBD104 kit contents
| Name | Acronym | Cap colour | No. of vials | Fill volume per vial (μl) |
|---|---|---|---|---|
| Native Barcode 01-12 | NB01-12 | White | 12 | 20 |
| Adapter Mix II | AMII | Green | 1 | 40 |
**EXP-NBD114 kit contents** 
| Name | Acronym | Cap colour | No. of vials | Fill volume per vial (μl) |
|---|---|---|---|---|
| Native Barcode 13-24 | NB13-24 | White | 12 | 20 |
| Adapter Mix II | AMII | Green | 1 | 40 |
Ligation Sequencing Kit contents (SQK-LSK109)
| Name | Acronym | Cap colour | No. of vials | Fill volume per vial (µl) |
|---|---|---|---|---|
| DNA CS | DCS | Yellow | 1 | 50 |
| Adapter Mix | AMX | Green | 1 | 40 |
| Ligation Buffer | LNB | Clear | 1 | 200 |
| L Fragment Buffer | LFB | White cap, orange stripe on label | 2 | 1,800 |
| S Fragment Buffer | SFB | Grey | 2 | 1,800 |
| Sequencing Buffer | SQB | Red | 2 | 300 |
| Elution Buffer | EB | Black | 1 | 200 |
| Loading Beads | LB | Pink | 1 | 360 |
Flow Cell Priming Kit contents (EXP-FLP002)

| Name | Acronym | Cap colour | No. of vials | Fill volume per vial (μl) |
|---|---|---|---|---|
| Flush Buffer | FB | Blue | 6 | 1,170 |
| Flush Tether | FLT | Purple | 1 | 200 |
Adapter Mix II Expansion contents (EXP-AMII001)
| Name | Acronym | Cap colour | No. of tubes | Fill volume per vial (μl) |
|---|---|---|---|---|
| Adapter Mix II | AMII | Green | 2 | 40 |
Adapter Mix II Expansion use
Protocols that use the Native Barcoding Expansions require 5 μl of AMII per reaction. Native Barcoding Expansions EXP-NBD104/NBD114 and EXP-NBD196 contain sufficient AMII for 6 and 12 reactions, respectively (or 12 and 24 reactions when sequencing on Flongle). This assumes that all barcodes are used in one sequencing run.
The Adapter Mix II expansion provides additional AMII for customers who are running subsets of barcodes, and allows a further 12 reactions (24 on Flongle).
3. Computer requirements and software
MinION Mk1B IT requirements
Sequencing on a MinION Mk1B requires a high-spec computer or laptop to keep up with the rate of data acquisition. For more information, refer to the MinION Mk1B IT requirements document.
Software for nanopore sequencing
MinKNOW
The MinKNOW software controls the nanopore sequencing device, collects sequencing data and basecalls in real time. You will be using MinKNOW for every sequencing experiment to sequence, basecall and demultiplex if your samples were barcoded.
For instructions on how to run the MinKNOW software, please refer to the MinKNOW protocol.
EPI2ME (optional)
The EPI2ME cloud-based platform performs further analysis of basecalled data, for example alignment to the Lambda genome, barcoding, or taxonomic classification. Currently, the FASTQ Barcoding workflow in EPI2ME is not compatible with dual-barcoded reads. EPI2ME installation and use For instructions on how to create an EPI2ME account and install the EPI2ME Desktop Agent, please refer to the EPI2ME Platform protocol.
Check your flow cell
We highly recommend that you check the number of pores in your flow cell prior to starting a sequencing experiment. This should be done within 12 weeks of purchasing for MinION/GridION/PromethION or within four weeks of purchasing Flongle Flow Cells. Oxford Nanopore Technologies will replace any flow cell with fewer than the number of pores in the table below, when the result is reported within two days of performing the flow cell check, and when the storage recommendations have been followed. To do the flow cell check, please follow the instructions in the Flow Cell Check document.
| Flow cell | Minimum number of active pores covered by warranty |
|---|---|
| Flongle Flow Cell | 50 |
| MinION/GridION Flow Cell | 800 |
| PromethION Flow Cell | 5000 |
4. End-prep
Material
- 100–200 fmol of each DNA sample to be barcoded in 45 µl
Instrumental
- Termociclador
- Cubeta con hielo
- Centrifuge capable of taking 96-well plates
- Magnetic separation rack, suitable for 1.5 ml Eppendorf tubes
Equipo opcional
- Fluorímetro Qubit™ (o equivalente para el control de calidad)
Optional fragmentation and size selection
By default, the protocol contains no DNA fragmentation step, however in some cases it may be advantageous to fragment your sample. For example, when working with lower amounts of input gDNA (100 ng – 500 ng), fragmentation will increase the number of DNA molecules and therefore increase throughput. Instructions are available in the DNA Fragmentation section of Extraction methods.
Additionally, we offer several options for size-selecting your DNA sample to enrich for long fragments - instructions are available in the Size Selection section of Extraction methods.
Prepare the DNA in nuclease-free water.
- Transfer 100–200 fmol DNA for each sample to be barcoded into a separate 1.5 ml Eppendorf DNA LoBind tube
- Adjust the volume to 45 μl with nuclease-free water
- Mix thoroughly by flicking the tube to avoid unwanted shearing
- Spin down briefly in a microfuge
In a 0.2 ml 96 well PCR plate, set up the end-repair / dA-tailing reactions as follows:
| Reagent | Volume |
|---|---|
| DNA sample | 45 µl |
| Ultra II End-prep reaction buffer | 7 µl |
| Ultra II End-prep enzyme mix | 3 µl |
| Nuclease-free water | 5 µl |
| Total | 60 µl |
Mix by pipetting.
Seal the plate with adhesive film or PCR strip caps, spin down in a centrifuge and incubate for 5 minutes at 20 °C and 5 minutes at 65 °C using the thermal cycler.
If condensation is observed in the plate after the thermocycling, briefly spin down the plate contents in a centrifuge.
Resuspend the AMPure XP beads by vortexing.
Add 60 µl of resuspended AMPure XP beads to the end-prep reaction and mix by pipetting.
Allow DNA to bind to beads for 5 minutes at room temperature.
Prepare sufficient fresh 70% ethanol in nuclease-free water.
Place on a magnetic rack, allow beads to pellet and pipette off supernatant.
Keep the tube on the magnet and wash the beads with 180 µl of freshly-prepared 70% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
Repeat the previous step.
Cover the plate with adhesive film and leave plate on magnet for 2 minutes to allow residual liquid to collect at the bottom. Remove the adhesive film, return the plate to the magnet and aspirate residual wash solution.
Briefly incubate the plate on a thermal cycler at 37° C with the lid open and the plate wells unsealed.
Remove the plate from the magnet and resuspend pellet in 31 µl nuclease-free water. Incubate for 2 minutes at room temperature.
Pellet the beads on a magnet until the eluate is clear and colourless.
Remove eluate once it is clear and colourless. Transfer each eluted sample to a new 96-well PCR plate.
Quantify 1 µl of end-prepped DNA using a Qubit fluorometer - recovery aim 70–140 fmol.
Take forward approximately 70–140 fmol of end-prepped DNA in 30 µl nuclease-free water into adapter ligation.
5. Ligation of Barcode Adapter
Material
- Barcode Adapter (BCA)
Consumibles
- NEB Blunt/TA Ligase Master Mix (NEB, cat # M0367)
- Agua sin nucleasas (p.ej. Thermo Scientific, AM9937)
- Etanol al 70 % recién preparado en agua sin nucleasas
- 10 mM Tris-HCl pH 8.5
- 0.2 ml 96-well PCR plate
Instrumental
- Centrifugadora de microplacas
- Hula mixer (mezclador de rotación suave)
- Mezclador vórtex
- Magnetic rack suitable for 96-well PCR plates, e.g. DynaMag™-96 Side Skirted Magnet (Thermo Fisher, cat # 12027)
- Cubeta con hielo
- Multichannel pipette and tips
Equipo opcional
- Bioanalizador Agilent (o equivalente)
Add the reagents to a fresh 96-well plate, in the order given below:
Between each addition, pipette mix 10-20 times.
| Reagent | Volume |
|---|---|
| End-prepped DNA | 30 µl |
| Barcode Adapter | 20 µl |
| Blunt/TA Ligase Master Mix | 50 µl |
| Total | 100 µl |
Mix by pipetting.
Seal the plate with adhesive film or PCR strip caps and briefly spin down in a plate spinner.
Incubate the reaction for 10 minutes at room temperature.
Resuspend the AMPure XP beads by vortexing.
Add 40 µl of resuspended AMPure XP beads to each sample and mix by pipetting up and down ten times.
Allow DNA to bind to beads for 5 minutes at room temperature.
Prepare sufficient fresh 70% ethanol in nuclease-free water.
Place on a magnetic rack, allow beads to pellet and pipette off supernatant.
Keep the tube on the magnet and wash the beads with 180 µl of freshly-prepared 70% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
Repeat the previous step.
Cover the plate with adhesive film and leave plate on magnet for 2 minutes to allow residual liquid to collect at the bottom. Remove the adhesive film, return the plate to the magnet and aspirate residual wash solution.
Briefly incubate the plate on a thermal cycler at 37° C with the lid open and the plate wells unsealed.
Remove the plate from the magnet and resuspend pellet in 25 µl nuclease-free water. Incubate for 2 minutes at room temperature.
Pellet the beads on a magnet until the eluate is clear and colourless.
Remove eluate once it is clear and colourless. Transfer each eluted sample to a new 96-well PCR plate.
Quantify 1 µl of end-prepped DNA using a Qubit fluorometer.
Dilute the library to 20–30 fmol with nuclease-free water or 10 mM Tris-HCl pH 8.5.
6. Barcoding PCR
Material
- PCR Barcodes (BC01-96, at 10 µM)
Consumibles
- LongAmp Taq 2X Master Mix (e.g. NEB, cat # M0287)
- Microesferas Agencourt AMPure XP (Beckman Coulter™, A63881)
- Etanol al 70 % recién preparado en agua sin nucleasas
- Agua sin nucleasas (p.ej. Thermo Scientific, AM9937)
- 0.2 ml 96-well PCR plate
- Tubos Eppendorf DNA LoBind de 1,5 ml
Instrumental
- Termociclador
- Centrifugadora de microplacas
- Magnetic rack suitable for 96-well PCR plates, e.g. DynaMag™-96 Side Skirted Magnet (Thermo Fisher, cat # 12027)
Equipo opcional
- Fluorímetro Qubit™ (o equivalente para el control de calidad)
Layout of barcodes in the 96 tube plate
The wells of the 96 tube plate correspond to the barcodes in the following way. All barcodes are supplied at 10 µM concentration and to be used at a final concentration of 0.2 µM.
Set up a barcoding PCR reaction as follows for each library:
The following is written for LongAmp Taq, but can be adapted for use with other polymerases.
Between each addition, pipette mix 10-20 times.
| Reagent | Volume |
|---|---|
| PCR Barcode (one of BC1-BC96, at 10 µM) | 1 µl |
| Adapter-ligated DNA | 2 µl |
| LongAmp Taq 2x master mix | 25 µl |
| Nuclease-free water | 22 µl |
| Total volume | 50 µl |
The amount of input DNA may need to be adjusted depending on application. For example, for sequencing human or larger genomes, we recommend putting ~50 ng DNA into a PCR reaction. For amplicons or smaller genomes, the 20 ng stated above is sufficient. If the amount of input material is altered, the number of PCR cycles may need to be adjusted to produce the same yield.
Mix by pipetting.
Seal the plate with adhesive film or PCR strip caps and briefly spin down in a plate spinner.
Amplify using the following cycling conditions:
| Cycle step | Temperature | Time | No. of cycles |
|---|---|---|---|
| Initial denaturation | 95 °C | 3 mins | 1 |
| Denaturation | 95 °C | 15 secs | 15-18 (b) |
| Annealing | 62 °C (a) | 15 secs (a) | 15-18 (b) |
| Extension | 65 °C (c) | dependent on length of target fragment (d) | 15-18 (b) |
| Final extension | 65 °C | dependent on length of target fragment (d) | 1 |
| Hold | 4 °C | ∞ |
a. This is specific to the Oxford Nanopore primer and should be maintained
b. Adjust accordingly if input quantities are altered
c. This temperature is determined by the type of polymerase that is being used (given here for LongAmp Taq polymerase)
d. Adjust accordingly for different lengths of amplicons and the type of polymerase that is being used (LongAmp Taq amplifies at a rate of 50 seconds per kb)
Resuspend the AMPure XP beads by vortexing.
Add 35 µl of of resuspended AMPure XP beads to each sample and mix by pipetting the entire combined volume up and down 10 times.
Incubate for 5 minutes at room temperature.
Prepare sufficient fresh 70% ethanol in nuclease-free water.
Pellet the beads on a magnet for at least 2 min, or until the supernatant is clear. Keep the plate on the magnet and pipette off the supernatant.
Wash each pellet of beads by adding 200 µl of freshly-prepared 70% ethanol. Resuspend each pellet thoroughly by pipetting the entire volume of buffer up and down ten times. Return the plate to the magnetic rack and allow the beads to pellet until the supernatant is clear. Remove the supernatant using a pipette and discard.
Repeat the previous step.
Seal the plate. Spin down and place the plate back on the magnet. Pipette off any residual supernatant.
Remove the plate from the magnetic rack and resuspend each pellet in 21 µl nuclease-free water, pipetting the entire volume up and down 10 times. Incubate for 2 minutes at room temperature.
Pellet the beads on a magnet until the eluate is clear and colourless.
Remove and retain 21 µl of each eluate into a well of a clean 96-well plate.
Quantify 1 µl of each barcoded DNA sample using a Qubit fluorometer.
If your DNA samples are of mixed sizes, we recommend either running a subset of samples on an Agilent Bioanalyzer, or estimating the DNA size from a gel.
Using the DNA mass (calculated using the Qubit fluorometer) and size distribution (calculated using a gel or Agilent Bioanalyzer), pool equimolar quantities of barcoded amplicons in batches of 96, ensuring that every sample within a given pool has a unique barcode.
Using the DNA mass (calculated using the Qubit fluorometer) and size distribution (calculated using a gel or Agilent Bioanalyzer), pool equimolar quantities of barcoded amplicons in batches of 96 or fewer, depending on how many of the 96 barcodes were used. Ensure that every sample within a given pool has a unique barcode.
Note that after the subsequent end-prep step, you will need 100–200 fmol of DNA for each pool of samples to take into the native barcode ligation step.
7. End-prep
Material
- Up to 24 pools of barcoded amplicons (with up to 96 samples in each pool), in 20 µl
Instrumental
- Termociclador
- Cubeta con hielo
- Centrifuge capable of taking 96-well plates
- Magnetic separation rack, suitable for 1.5 ml Eppendorf tubes
Equipo opcional
- Fluorímetro Qubit™ (o equivalente para el control de calidad)
Mix the following reagents in a separate 0.2 ml thin-walled PCR tube for each pool of samples:
| Reagent | Volume |
|---|---|
| Pool of barcoded DNA samples | 20 µl |
| Ultra II End-prep reaction buffer | 7 µl |
| Ultra II End-prep enzyme mix | 3 µl |
| Nuclease-free water | 30 µl |
| Total | 60 µl |
Mix by pipetting.
Using a thermal cycler, incubate at 20° C for 15 minutes and 65° C for 5 mins.
Transfer each sample to a separate 1.5 ml Eppendorf DNA LoBind tube.
Resuspend the AMPure XP beads by vortexing.
Add 60 µl of resuspended AMPure XP beads to the end-prep reaction and mix by pipetting.
Allow DNA to bind to beads for 5 minutes at room temperature.
Prepare sufficient fresh 70% ethanol in nuclease-free water.
Keep the tube on the magnet and wash the beads with 180 µl of freshly-prepared 70% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
Keep the tube on the magnet and wash the beads with 200 µl of freshly prepared 70% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
Repeat the previous step.
Spin down and place the tube back on the magnet. Pipette off any residual ethanol. Allow to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
Remove the tube from the magnetic rack and resuspend the pellet in 23.5 µl nuclease-free water. Incubate for 2 minutes at room temperature.
Pellet the beads on a magnet until the eluate is clear and colourless.
Remove and retain 23.5 µl of eluate into a clean 1.5 ml Eppendorf DNA LoBind tube.
Quantify 1 µl of end-prepped DNA using a Qubit fluorometer.
Take forward 100–200 fmol of end-prepped DNA in 22.5 µl into native barcode ligation.
8. Native barcode ligation
Material
- Native Barcoding Expansion 1-12 (EXP-NBD104) and 13-24 (EXP-NBD114) if multiplexing more than 12 samples
Consumibles
- Etanol al 70 % recién preparado en agua sin nucleasas
- Tubos Eppendorf DNA LoBind de 1,5 ml
- Agua sin nucleasas (p.ej. Thermo Scientific, AM9937)
- NEB Blunt/TA Ligase Master Mix (NEB, M0367)
Instrumental
- Magnetic separation rack, suitable for 1.5 ml Eppendorf tubes
- Hula mixer (mezclador de rotación suave)
- Mezclador vórtex
- Cubeta con hielo
- Microcentrífuga
- Pipeta y puntas P1000
- Pipeta y puntas P100
- Pipeta y puntas P10
Equipo opcional
- Fluorímetro Qubit™ (o equivalente para el control de calidad)
Thaw the native barcodes at room temperature. Use one barcode per sample. Individually mix the barcodes by pipetting, spin down, and place them on ice.
Select a unique barcode for every sample to be run together on the same flow cell, from the provided 24 barcodes. Up to 24 samples can be barcoded and combined in one experiment.
Add the reagents in the order given below, mixing by flicking the tube between each sequential addition:
| Reagent | Volume |
|---|---|
| End-prepped DNA | 22.5 µl |
| Native Barcode | 2.5 µl |
| Blunt/TA Ligase Master Mix | 25 µl |
| Total | 50 µl |
Mix well by pipetting and spin down.
Incubate the reaction for 10 minutes at room temperature.
Resuspend the AMPure XP beads by vortexing.
Add 20 µl of resuspended AMPure XP beads to the reaction and mix by gently flicking the tube.
Incubate on a Hula mixer (rotator mixer) for 5 minutes at room temperature.
Prepare sufficient fresh 70% ethanol in nuclease-free water.
Spin down the sample and pellet on a magnet. Keep the tube on the magnet, and pipette off the supernatant when clear and colourless.
Keep the tube on the magnet and wash the beads with 200 µl of freshly prepared 70% ethanol without disturbing the pellet. Remove the ethanol using a pipette and discard.
Repeat the previous step.
Spin down and place the tube back on the magnet. Pipette off any residual ethanol. Allow to dry for ~30 seconds, but do not dry the pellet to the point of cracking.
Remove the tube from the magnetic rack and resuspend pellet in 21 µl nuclease-free water. Incubate for 2 minutes at room temperature.
Pellet the beads on a magnet until the eluate is clear and colourless.
Remove and retain 21 µl of eluate into a clean 1.5 ml Eppendorf DNA LoBind tube.
Quantify 1 µl of eluted sample using a Qubit fluorometer.
Analyse 1 µl of sample using the Agilent Bioanalyzer. Determine the average amplicon size from this data, and use this to calculate the input sample volume for the next step.
Pool equimolar amounts of each barcoded sample into a 1.5 ml Eppendord DNA LoBind tube, ensuring that sufficient sample is combined to produce a pooled sample of 0.2 pmol total.
Quantify 1 µl of pooled and barcoded DNA using a Qubit fluorometer.
Dilute 100–200 fmol pooled sample to 65 µl in nuclease-free water.
9. Adapter ligation and clean-up
Material
- Long Fragment Buffer (LFB)
- Short Fragment Buffer (SFB)
- Elution Buffer (EB)
- Adapter Mix II (AMII)
Instrumental
- Microcentrífuga
- Gradilla magnética
- Mezclador vórtex
- Hula mixer (mezclador de rotación suave)
Equipo opcional
- Fluorímetro Qubit™ (o equivalente para el control de calidad)
Adapter Mix II Expansion use
Protocols that use the Native Barcoding Expansions require 5 μl of AMII per reaction. Native Barcoding Expansions EXP-NBD104/NBD114 and EXP-NBD196 contain sufficient AMII for 6 and 12 reactions, respectively (or 12 and 24 reactions when sequencing on Flongle). This assumes that all barcodes are used in one sequencing run.
The Adapter Mix II expansion provides additional AMII for customers who are running subsets of barcodes, and allows a further 12 reactions (24 on Flongle).
Depending on the wash buffer (LFB or SFB) used, the clean-up step after adapter ligation is designed to either enrich for DNA fragments of >3 kb, or purify all fragments equally.
- To enrich for DNA fragments of 3 kb or longer, use Long Fragment Buffer (LFB)
- To retain DNA fragments of all sizes, use Short Fragment Buffer (SFB)
Thaw the Elution Buffer (EB) and NEBNext Quick Ligation Reaction Buffer (5x) at room temperature, mix by vortexing, spin down and place on ice. Check the contents of each tube are clear of any precipitate.
Spin down the T4 Ligase and the Adapter Mix II (AMII), and place on ice.
To enrich for DNA fragments of 3 kb or longer, thaw one tube of Long Fragment Buffer (LFB) at room temperature, mix by vortexing, spin down and place on ice.
To retain DNA fragments of all sizes, thaw one tube of Short Fragment Buffer (SFB) at room temperature, mix by vortexing, spin down and place on ice.
Taking the pooled and barcoded DNA, perform adapter ligation as follows, mixing by flicking the tube between each sequential addition.
| Reagent | Volume |
|---|---|
| 100–200 fmol pooled barcoded sample | 65 µl |
| Adapter Mix II (AMII) | 5 µl |
| NEBNext Quick Ligation Reaction Buffer (5X) | 20 µl |
| Quick T4 DNA Ligase | 10 µl |
| Total | 100 µl |
Ensure the components are thoroughly mixed by pipetting, and spin down.
Incubate the reaction for 10 minutes at room temperature.
Resuspend the AMPure XP beads by vortexing.
Add 50 µl of resuspended AMPure XP beads to the reaction and mix by pipetting.
Incubate on a Hula mixer (rotator mixer) for 5 minutes at room temperature.
Place on a magnetic rack, allow beads to pellet and pipette off supernatant.
Wash the beads by adding either 250 μl Long Fragment Buffer (LFB) or 250 μl Short Fragment Buffer (SFB). Flick the beads to resuspend, spin down, then return the tube to the magnetic rack and allow the beads to pellet. Remove the supernatant using a pipette and discard.
Repeat the previous step.
Spin down and place the tube back on the magnet. Pipette off any residual supernatant.
Remove the tube from the magnetic rack and resuspend the pellet in 15 µl Elution Buffer (EB). Spin down and incubate for 10 minutes at room temperature. For high molecular weight DNA, incubating at 37°C can improve the recovery of long fragments.
Pellet the beads on a magnet until the eluate is clear and colourless, for at least 1 minute.
Remove and retain 15 µl of eluate containing the DNA library into a clean 1.5 ml Eppendorf DNA LoBind tube.
Dispose of the pelleted beads
Quantify 1 µl of adapter ligated DNA using a Qubit fluorometer - recovery aim 50–100 fmol.
We recommend loading 5-50 fmol of the final prepared library onto a flow cell.
Loading more than the maximal recommended amount of DNA can have a detrimental effect on output as higher quantities of DNA results in a larger number of ligated DNA ends with loaded motor protein. This depletes fuel in the Sequencing Buffer, regardless of whether or not the DNA fragments are being sequenced. This leads to fuel depletion and speed drop-off early in the sequencing run. Dilute the libraries in Elution Buffer if required.
If you are using the Flongle for sample prep development, we recommend loading 3-20 fmol instead.
The prepared library is used for loading onto the flow cell. Store the library on ice or at 4°C until ready to load.
Library storage recommendations
We recommend storing libraries in Eppendorf DNA LoBind tubes at 4°C for short term storage or repeated use, for example, reloading flow cells between washes. For single use and long-term storage of more than 3 months, we recommend storing libraries at -80°C in Eppendorf DNA LoBind tubes. For further information, please refer to the DNA library stability Know-How document.
If quantities allow, the library may be diluted in Elution Buffer (EB) for splitting across multiple flow cells.
Additional buffer for doing this can be found in the Sequencing Auxiliary Vials expansion (EXP-AUX001), available to purchase separately. This expansion also contains additional vials of Sequencing Buffer (SQB) and Loading Beads (LB), required for loading the libraries onto flow cells.
10. Priming and loading the SpotON flow cell
Material
- Flow Cell Priming Kit (EXP-FLP002)
- Loading Beads (LB)
- Sequencing Buffer (SQB)
Consumibles
- Tubos Eppendorf DNA LoBind de 1,5 ml
- Agua sin nucleasas (p.ej. Thermo Scientific, AM9937)
Instrumental
- MinION Mk1B or Mk1C
- SpotON Flow Cell
- MinION/GridION Flow Cell Light Shield
- Pipeta y puntas P1000
- Pipeta y puntas P100
- Pipeta y puntas P20
- Pipeta y puntas P10
Priming and loading a flow cell
We recommend all new users watch the 'Priming and loading your flow cell' video before your first run.
Thaw the Sequencing Buffer (SQB), Loading Beads (LB), Flush Tether (FLT) and one tube of Flush Buffer (FB) at room temperature before mixing the reagents by vortexing, and spin down at room temperature.
To prepare the flow cell priming mix, add 30 µl of thawed and mixed Flush Tether (FLT) directly to the tube of thawed and mixed Flush Buffer (FB), and mix by vortexing at room temperature.
Open the MinION device lid and slide the flow cell under the clip.
Press down firmly on the flow cell to ensure correct thermal and electrical contact.
Complete a flow cell check to assess the number of pores available before loading the library.
This step can be omitted if the flow cell has been checked previously.
See the flow cell check instructions in the MinKNOW protocol for more information.
Slide the flow cell priming port cover clockwise to open the priming port.
Take care when drawing back buffer from the flow cell. Do not remove more than 20-30 µl, and make sure that the array of pores are covered by buffer at all times. Introducing air bubbles into the array can irreversibly damage pores.
After opening the priming port, check for a small air bubble under the cover. Draw back a small volume to remove any bubbles:
- Set a P1000 pipette to 200 µl
- Insert the tip into the priming port
- Turn the wheel until the dial shows 220-230 µl, to draw back 20-30 µl, or until you can see a small volume of buffer entering the pipette tip
Note: Visually check that there is continuous buffer from the priming port across the sensor array.
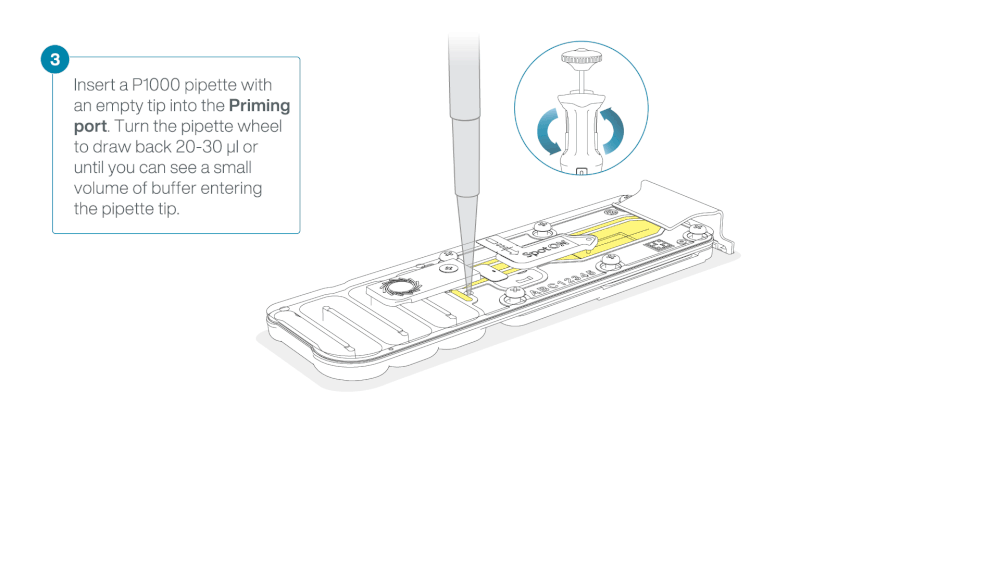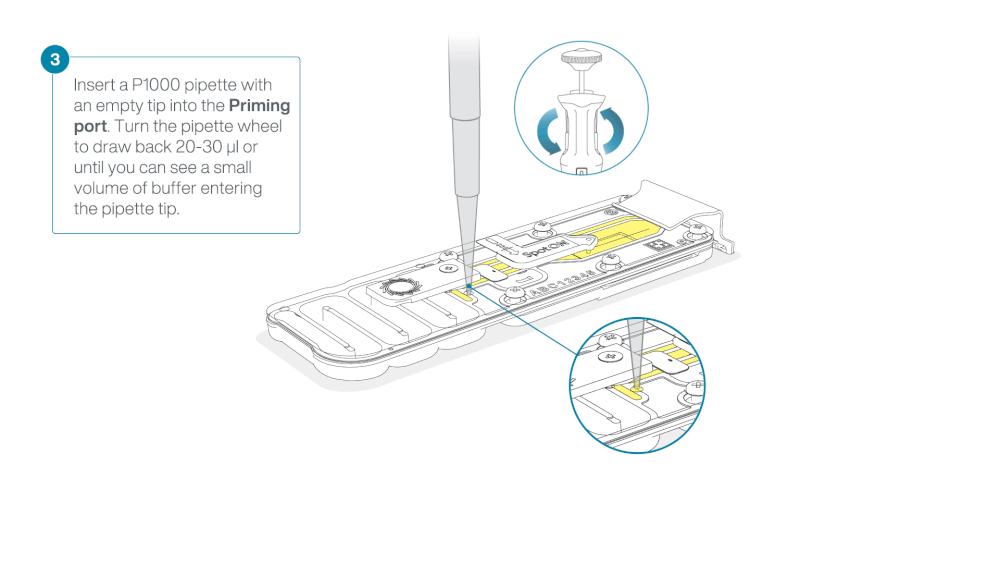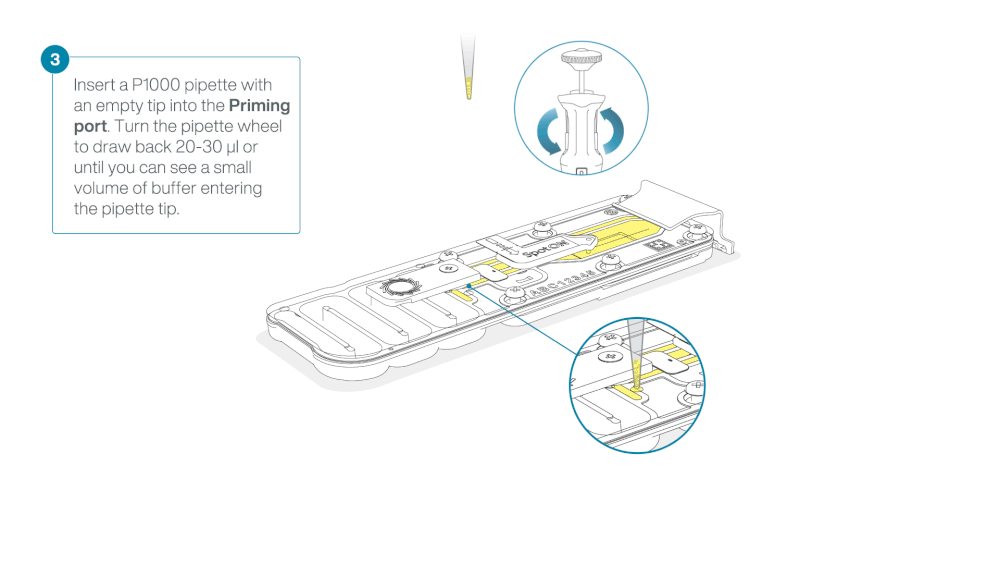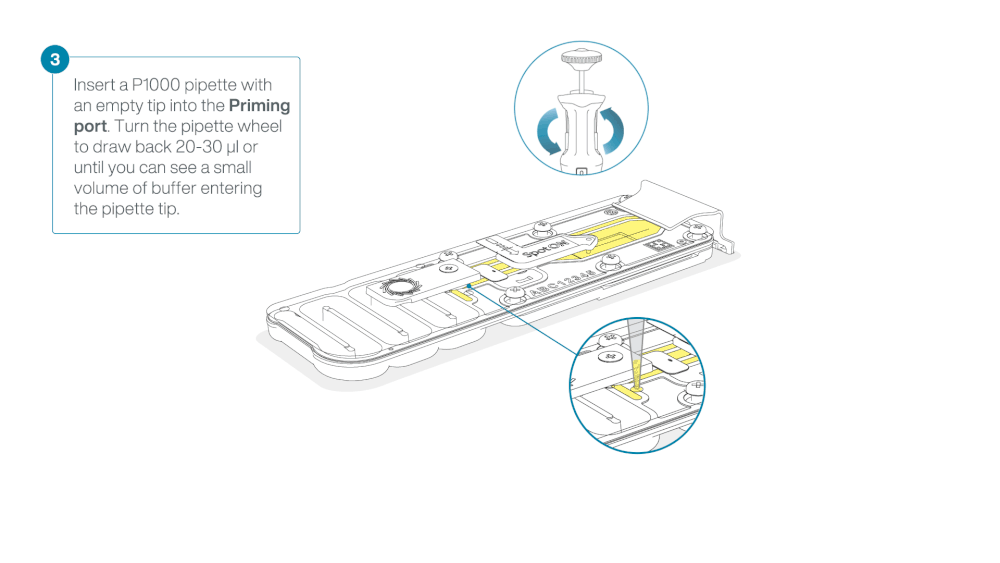
Load 800 µl of the priming mix into the flow cell via the priming port, avoiding the introduction of air bubbles. Wait for five minutes. During this time, prepare the library for loading by following the steps below.
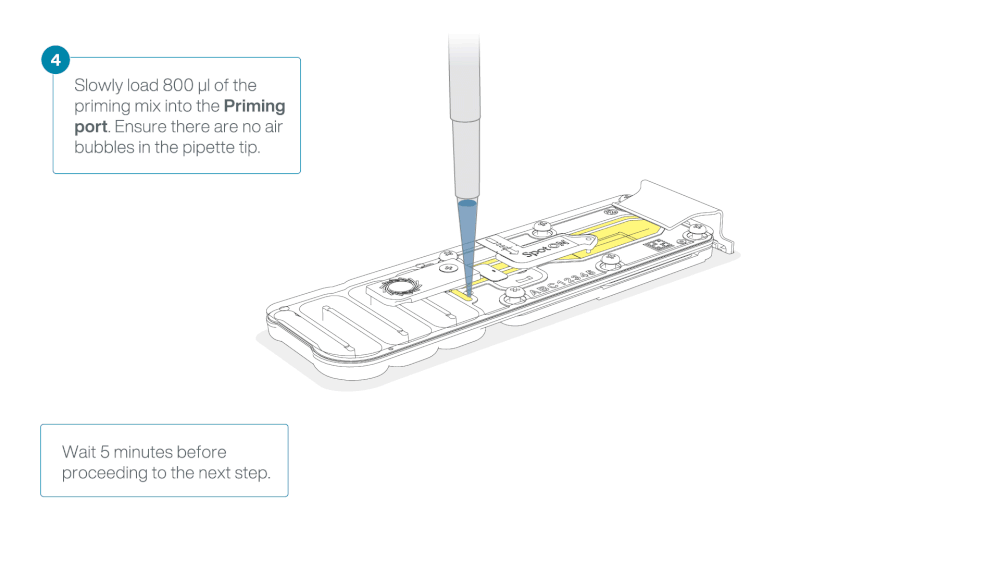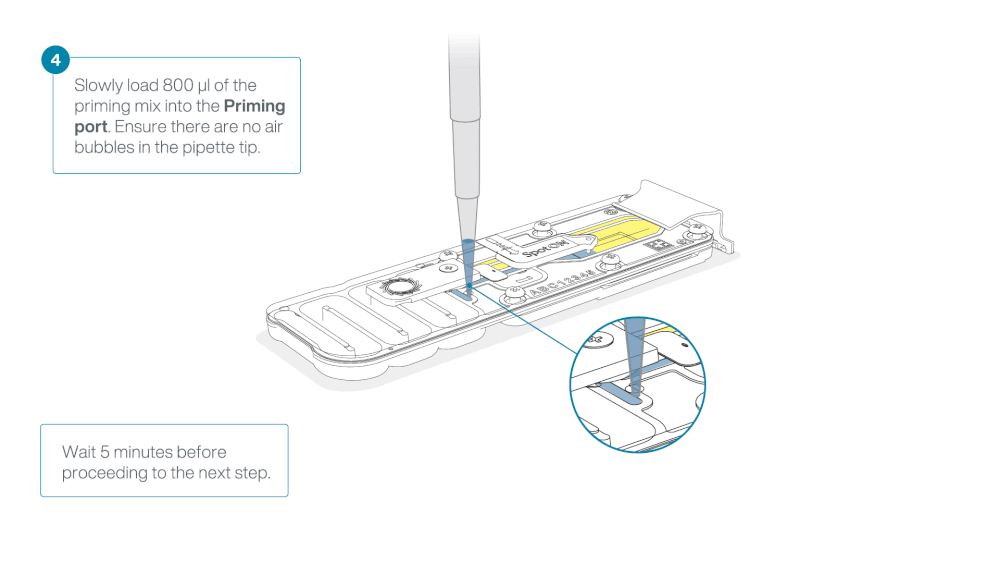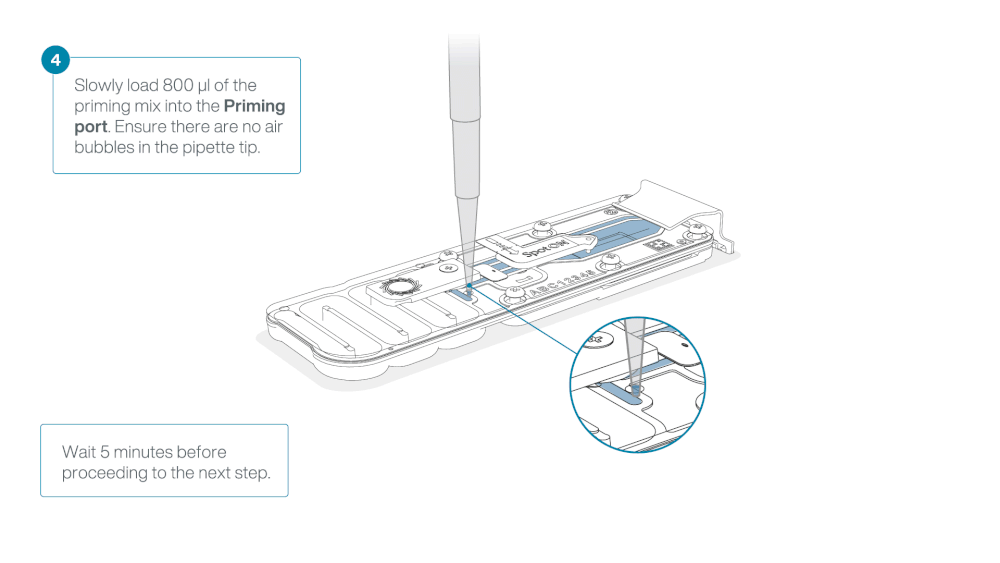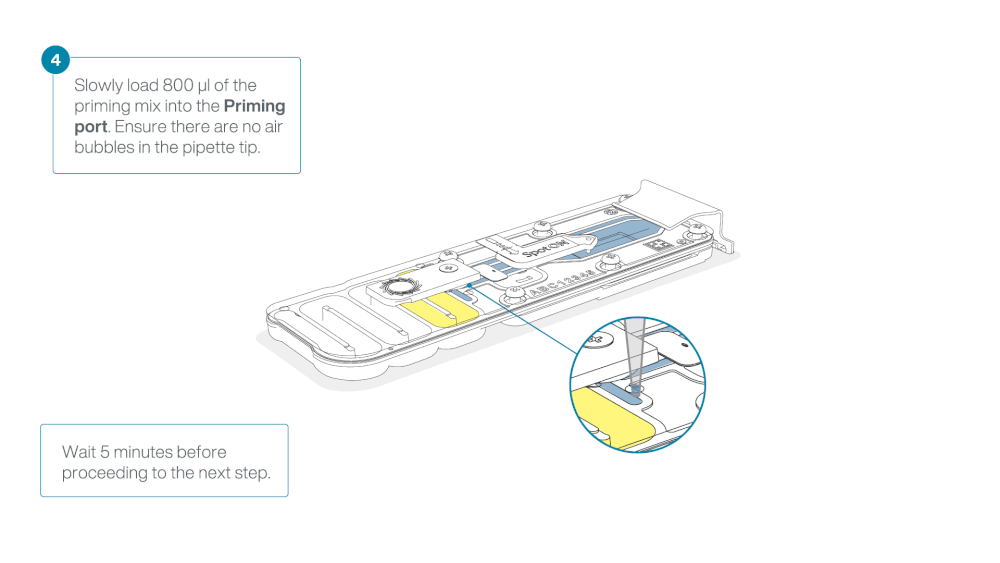
Thoroughly mix the contents of the Loading Beads (LB) by pipetting.
The Loading Beads (LB) tube contains a suspension of beads. These beads settle very quickly. It is vital that they are mixed immediately before use.
In a new tube, prepare the library for loading as follows:
| Reagent | Volume per flow cell |
|---|---|
| Sequencing Buffer (SQB) | 37.5 µl |
| Loading Beads (LB), mixed immediately before use | 25.5 µl |
| DNA library | 12 µl |
| Total | 75 µl |
Note: Load the library onto the flow cell immediately after adding the Sequencing Buffer (SQB) and Loading Beads (LB) because the fuel in the buffer will start to be consumed by the adapter.
Complete the flow cell priming:
- Gently lift the SpotON sample port cover to make the SpotON sample port accessible.
- Load 200 µl of the priming mix into the flow cell priming port (not the SpotON sample port), avoiding the introduction of air bubbles.

Mix the prepared library gently by pipetting up and down just prior to loading.
Add 75 μl of the prepared library to the flow cell via the SpotON sample port in a dropwise fashion. Ensure each drop flows into the port before adding the next.

Gently replace the SpotON sample port cover, making sure the bung enters the SpotON port and close the priming port.

Install the light shield on your flow cell as soon as library has been loaded for optimal sequencing output.
We recommend leaving the light shield on the flow cell when library is loaded, including during any washing and reloading steps. The shield can be removed when the library has been removed from the flow cell.
Place the light shield onto the flow cell, as follows:
Carefully place the leading edge of the light shield against the clip. Note: Do not force the light shield underneath the clip.
Gently lower the light shield onto the flow cell. The light shield should sit around the SpotON cover, covering the entire top section of the flow cell.

The MinION Flow Cell Light Shield is not secured to the flow cell and careful handling is required after installation.
Close the device lid and set up a sequencing run on MinKNOW.
11. Data acquisition and basecalling
Overview of nanopore data analysis
For a full overview of nanopore data analysis, which includes options for basecalling and post-basecalling analysis, please refer to the Data Analysis document.
How to start sequencing
The sequencing device control, data acquisition and real-time basecalling are carried out by the MinKNOW software. Please ensure MinKNOW is installed on your computer or device. There are multiple options for how to carry out sequencing:
1. Data acquisition and basecalling in real-time using MinKNOW on a computer
Follow the instructions in the MinKNOW protocol beginning from the "Starting a sequencing run" section until the end of the "Completing a MinKNOW run" section.
2. Data acquisition and basecalling in real-time using the MinION Mk1B/Mk1D device
Follow the instructions in the MinION Mk1B user manual or the MinION Mk1D user manual.
3. Data acquisition and basecalling in real-time using the MinION Mk1C device
Follow the instructions in the MinION Mk1C user manual.
4. Data acquisition and basecalling in real-time using the GridION device
Follow the instructions in the GridION user manual.
5. Data acquisition and basecalling in real-time using the PromethION device
Follow the instructions in the PromethION user manual or the PromethION 2 Solo user manual.
6. Data acquisition using MinKNOW on a computer and basecalling at a later time using MinKNOW
Follow the instructions in the MinKNOW protocol beginning from the "Starting a sequencing run" section until the end of the "Completing a MinKNOW run" section. When setting your experiment parameters, set the Basecalling tab to OFF. After the sequencing experiment has completed, follow the instructions in the Post-run analysis section of the MinKNOW protocol.
Settings in MinKNOW
When setting up your sequencing experiment in MinKNOW, select the SQK-LSK109 kit only (no Barcoding Expansion Packs) in Kit Selection: 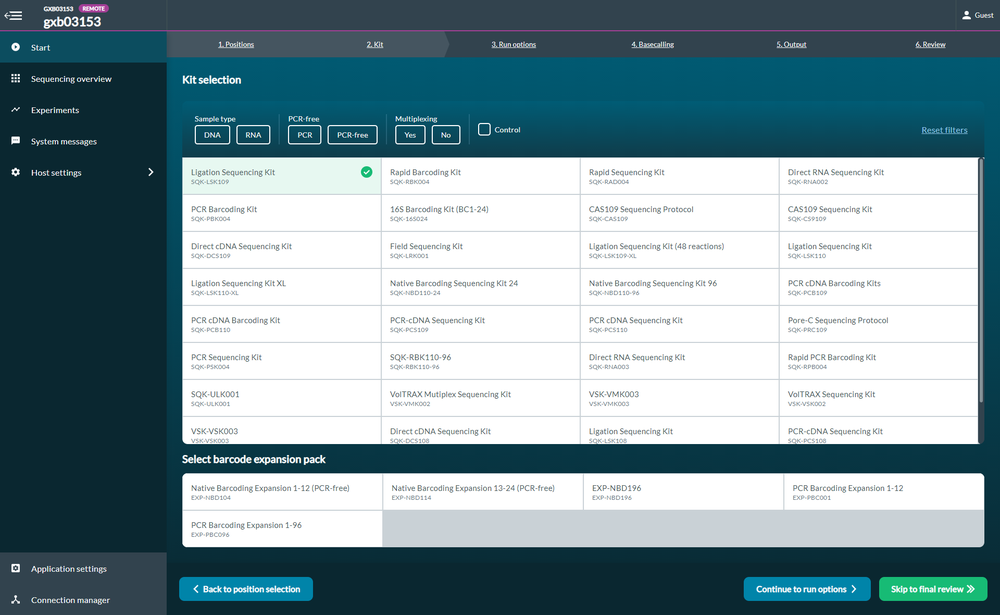
12. Downstream analysis
Barcode demultiplexing in Guppy
To demultiplex your reads by barcode, use the dual barcoding configuration file, specifying the barcode kit as "EXP-DUAL00":
guppy_barcoder --input_path <folder containing FASTQ and/or FASTA files> --save_path <output folder> --config configuration_dual.cfg --barcode_kits "EXP-DUAL00"
For more information and instructions on how to use Guppy, please refer to the relevant sections of the protocol: Barcode demultiplexing Quick Start
Other data analysis options
1. EPI2ME platform
The EPI2ME platform is a cloud-based data analysis service developed by Metrichor Ltd., a subsidiary of Oxford Nanopore Technologies. The EPI2ME platform offers a range of analysis workflows, e.g. for metagenomic identification, barcoding, alignment, and structural variant calling. The analysis requires no additional equipment or compute power, and provides an easy-to-interpret report with the results. For instructions on how to run an analysis workflow in EPI2ME, please follow the instructions in the EPI2ME protocol, beginning at the "Starting an EPI2ME workflow" step. Please note that the FASTQ Barcoding workflow in EPI2ME is currently not compatible with dual-barcoded reads.
2. Bioinformatics tutorials
For more in-depth data analysis, Oxford Nanopore Technologies offers a range of bioinformatics tutorials, which are available in the Bioinformatics resource section of the Community. The tutorials take the user through installing and running pre-built analysis pipelines, which generate a report with the results. The tutorials are aimed at biologists who would like to analyse data without the help of a dedicated bioinformatician, and who are comfortable using the command line.
3. Research analysis tools
Oxford Nanopore Technologies' Research division has created a number of analysis tools, which are available in the Oxford Nanopore GitHub repository. The tools are aimed at advanced users, and contain instructions for how to install and run the software. They are provided as-is, with minimal support.
4. Community-developed analysis tools
If a data analysis method for your research question is not provided in any of the resources above, please refer to the Community-developed data analysis tool library. Numerous members of the Nanopore Community have developed their own tools and pipelines for analysing nanopore sequencing data, most of which are available on GitHub. Please be aware that these tools are not supported by Oxford Nanopore Technologies, and are not guaranteed to be compatible with the latest chemistry/software configuration.
13. Flow cell reuse and returns
Material
- Kit Flow Cell Wash (EXP-WSH004)
After your sequencing experiment is complete, if you would like to reuse the flow cell, please follow the Flow Cell Wash Kit protocol and store the washed flow cell at +2°C to +8°C.
The Flow Cell Wash Kit protocol is available on the Nanopore Community.
We recommend you to wash the flow cell as soon as possible after you stop the run. However, if this is not possible, leave the flow cell on the device and wash it the next day.
Alternatively, follow the returns procedure to send the flow cell back to Oxford Nanopore.
Instructions for returning flow cells can be found here.
If you encounter issues or have questions about your sequencing experiment, please refer to the Troubleshooting Guide that can be found in the online version of this protocol.
14. Issues during DNA/RNA extraction and library preparation
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Low sample quality
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Low DNA purity (Nanodrop reading for DNA OD 260/280 is <1.8 and OD 260/230 is <2.0–2.2) | The DNA extraction method does not provide the required purity | The effects of contaminants are shown in the Contaminants document. Please try an alternative extraction method that does not result in contaminant carryover. Consider performing an additional SPRI clean-up step. |
| Low RNA integrity (RNA integrity number <9.5 RIN, or the rRNA band is shown as a smear on the gel) | The RNA degraded during extraction | Try a different RNA extraction method. For more info on RIN, please see the RNA Integrity Number document. Further information can be found in the DNA/RNA Handling page. |
| RNA has a shorter than expected fragment length | The RNA degraded during extraction | Try a different RNA extraction method. For more info on RIN, please see the RNA Integrity Number document. Further information can be found in the DNA/RNA Handling page. We recommend working in an RNase-free environment, and to keep your lab equipment RNase-free when working with RNA. |
Low DNA recovery after AMPure bead clean-up
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Low recovery | DNA loss due to a lower than intended AMPure beads-to-sample ratio | 1. AMPure beads settle quickly, so ensure they are well resuspended before adding them to the sample. 2. When the AMPure beads-to-sample ratio is lower than 0.4:1, DNA fragments of any size will be lost during the clean-up. |
| Low recovery | DNA fragments are shorter than expected | The lower the AMPure beads-to-sample ratio, the more stringent the selection against short fragments. Please always determine the input DNA length on an agarose gel (or other gel electrophoresis methods) and then calculate the appropriate amount of AMPure beads to use. 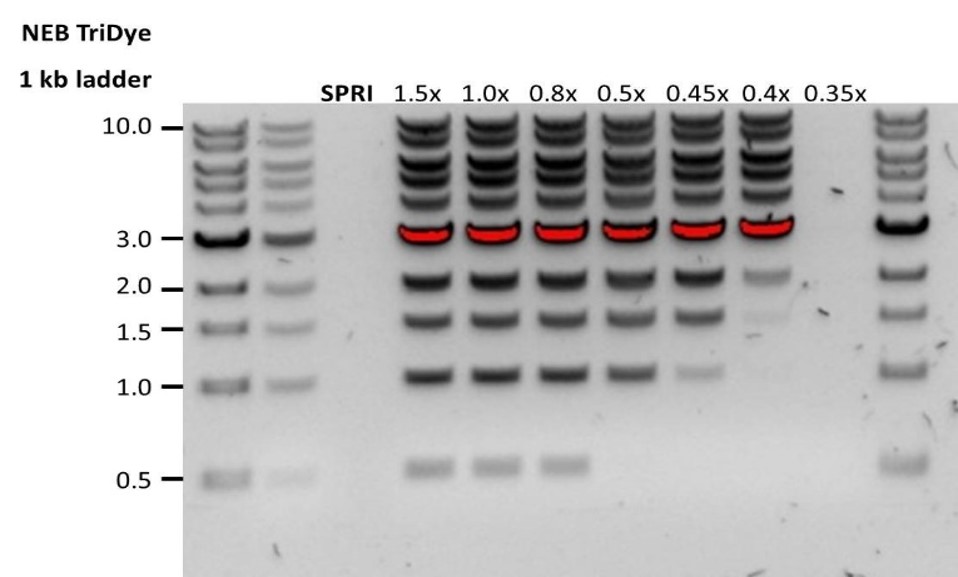 |
| Low recovery after end-prep | The wash step used ethanol <70% | DNA will be eluted from the beads when using ethanol <70%. Make sure to use the correct percentage. |
15. Issues during the sequencing run
Below is a list of the most commonly encountered issues, with some suggested causes and solutions.
We also have an FAQ section available on the Nanopore Community Support section.
If you have tried our suggested solutions and the issue still persists, please contact Technical Support via email (support@nanoporetech.com) or via LiveChat in the Nanopore Community.
Fewer pores at the start of sequencing than after Flow Cell Check
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | An air bubble was introduced into the nanopore array | After the Flow Cell Check it is essential to remove any air bubbles near the priming port before priming the flow cell. If not removed, the air bubble can travel to the nanopore array and irreversibly damage the nanopores that have been exposed to air. The best practice to prevent this from happening is demonstrated in this video. |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | The flow cell is not correctly inserted into the device | Stop the sequencing run, remove the flow cell from the sequencing device and insert it again, checking that the flow cell is firmly seated in the device and that it has reached the target temperature. If applicable, try a different position on the device (GridION/PromethION). |
| MinKNOW reported a lower number of pores at the start of sequencing than the number reported by the Flow Cell Check | Contaminations in the library damaged or blocked the pores | The pore count during the Flow Cell Check is performed using the QC DNA molecules present in the flow cell storage buffer. At the start of sequencing, the library itself is used to estimate the number of active pores. Because of this, variability of about 10% in the number of pores is expected. A significantly lower pore count reported at the start of sequencing can be due to contaminants in the library that have damaged the membranes or blocked the pores. Alternative DNA/RNA extraction or purification methods may be needed to improve the purity of the input material. The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
MinKNOW script failed
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Script failed" | Restart the computer and then restart MinKNOW. If the issue persists, please collect the MinKNOW log files and contact Technical Support. If you do not have another sequencing device available, we recommend storing the flow cell and the loaded library at 4°C and contact Technical Support for further storage guidance. |
Pore occupancy below 40%
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Pore occupancy <40% | Not enough library was loaded on the flow cell | Ensure you load the recommended amount of good quality library in the relevant library prep protocol onto your flow cell. Please quantify the library before loading and calculate mols using tools like the Promega Biomath Calculator, choosing "dsDNA: µg to pmol" |
| Pore occupancy close to 0 | The Ligation Sequencing Kit was used, and sequencing adapters did not ligate to the DNA | Make sure to use the NEBNext Quick Ligation Module (E6056) and Oxford Nanopore Technologies Ligation Buffer (LNB, provided in the sequencing kit) at the sequencing adapter ligation step, and use the correct amount of each reagent. A Lambda control library can be prepared to test the integrity of the third-party reagents. |
| Pore occupancy close to 0 | The Ligation Sequencing Kit was used, and ethanol was used instead of LFB or SFB at the wash step after sequencing adapter ligation | Ethanol can denature the motor protein on the sequencing adapters. Make sure the LFB or SFB buffer was used after ligation of sequencing adapters. |
| Pore occupancy close to 0 | No tether on the flow cell | Tethers are adding during flow cell priming (FLT/FCT tube). Make sure FLT/FCT was added to FB/FCF before priming. |
Shorter than expected read length
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Shorter than expected read length | Unwanted fragmentation of DNA sample | Read length reflects input DNA fragment length. Input DNA can be fragmented during extraction and library prep. 1. Please review the Extraction Methods in the Nanopore Community for best practice for extraction. 2. Visualise the input DNA fragment length distribution on an agarose gel before proceeding to the library prep. 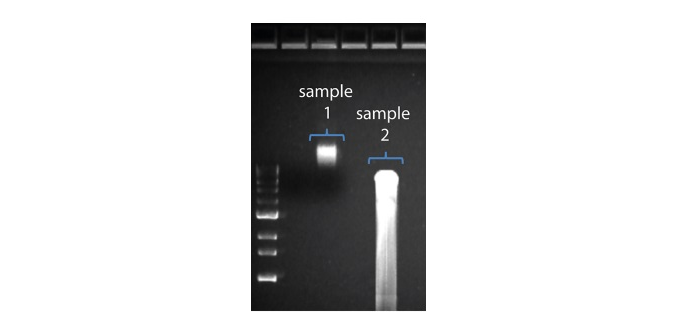 In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented. In the image above, Sample 1 is of high molecular weight, whereas Sample 2 has been fragmented.3. During library prep, avoid pipetting and vortexing when mixing reagents. Flicking or inverting the tube is sufficient. |
Large proportion of unavailable pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
Large proportion of unavailable pores (shown as blue in the channels panel and pore activity plot) 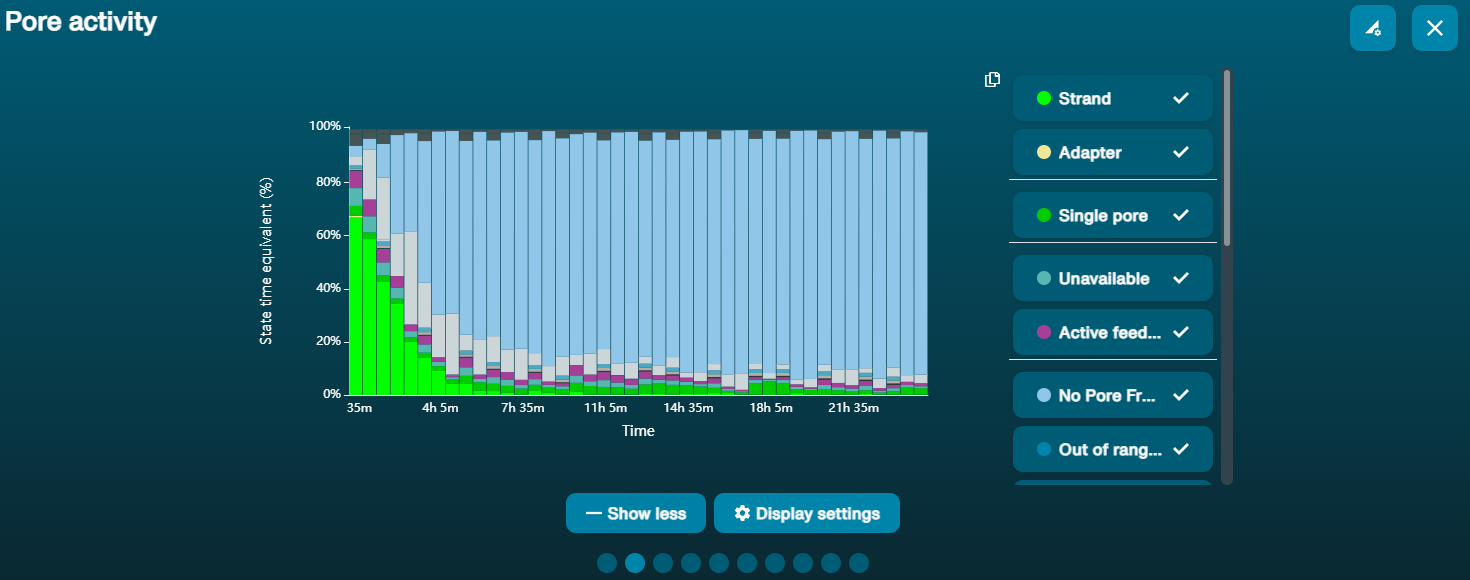 The pore activity plot above shows an increasing proportion of "unavailable" pores over time. The pore activity plot above shows an increasing proportion of "unavailable" pores over time. | Contaminants are present in the sample | Some contaminants can be cleared from the pores by the unblocking function built into MinKNOW. If this is successful, the pore status will change to "sequencing pore". If the portion of unavailable pores stays large or increases: 1. A nuclease flush using the Flow Cell Wash Kit (EXP-WSH004) can be performed, or 2. Run several cycles of PCR to try and dilute any contaminants that may be causing problems. |
Large proportion of inactive pores
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Large proportion of inactive/unavailable pores (shown as light blue in the channels panel and pore activity plot. Pores or membranes are irreversibly damaged) | Air bubbles have been introduced into the flow cell | Air bubbles introduced through flow cell priming and library loading can irreversibly damage the pores. Watch the Priming and loading your flow cell video for best practice |
| Large proportion of inactive/unavailable pores | Certain compounds co-purified with DNA | Known compounds, include polysaccharides, typically associate with plant genomic DNA. 1. Please refer to the Plant leaf DNA extraction method. 2. Clean-up using the QIAGEN PowerClean Pro kit. 3. Perform a whole genome amplification with the original gDNA sample using the QIAGEN REPLI-g kit. |
| Large proportion of inactive/unavailable pores | Contaminants are present in the sample | The effects of contaminants are shown in the Contaminants Know-how piece. Please try an alternative extraction method that does not result in contaminant carryover. |
Reduction in sequencing speed and q-score later into the run
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Reduction in sequencing speed and q-score later into the run | For Kit 9 chemistry (e.g. SQK-LSK109), fast fuel consumption is typically seen when the flow cell is overloaded with library (please see the appropriate protocol for your DNA library to see the recommendation). | Add more fuel to the flow cell by following the instructions in the MinKNOW protocol. In future experiments, load lower amounts of library to the flow cell. |
Temperature fluctuation
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Temperature fluctuation | The flow cell has lost contact with the device | Check that there is a heat pad covering the metal plate on the back of the flow cell. Re-insert the flow cell and press it down to make sure the connector pins are firmly in contact with the device. If the problem persists, please contact Technical Services. |
Failed to reach target temperature
| Observation | Possible cause | Comments and actions |
|---|---|---|
| MinKNOW shows "Failed to reach target temperature" | The instrument was placed in a location that is colder than normal room temperature, or a location with poor ventilation (which leads to the flow cells overheating) | MinKNOW has a default timeframe for the flow cell to reach the target temperature. Once the timeframe is exceeded, an error message will appear and the sequencing experiment will continue. However, sequencing at an incorrect temperature may lead to a decrease in throughput and lower q-scores. Please adjust the location of the sequencing device to ensure that it is placed at room temperature with good ventilation, then re-start the process in MinKNOW. Please refer to this link for more information on MinION temperature control. |
Guppy – no input .fast5 was found or basecalled
| Observation | Possible cause | Comments and actions |
|---|---|---|
| No input .fast5 was found or basecalled | input_path did not point to the .fast5 file location | The --input_path has to be followed by the full file path to the .fast5 files to be basecalled, and the location has to be accessible either locally or remotely through SSH. |
| No input .fast5 was found or basecalled | The .fast5 files were in a subfolder at the input_path location | To allow Guppy to look into subfolders, add the --recursive flag to the command |
Guppy – no Pass or Fail folders were generated after basecalling
| Observation | Possible cause | Comments and actions |
|---|---|---|
| No Pass or Fail folders were generated after basecalling | The --qscore_filtering flag was not included in the command | The --qscore_filtering flag enables filtering of reads into Pass and Fail folders inside the output folder, based on their strand q-score. When performing live basecalling in MinKNOW, a q-score of 7 (corresponding to a basecall accuracy of ~80%) is used to separate reads into Pass and Fail folders. |
Guppy – unusually slow processing on a GPU computer
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Unusually slow processing on a GPU computer | The --device flag wasn't included in the command | The --device flag specifies a GPU device to use for accelerate basecalling. If not included in the command, GPU will not be used. GPUs are counted from zero. An example is --device cuda:0 cuda:1, when 2 GPUs are specified to use by the Guppy command. |
.gif)







