Kit de lavado de celdas de flujo (EXP-WSH004 o EXP-WSH004-XL) (WFC_9120_v1_revS_25Jul2025)
MinION: Protocol
Kit de lavado de celdas de flujo (EXP-WSH004 o EXP-WSH004-XL) V WFC_9120_v1_revS_25Jul2025
De uso exclusivo en investigación
FOR RESEARCH USE ONLY.
Contents
Introducción
Preparativos
- 3. Consideraciones y reconocimiento de las celdas de flujo MinION y GridION
- 4. Preparar los reactivos
Lavar la celda de flujo
Opciones tras el lavado
Descripción general
De uso exclusivo en investigación
1. Descripción general del protocolo
Introducción al kit de lavado
Este kit hace posible ejecutar bibliotecas de secuenciación de manera consecutiva en la misma celda de flujo. Primero se elimina y se digiere la biblioteca de ADN presente en la celda; a continuación, se restablece el sistema, de modo que quede listo para cargar una biblioteca nueva. El kit es compatible con las celdas de flujo R9.4.1 y R10.4.1.
También se utiliza con las celdas de flujo de ARN para purgar las bibliotecas correspondientes, pero no elimina los bloqueos asociados ni restablece los nanoporos.
Gracias a este kit, la misma celda se reutiliza varias veces*, lo que prolonga el tiempo de ejecución disponible.
*El número de lavados y de reutilizaciones dependerá de varios factores. Por lo general, las cifras oscilan entre 3 y 6 por celda de flujo. Encontrará más información sobre el tema y los factores limitantes, en el documento técnico Kit de lavado de celdas de flujo (EXP-WSH004 o EXP-WSH004-XL).
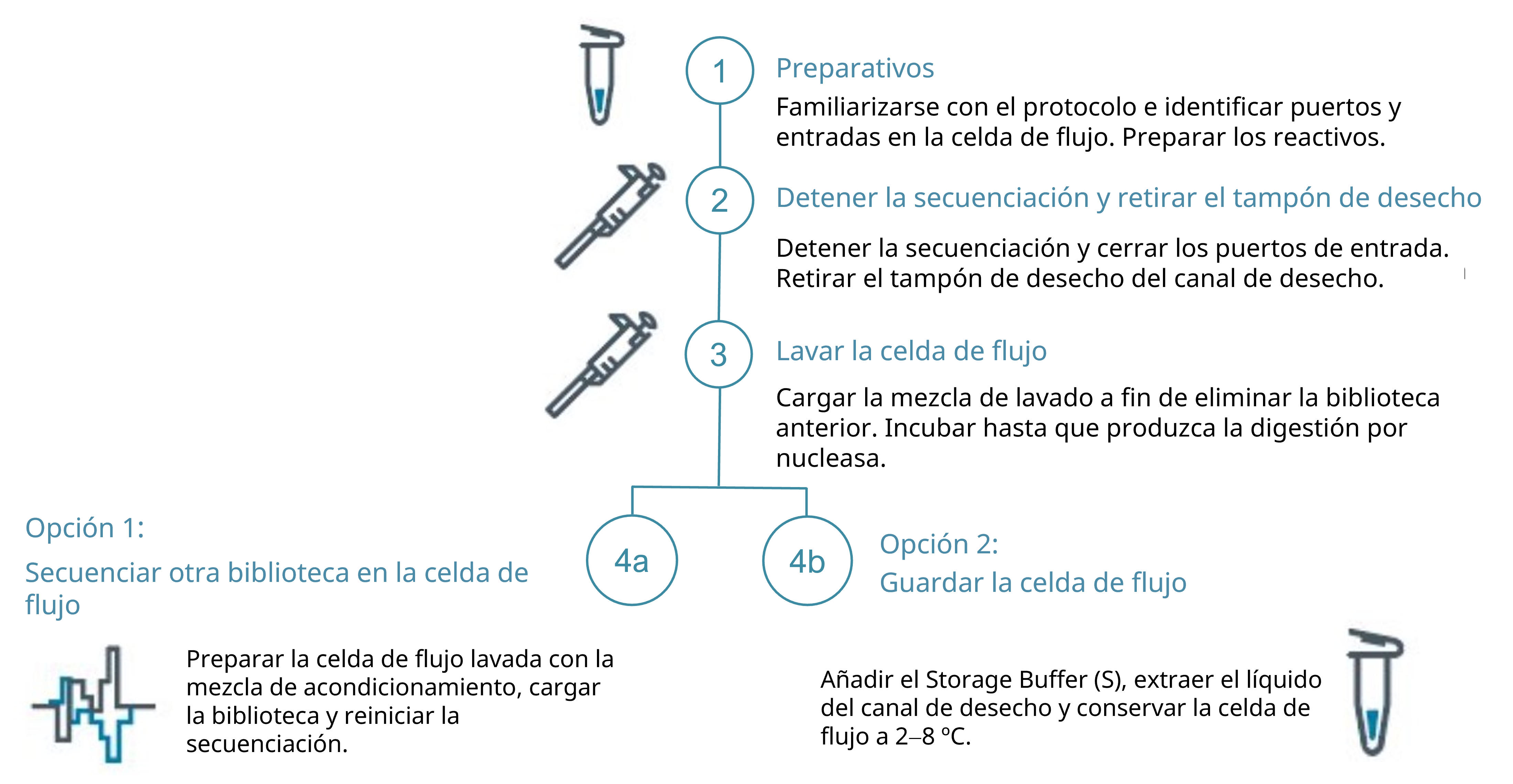
Tras el paso de lavado, dispone de dos opciones de reutilización:
- Cargar una biblioteca nueva e iniciar la secuenciación de inmediato.
- Introducir Storage Buffer (S) del kit de lavado en la celda de flujo y guardarla a 2–8 °C.
Documento técnico Kit de lavado de celdas de flujo (EXP-WSH004 o EXP-WSH004)
Encontrará más información sobre el funcionamiento del kit, así como datos de rendimiento y recomendaciones adicionales en el documento técnico Kit de lavado de celdas de flujo (EXP-WSH004 o EXP-WSH004-XL).
2. Material y consumibles
Material
- Flow Cell Wash Kit (EXP-WSH004) o Flow Cell Wash Kit XL (EXP-WSH004-XL)
- Reactivos de acondicionamiento presentes en el kit de secuenciación o en los siguientes kits:
- Sequencing Auxiliary Vials V14 (EXP-AUX003)
- Flow Cell Priming Kit (EXP-FLP004)
Instrumental
- Pipeta y puntas P1000
- Pipeta y puntas P20
- Cubeta con hielo
Recarga de la biblioteca
Tras lavar la celda de flujo, se necesitan tampones adicionales para recargar la biblioteca; dichos tampones se encuentran en uno de los siguientes kits complementarios:
Sequencing Auxiliary Vials V14 (EXP-AUX003), que contiene viales de Sequencing Buffer (SB), Elution Buffer (EB), Library Solution (LIS) y Library Beads (LIB), junto con los reactivos de acondicionamiento del Kit 14: Flow Cell Flush (FCF) y Flow Cell Tether (FCT), compatibles exclusivamente con la química del Kit 14.
Flow Cell Priming Kit (EXP-FLP004), que contiene los dos reactivos utilizados en el acondicionamiento de celdas de flujo del Kit 14: Flow Cell Flush (FCF) y Flow Cell Tether (FCT), compatibles exclusivamente con la química del Kit 14.
Contenido del kit Flow Cell Wash (EXP-WSH004)
- Wash Mix (WMX) – contiene DNasa I.
- Wash Diluent (DIL) – contiene el tampón de exonucleasa que potencia la actividad de la DNasa I.
- Storage Buffer (S) – mantiene las celdas de flujo en condiciones adecuadas hasta el siguiente uso.
Contenido del kit Flow Cell Wash Kit XL(EXP-WSH004-XL)
- Wash Mix (WMX) – contiene DNasa I.
- Wash Diluent (DIL) – contiene el tampón de exonucleasa que potencia la actividad de la DNasa I.
- Storage Buffer (S) – mantiene las celdas de flujo en condiciones adecuadas hasta el siguiente uso.
3. Consideraciones y reconocimiento de las celdas de flujo MinION y GridION
Mantener la celda de flujo en el dispositivo durante todo el proceso.
Recomendamos encarecidamente mantener la celda de flujo insertada en el dispositivo a fin de evitar desconexiones y mantener un control adecuado de la temperatura durante todo el proceso.
La pantalla protectora durante el lavado.
Lo ideal es que la pantalla protectora permanezca insertada en la celda de flujo durante el lavado. No obstante, si prefiere retirarla a fin de mejorar la visibilidad de la matriz, es posible hacerlo.
Siga las directrices del protocolo y no olvide volver a colocarla antes de reanudar o iniciar un nuevo experimento de secuenciación.
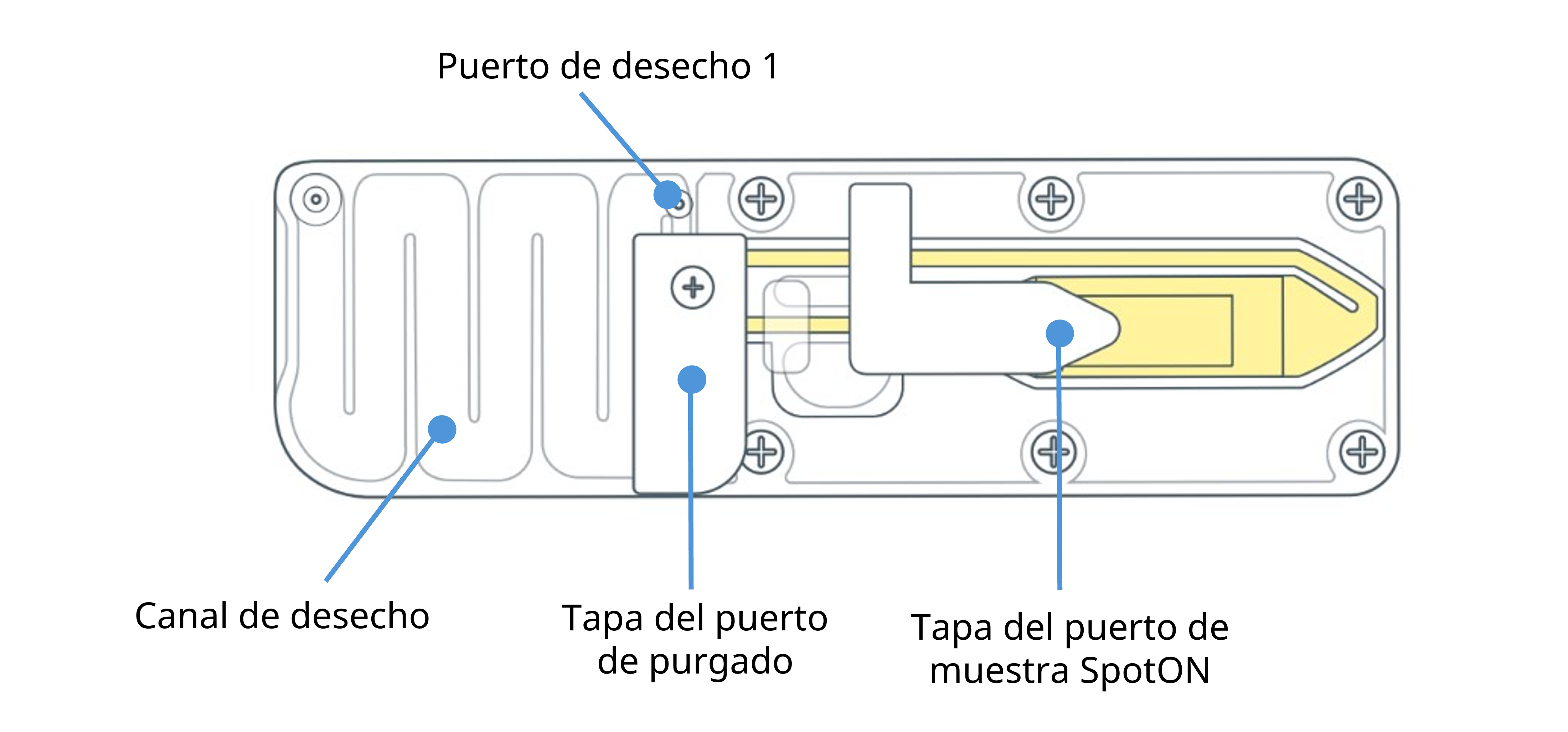
Familiarícese con las partes de las celdas de flujo MinION/GridION.
Si es necesario, recupere la biblioteca de la(s) celda(s) de flujo
Si tiene previsto recuperar la biblioteca antes de proceder con el lavado, realice este paso en primer lugar.
Las instrucciones se encuentran en el protocolo de recuperación de bibliotecas de celdas de flujo.
Mire el video sobre cómo lavar y recargar celdas de flujo MinION/GridION
que muestra cómo lavar una celda de flujo tras un experimento de secuenciación y cómo cargar una nueva biblioteca. Recomendamos mirar sin falta este video de demostración antes de realizar el procedimiento:
Vídeo sobre cómo lavar y cargar celdas de flujo MinION/GridION
4. Preparar los reactivos
Material
- Flow Cell Wash Kit (EXP-WSH004) o Flow Cell Wash Kit XL (EXP-WSH004-XL)
Consumibles
- Tubos Eppendorf DNA LoBind de 1,5 ml
Instrumental
- Pipeta y puntas P1000
- Pipeta y puntas P20
- Mezclador vórtex
- Microcentrífuga
- Cubeta con hielo
Recomendamos vivamente preparar los reactivos antes de cada uso.
Si prepara previamente la mezcla Flow Cell Wash Mix no la guarde durante más de un día, ya que ello conlleva resultados subóptimos.
Colocar el tubo Wash Mix (WMX) en hielo. No mezclar en vórtex.
Descongelar un vial de Wash Diluent (DIL) a temperatura ambiente.
Mezclar su contenido minuciosamente mediante vórtex, centrifugar brevemente y colocar en hielo.
En un tubo Eppendorf DNA LoBind nuevo de 1,5 ml preparar la siguiente mezcla:
| Reactivo | Volumen necesario por celda de flujo |
|---|---|
| Wash Mix (WMX) | 2 μl |
| Wash Diluent (DIL) | 398 μl |
| Total | 400 μl |
Mezclar bien con la pipeta y colocar los tubo(s) en hielo.
Nota: no mezclar en vortex el tubo Flow Cell Wash Mix.
Colocar el tubo en hielo hasta que se necesite.
5. Interrumpir/detener la secuenciación y retirar el tampón de desecho
Instrumental
- Pipeta y puntas P1000
Mantener la celda de flujo en el dispositivo durante todo el proceso.
Recomendamos mantener la celda de flujo insertada en el dispositivo, a fin de evitar desconexiones, reducir al mínimo posibles daños o errores y mejorar la usabilidad.
Detener o interrumpir el experimento de secuenciación en MinKNOW
La celda de flujo no debe secuenciar durante ninguno de los pasos de lavado. Manipularla mientras secuencia provoca daños y pérdida de poros.
Cierre las tapas del puerto de purgado y del puerto de muestra SpotON.
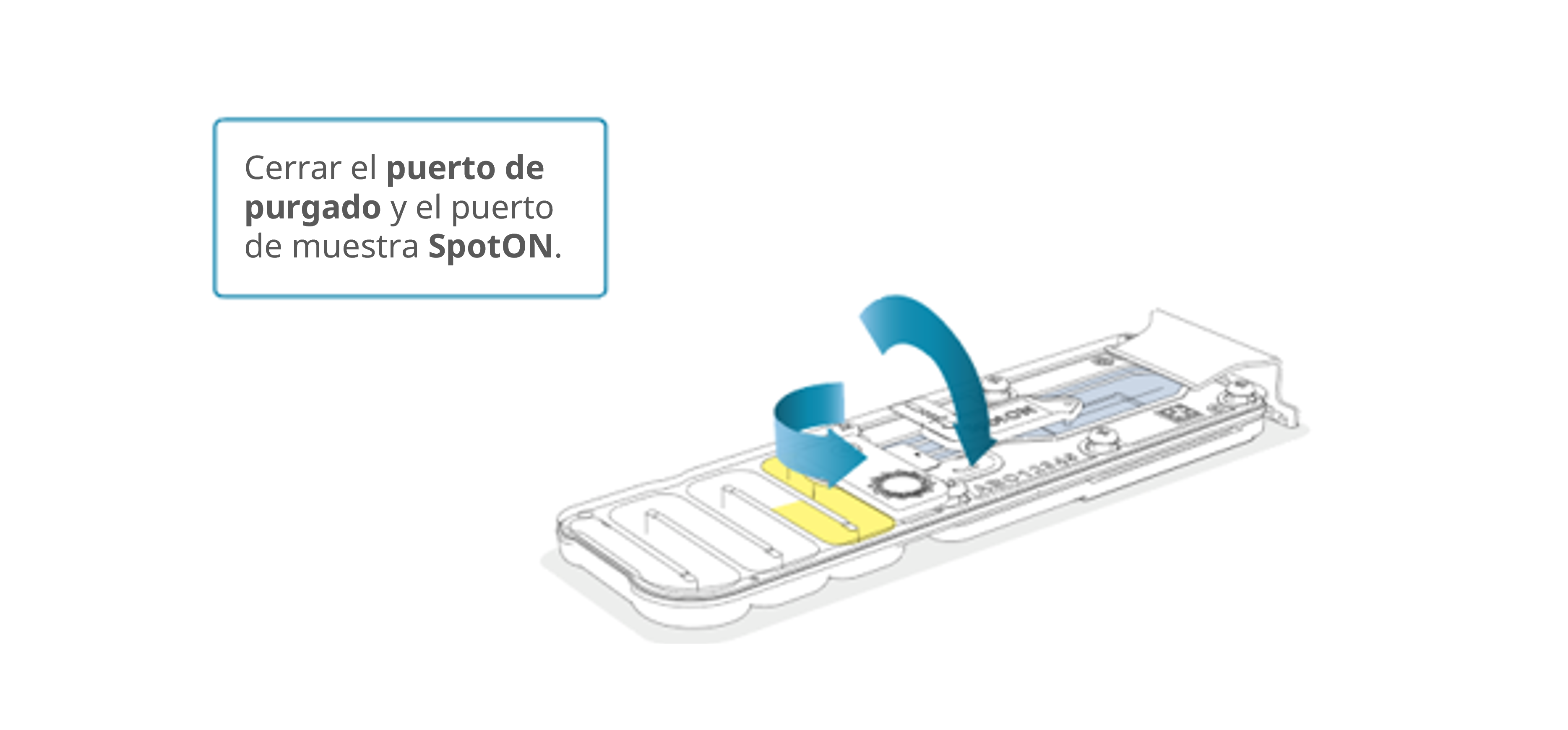
Importante: es fundamental que el puerto de purgado y el puerto de muestra SpotON estén cerrados antes de retirar el tampón de desecho y así evitar que el aire atraviese la zona de la matriz de sensores, lo que provocaría una pérdida significativa de canales de secuenciación.
Retirar el resto del tampón del puerto de residuos 1 con una pipeta P1000.
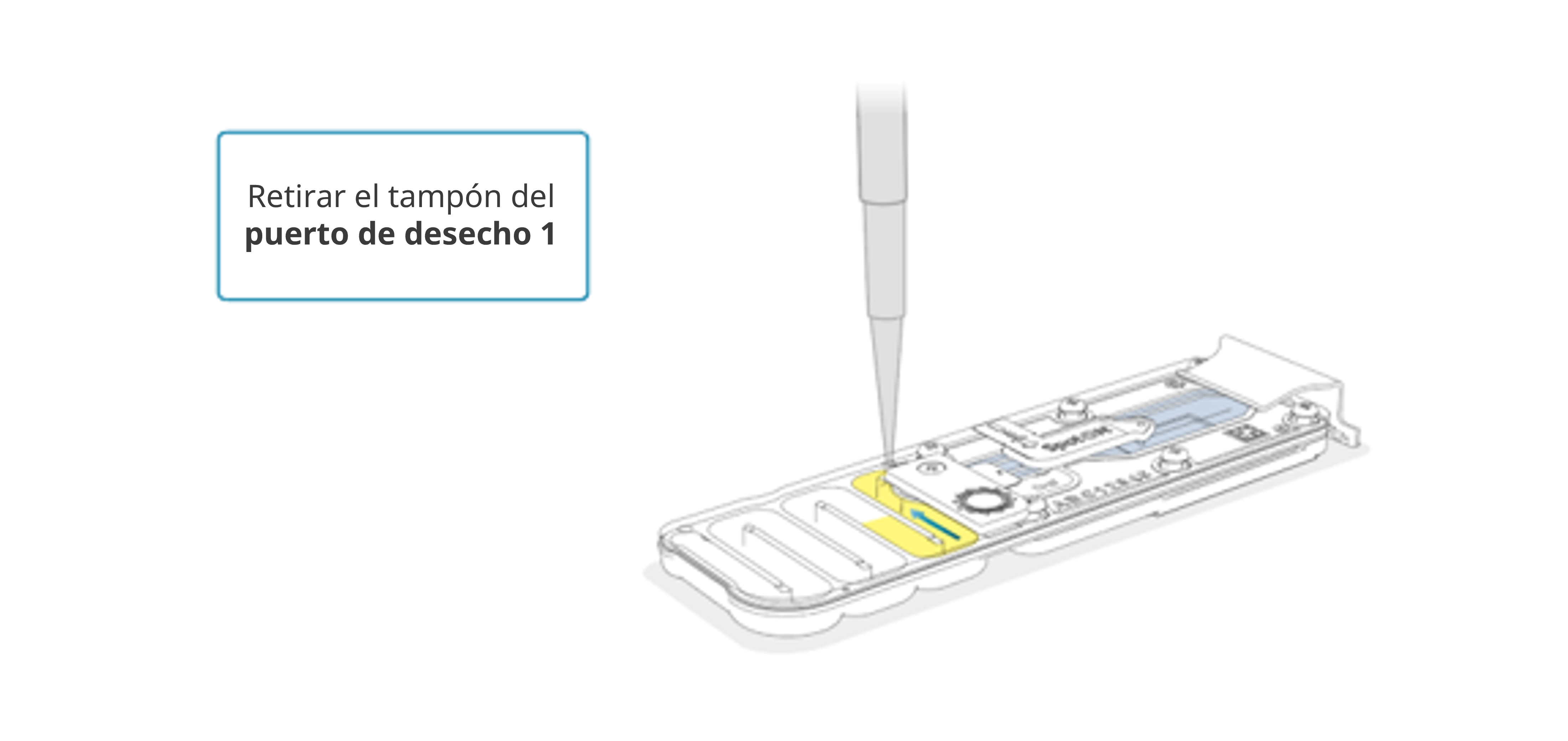
Retirar todo el tampón del canal de residuos. Si es necesario, deseche el volumen aspirado y repita la operación hasta extraer todo el contenido.
- Ajustar una pipeta P1000 a 1000 μl.
- Insertar la punta en el puerto de residuos 1.
- Aspirar lentamente hasta retirar el tampón.
Importante: dado que tanto el puerto de purgado como el puerto de muestra SpotON están cerrados, no debería salir ningún líquido de la zona de la matriz de sensores.
A continuación, lavar la celda de flujo.
6. Lavar la celda de flujo
Material
- Mezcla de lavado (del paso anterior)
Instrumental
- Pipeta y puntas P1000
- Pipeta y puntas P200
- Cubeta con hielo
Girar la tapa del puerto de purgado en el sentido de las agujas del reloj.
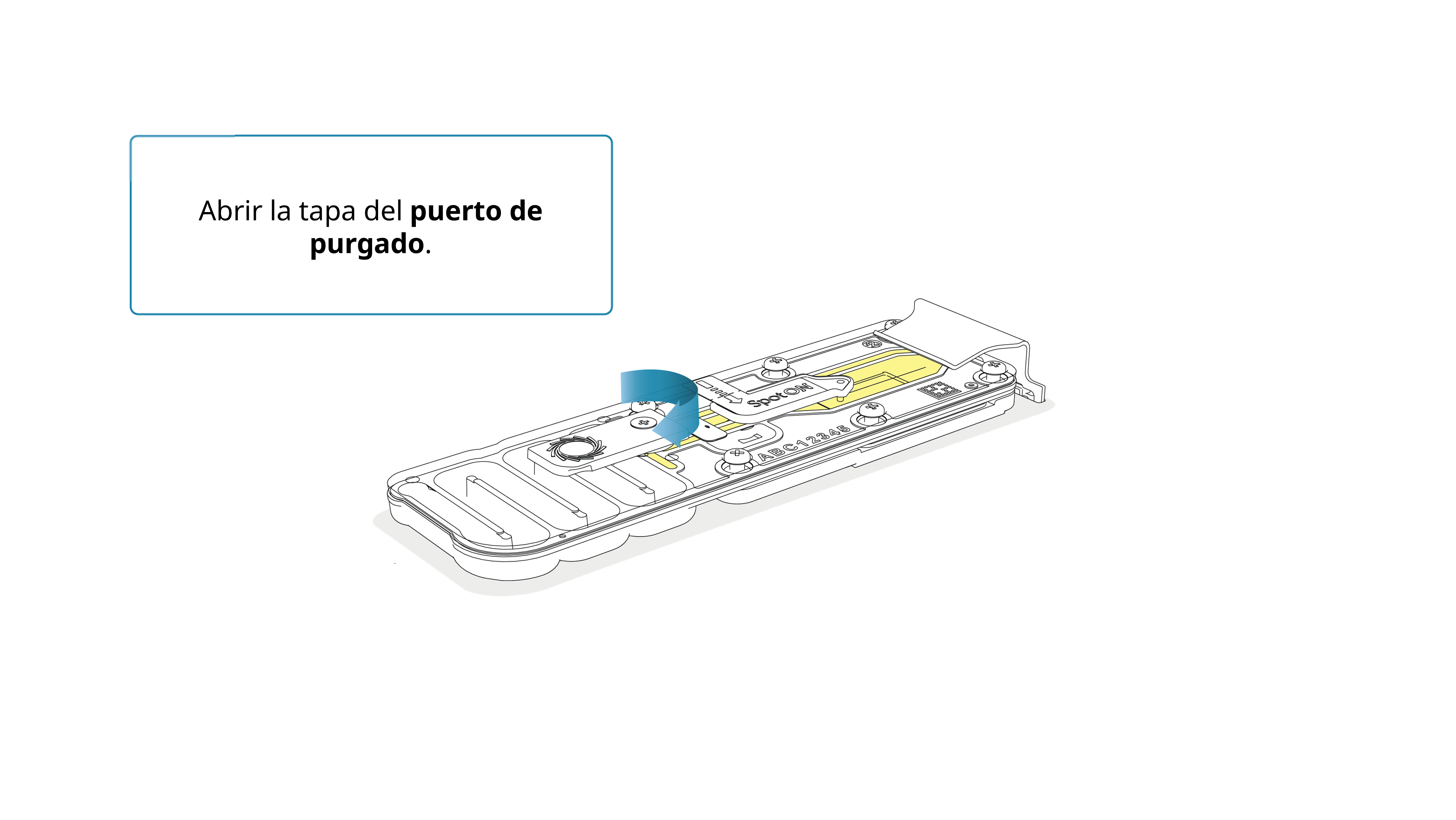
Tenga cuidado al extraer tampón. No retire más de 20-30 μl y compruebe que cubra la matriz de poros en todo momento. La introducción de burbujas de aire en la matriz puede dañar los poros de manera irreversible.
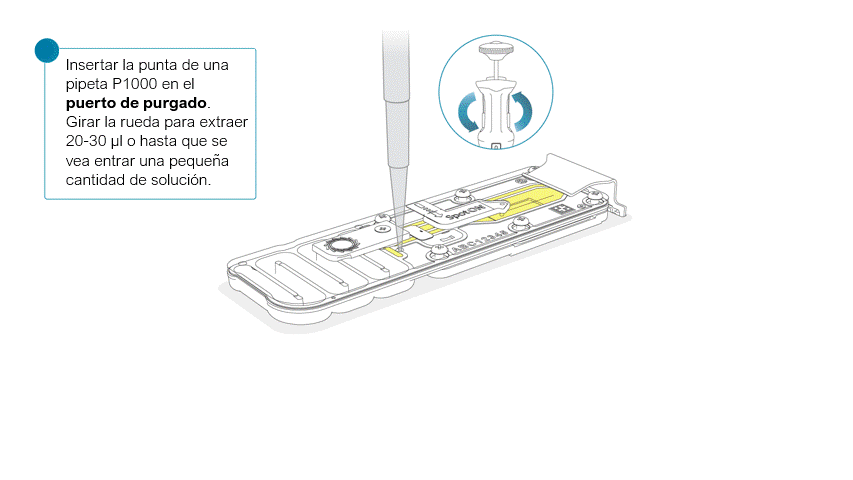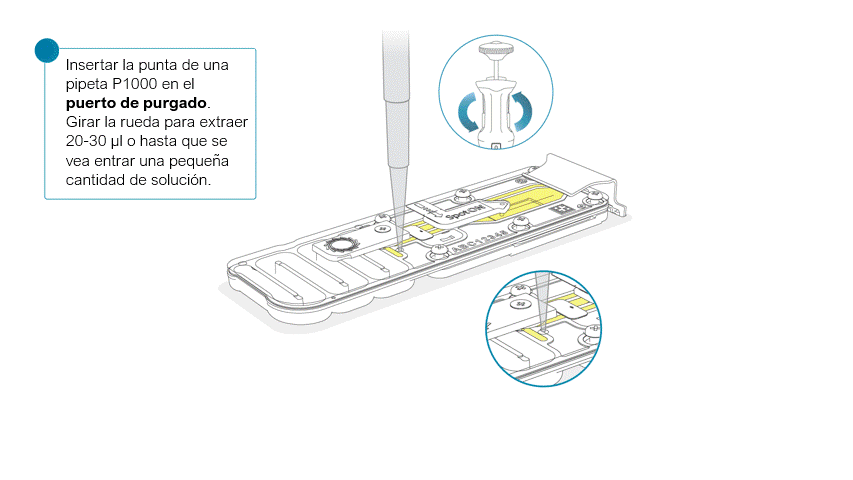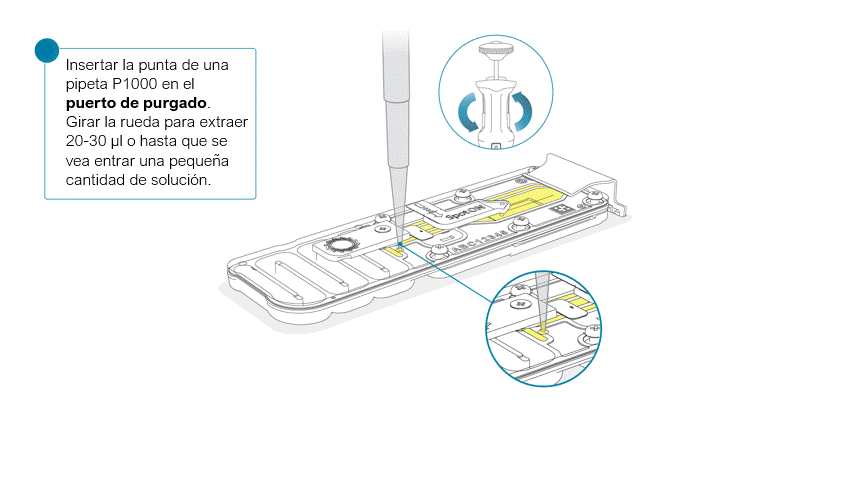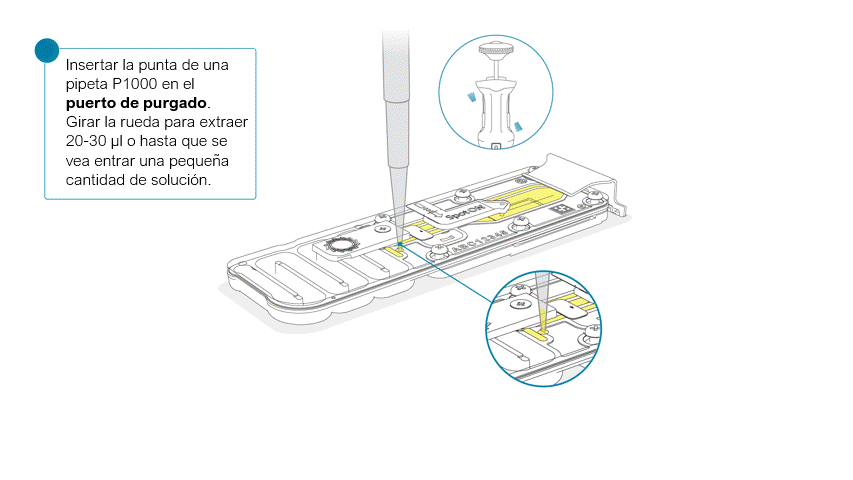
Una vez abierta, comprobar si debajo hay burbujas de aire. Extraer una pequeña cantidad de líquido para eliminar posibles burbujas:
- Ajustar una pipeta P1000 a 200 μl.
- Introducir la punta en el puerto de purgado.
- Girar la rueda hasta que el indicador de volumen marque 220-230 μl o hasta que se vea una pequeña cantidad de líquido entrar en la punta de la pipeta.
- Compruebe que haya un flujo continuo de tampón circulando desde el puerto de purgado a través de la matriz de poros.
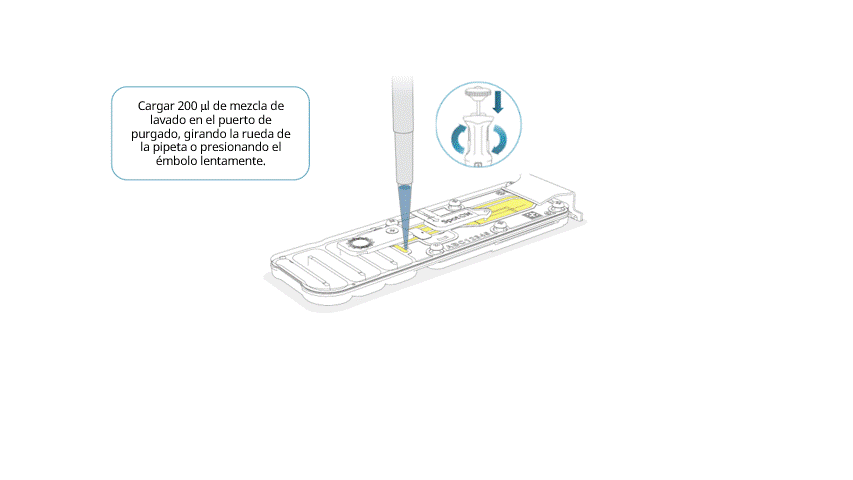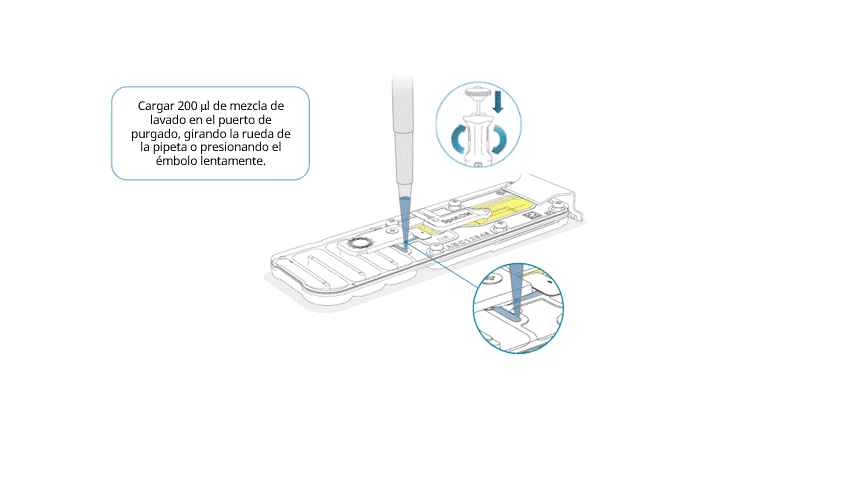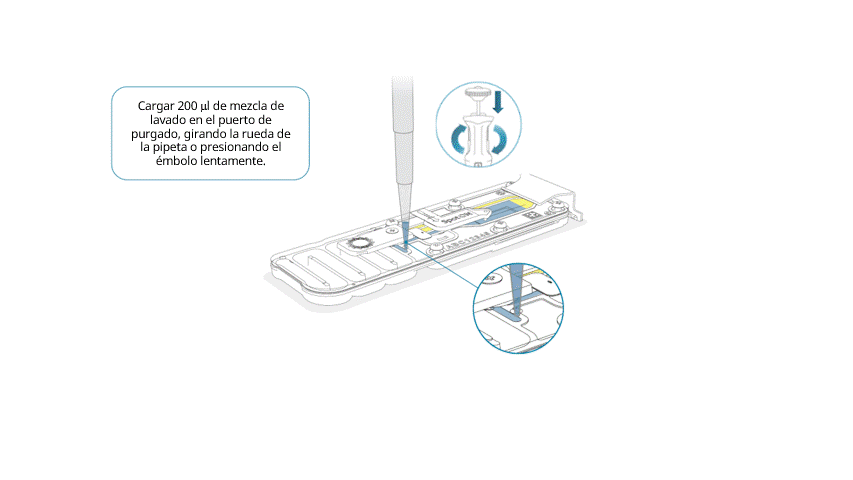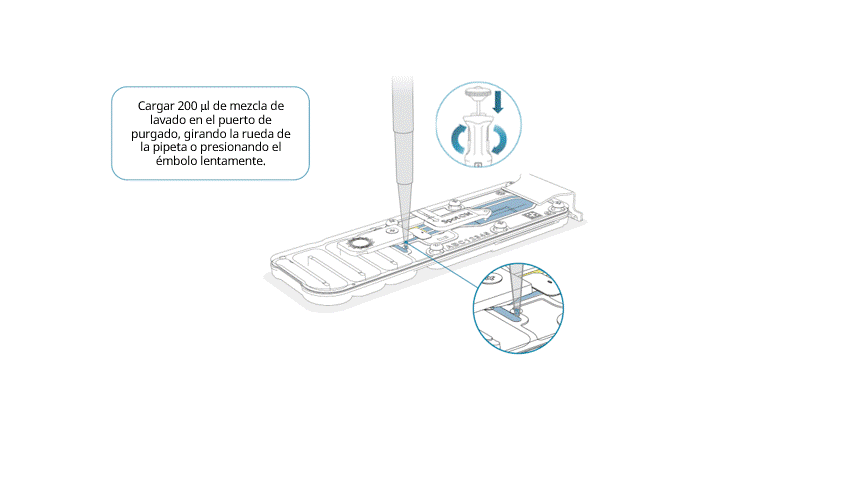
Cargar lentamente 200 μl de mezcla de lavado en el puerto de purgado, de la siguiente manera:
- Con una pipeta P1000, extraer 200 µl de mezcla de lavado.
- Introducir la punta de la pipeta en el puerto de purgado y comprobar que no haya burbujas de aire en la punta.
- Girar lentamente la rueda de la pipeta (si es posible) o presionar el émbolo muy lentamente y dejar una cantidad pequeña de tampón en la punta.
Importante: la adición de la mezcla de lavado en la celda de flujo debe realizarse durante al menos 5 segundos, muy lentamente y a un ritmo constante.
Ajustar un temporizador e incubar la mezcla de lavado en la celda de flujo durante 5 minutos.
Una vez finalizada la incubación, repetir el paso de carga de la mezcla de lavado (paso 3) con otros 200 µl de la misma mezcla.
Cerrar el puerto de purgado y esperar 1 hora.

Retirar el resto del tampón del puerto de desecho 1 con una pipeta P1000.

Ya se ha lavado la celda de flujo; continuar con el siguiente paso.
Continuar con:
- Opción 1: secuenciar una segunda biblioteca
- Opción 2: conservar la celda de flujo hasta más adelante
Importante: no guardar la celda de flujo con la mezcla de lavado en la matriz de poros, ya que existe el riesgo de que se pierdan poros de manera irreversible.
7. Opción 1: secuenciar una segunda biblioteca
Material
- Reactivos de acondicionamiento presentes en el kit de secuenciación o en los siguientes kits:
- Sequencing Auxiliary Vials V14 (EXP-AUX003)
- Flow Cell Priming Kit V14 (EXP-FLP004)
Instrumental
- Pipeta y puntas P1000
- Pipeta y puntas P20
- Cubeta con hielo
Los reactivos de secuenciación descritos en este procedimiento son compatibles con la química V14 más reciente.
Si utiliza una versión anterior de la química o un kit con reactivos de secuenciación determinados, verifique la compatibilidad con el procedimiento que vaya a realizar.
Los tampones utilizados en este proceso son incompatibles con la comprobación de la celda de flujo previa a la carga de la biblioteca siguiente. No obstante, una vez iniciada la secuenciación, el primer escaneo de poros informará sobre el número de nanoporos disponibles.
Descongelar los viales Sequencing Buffer (SB), Library Beads (LIB) o Library Solution (LIS) —si se requiere—, Flow Cell Tether (FCT) y un tubo de Flow Cell Flush (FCF) a temperatura ambiente y mezclar en vórtex. Centrifugar brevemente y poner en hielo.
A fin de optimizar el rendimiento de la secuenciación y mejorar la producción de las celdas de flujo R10.4.1 (FLO-MIN114), recomendamos añadir seroalbúmina bovina (BSA) a la mezcla de acondicionamiento a una concentración final de 0,2 mg/ml.
No recomendamos utilizar ningún otro tipo de albúmina (p. ej., seroalbúmina humana recombinante).
Preparar la mezcla de acondicionamiento con BSA. Añadir los siguientes reactivos y mezclar minuciosamente con la pipeta:
| Reactivo | Volumen por celda de flujo |
|---|---|
| Flow Cell Flush (FCF) | 1 170 µl |
| Seroalbúmina bovina (BSA) a 50 mg/ml | 5 μl |
| Flow Cell Tether (FCT) | 30 µl |
| Total | 1 205 µl |
Girar la tapa del puerto de purgado en el sentido de las agujas del reloj.
Una vez abierta, comprobar si debajo hay burbujas de aire. Extraer una pequeña cantidad de líquido para eliminar posibles burbujas:
- Ajustar una pipeta P1000 a 200 μl.
- Introducir la punta en el puerto de purgado.
- Girar la rueda hasta que el dial marque 220-230 µl o hasta que se vea una pequeña cantidad de tampón entrar en la punta
Importante: tenga cuidado al extraer el tampón; no retire más de 20-30 µl. Comprobar que un flujo continuo de solución circule desde el puerto de purgado a través de la matriz de poros.
Cargar 800 μl de mezcla de acondicionamiento por el puerto de purgado, de la siguiente manera:
- Con una pipeta P1000, extraer 800 µl de mezcla de acondicionamiento.
- Introducir la punta de la pipeta en el puerto de purgado y comprobar que no haya burbujas de aire en la punta.
- Girar lentamente la rueda de la pipeta (si es posible) o presionar el émbolo muy lentamente, tal como se muestra en el vídeo de demostración, y dejar una cantidad pequeña de tampón en la punta.
Importante: la adición de los 800 µl de mezcla de lavado en la celda de flujo debe realizarse durante al menos 10 segundos, muy lentamente y a un ritmo constante.
Es fundamental esperar cinco minutos entre cada purgado con la mezcla de acondicionamiento para eliminar la nucleasa de manera eficaz.
Cerrar el puerto de purgado y esperar cinco minutos.
Durante este tiempo, preparar la biblioteca siguiendo los pasos a continuación.
Mezclar minuciosamente con la pipeta el contenido del vial Library Beads (LIB).
El vial Library Beads (LIB) contiene microesferas en suspensión que sedimentan muy rápido; por eso es crucial mezclarlas justo antes de su uso.
Recomendamos utilizar Library Beads (LIB) en la mayoría de experimentos de secuenciación. No obstante, el reactivo Library Solution (LIS) está disponible en caso de que se utilicen bibliotecas más viscosas.
En un tubo nuevo, preparar la biblioteca conforme a la sección "Preparación y carga de la celda de flujo MinION/GridION" del protocolo correspondiente, de modo que se utilicen los reactivos y volúmenes correctos.
| Reactivo | Volumen por celda de flujo |
|---|---|
| Sequencing Buffer (SB) | 37,5 µl |
| Library Beads (LIB) bien mezclado antes de su uso o Library Solution (LIS) si se utiliza | 25,5 µl |
| Biblioteca de ADN | 12 µl |
| Total | 75 µl |
Retirar el resto del tampón del puerto de desecho 1 con una pipeta P1000.

Retirar todo el tampón del canal de desecho. Si es necesario, deseche el volumen aspirado y repita la operación hasta extraer todo el contenido.
- Ajustar una pipeta P1000 a 1000 μl.
- Insertar la punta en el puerto de desecho 1.
- Aspirar lentamente hasta retirar el tampón.
Importante: dado que tanto el puerto de purgado como el puerto de muestra SpotON están cerrados, no debería salir ningún líquido de la zona de la matriz de sensores.
Cargar lentamente 200 μl de mezcla de acondicionamiento por el puerto de purgado, de la siguiente manera:
- Abrir el puerto de purgado y levantar con cuidado el puerto de muestra SpotON.
- Con una pipeta P1000, extraer 200 µl de mezcla de acondicionamiento.
- Introducir la punta de la pipeta en el puerto de purgado y comprobar que no haya burbujas de aire en la punta.
- Girar lentamente la rueda de la pipeta (si es posible) o presionar el émbolo muy lentamente, tal como se muestra en el vídeo de demostración, y dejar una cantidad pequeña de tampón en la punta.
Importante: la adición de los 200 µl de mezcla de lavado debe realizarse durante al menos 5 segundos, muy lentamente y a un ritmo constante.
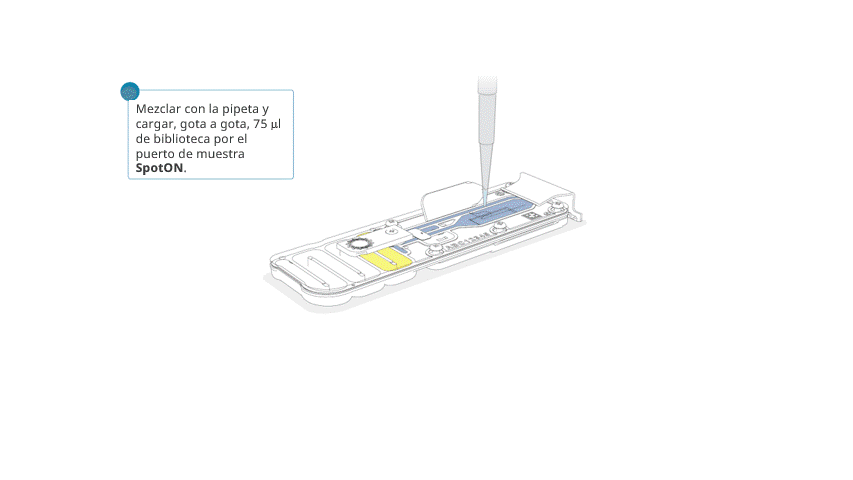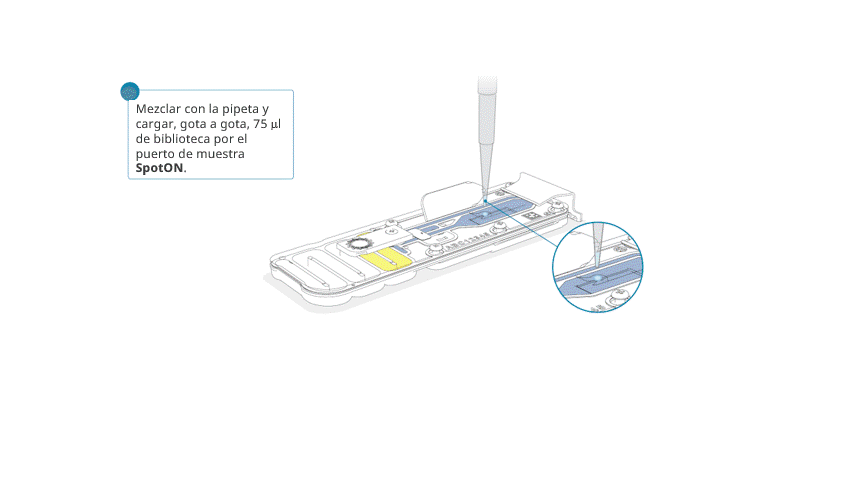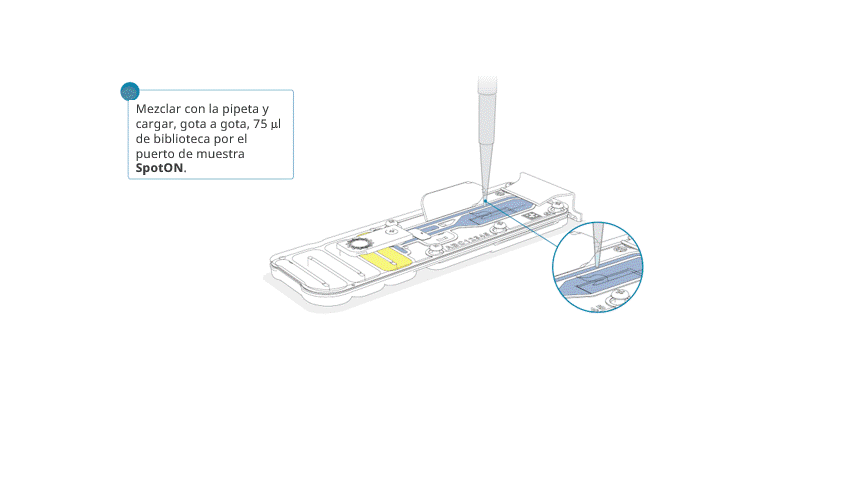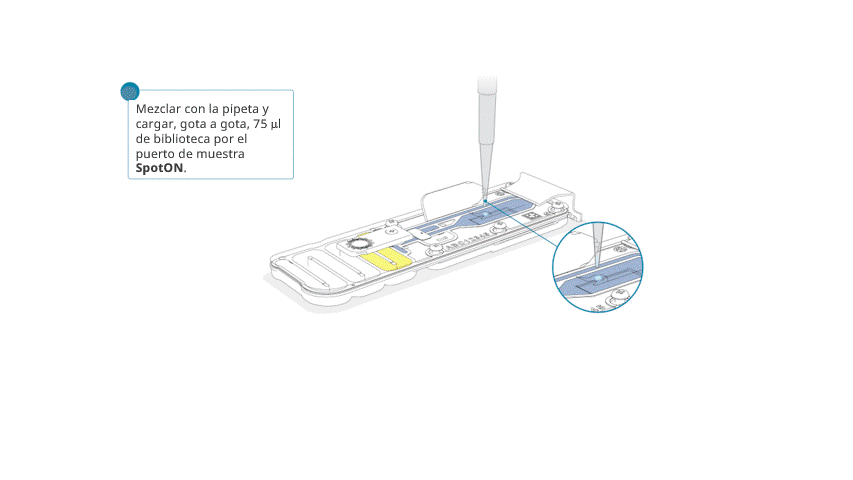
Aspirar y dispensar suavemente la biblioteca de secuenciación antes de cargarla.
Añadir, gota a gota, 75 μl de la biblioteca preparada por el puerto SpotON. Esperar a que cada gota fluya hacia adentro del puerto antes de añadir la siguiente.
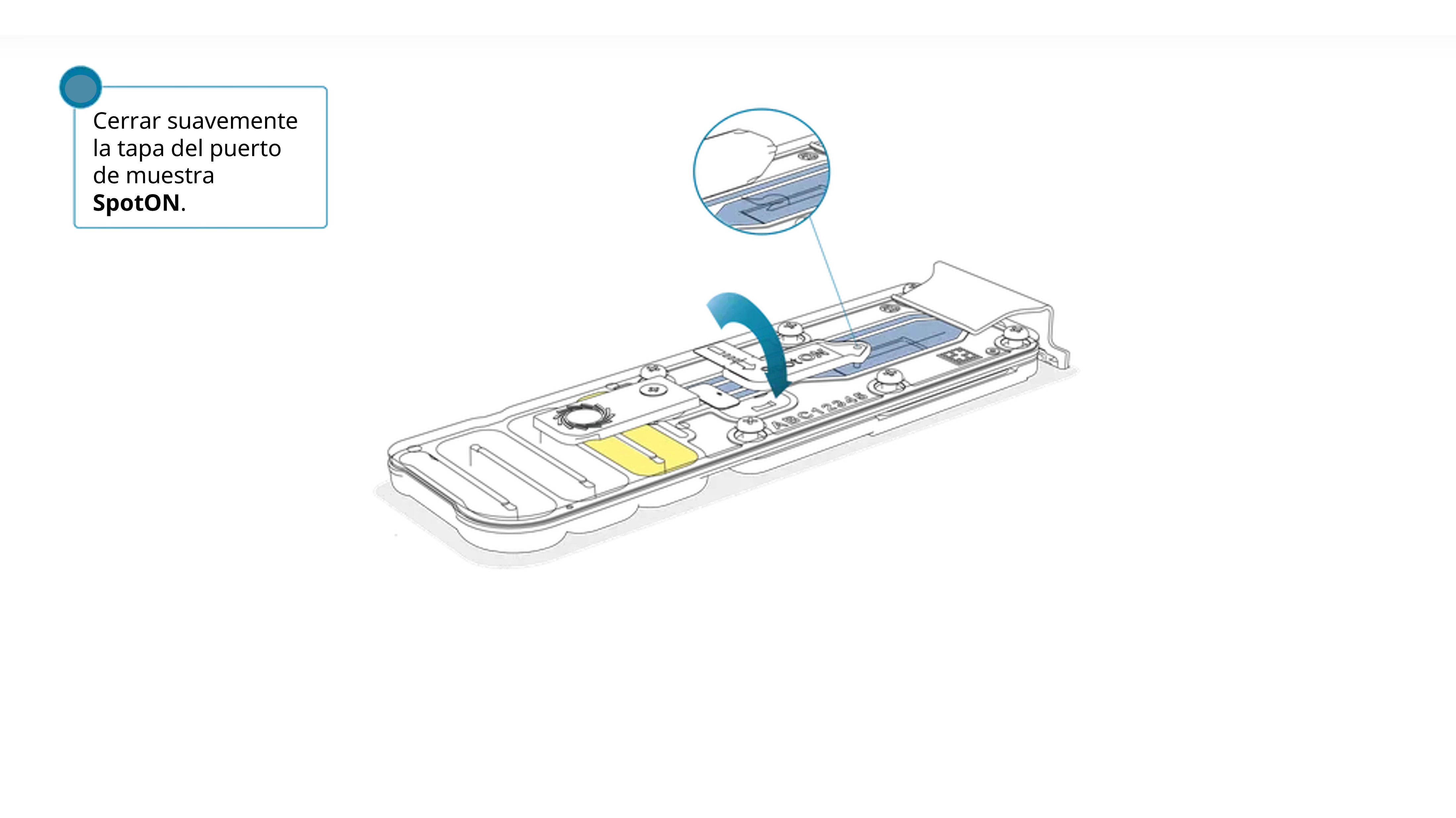
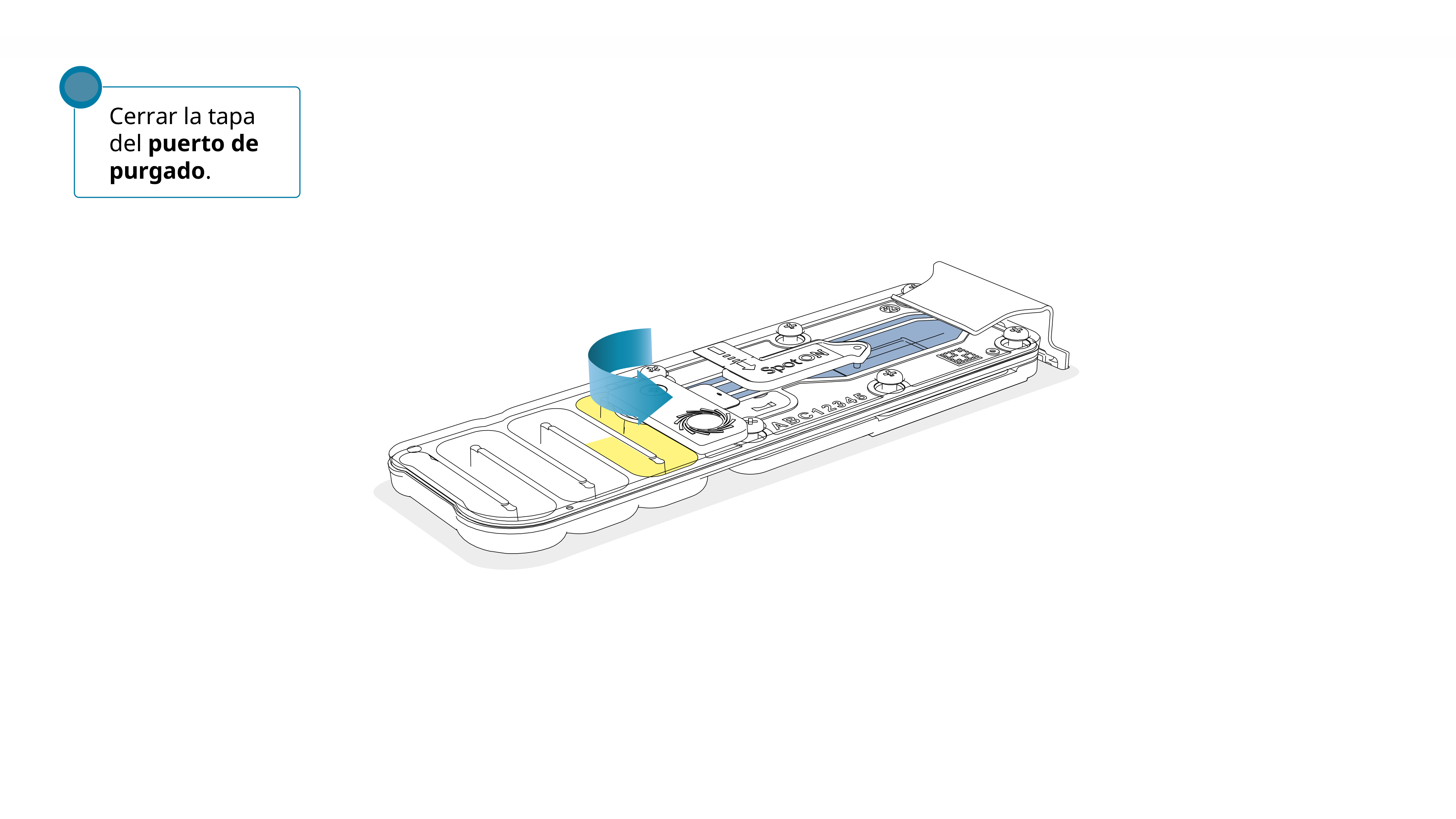
Volver a colocar la tapa con cuidado, comprobar que el tapón encaje en el agujero y cerrar el puerto de purgado.


Colocar la pantalla protectora sobre la celda de flujo en cuanto se haya cargado la biblioteca favorecerá un rendimiento de secuenciación óptimo.
Recomendamos no retirarla mientras la biblioteca esté cargada ni durante los lavados o recargas. Se retira una vez se haya extraído la biblioteca.
Colocar la pantalla protectora de la celda de flujo, como se indica a continuación:
Apoyar con cuidado el borde delantero contra el clip. Importante: no forzar la pieza para que pase por debajo.
Al posar la pantalla protectora sobre la celda de flujo, la pieza debe asentarse alrededor de la tapa SpotON y cubrir por completo la sección superior de la celda de flujo.
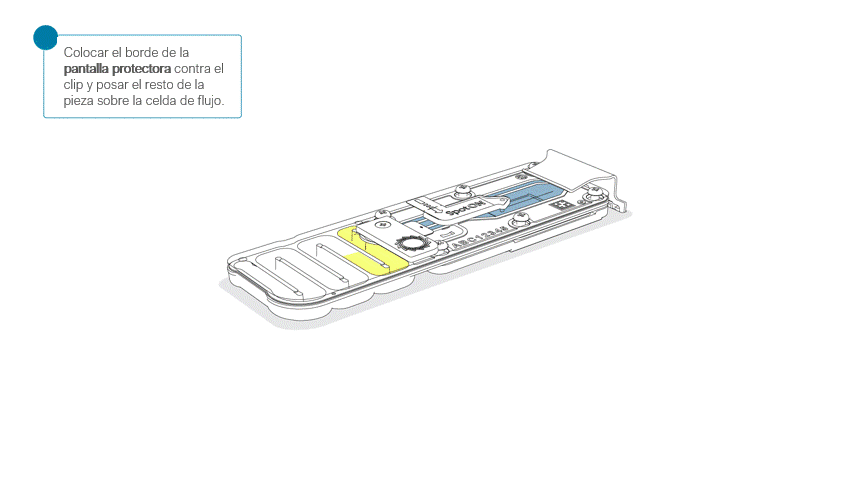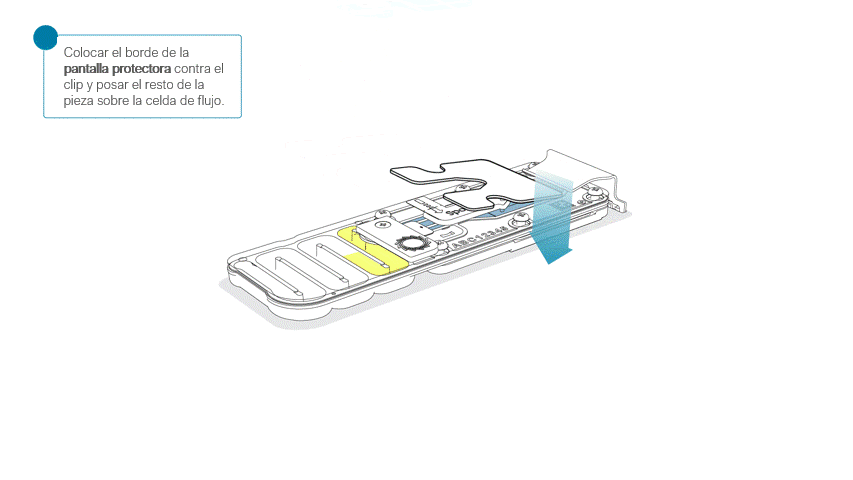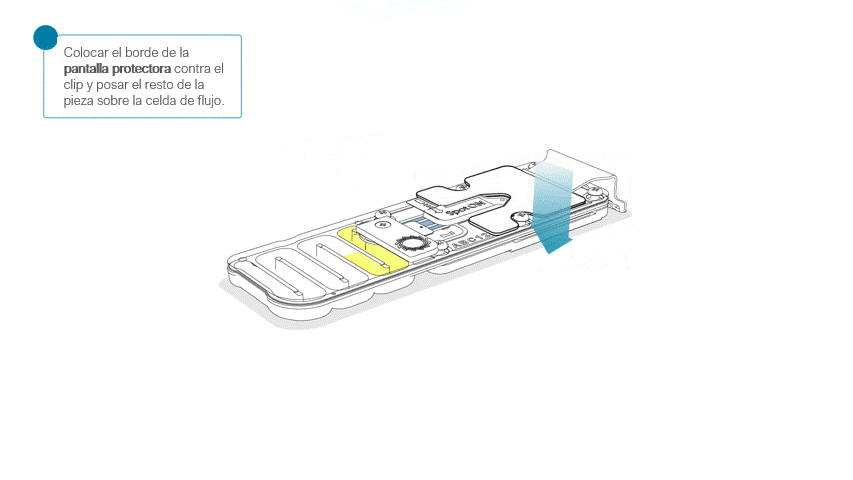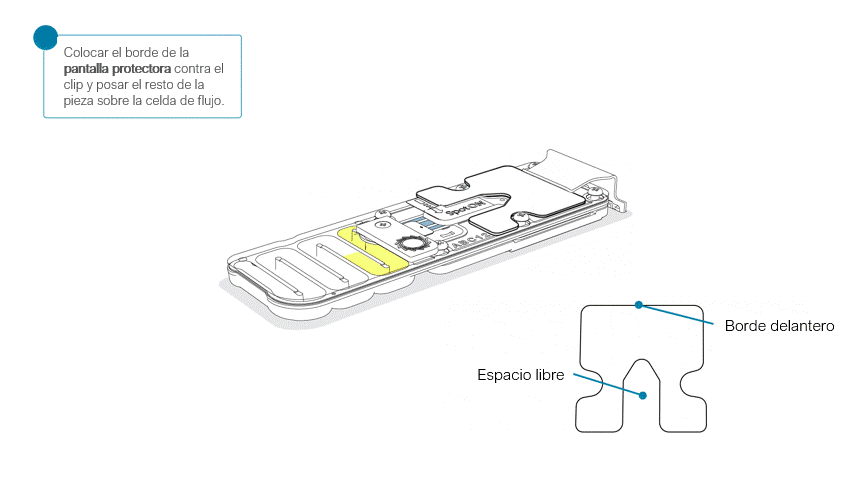
La pantalla protectora no queda sujeta a la celda de flujo MinION. Tras su colocación, combiene manipular la celda con cuidado.
Cerrar la tapa del dispositivo y continuar con el experimento de secuenciación en MinKNOW.
Recomendaciones de almacenamiento de la biblioteca
Recomendamos guardar la biblioteca en tubos Eppendorf DNA LoBind a 4 °C, durante periodos cortos o en caso de uso repetido, por ejemplo, durante la recarga de celdas de flujo entre lavados. En caso de uso individual o de conservación a largo plazo (más de 3 meses), recomendamos guardar las bibliotecas a –80 °C en tubos Eppendorf DNA LoBind. Encontrará más información en el documento técnico sobre estabilidad en bibliotecas de ADN.
8. Opción 2: conservar la celda de flujo hasta más adelante
Material
- Flow Cell Wash Kit (EXP-WSH004) o Flow Cell Wash Kit XL (EXP-WSH004-XL)
Instrumental
- Pipeta y puntas P1000
- Pipeta y puntas P20
El Storage Buffer (S) se utiliza en el purgado de las celdas de flujo así como en la comprobación del número de poros disponibles antes de cargar otra biblioteca.
Descongelar un vial de Storage Buffer (S) a temperatura ambiente.
Girar la tapa del puerto de purgado en el sentido de las agujas del reloj.
Una vez abierto, comprobar si debajo hay burbujas de aire. Extraer una pequeña cantidad de líquido para eliminar posibles burbujas:
- Ajustar una pipeta P1000 a 200 μl.
- Introducir la punta en el puerto de purgado.
- Girar la rueda hasta que el dial marque 220-230 µl o hasta que se vea una cantidad pequeña de tampón entrar en la punta.
Importante: tenga cuidado al extraer el tampón; no retire más de 20-30 µl y compruebe que circule un flujo continuo desde el puerto de purgado a través de la matriz de poros.
Cargar 500 μl de Storage Buffer (S) de la siguiente manera:
- Con una pipeta P1000, extraer 500 µl de Storage Buffer (S).
- Introducir la punta de la pipeta en el puerto de purgado y comprobar que no haya burbujas de aire en la punta.
- Girar lentamente la rueda de la pipeta (si es posible) o presionar el émbolo muy lentamente y dejar una cantidad pequeña de tampón en la punta.
Importante: la adición de Storage Buffer (S) en la celda de flujo debe realizarse durante al menos 5 segundos, muy lentamente y a un ritmo constante.
Retirar el resto del tampón del puerto de desecho 1 con una pipeta P1000.

Retirar todo el tampón del canal de desecho. Si es necesario, descarte el volumen aspirado y repita la operación hasta extraer todo el contenido.
- Cerrar el puerto de purgado.
- Ajustar una pipeta P1000 a 1000 μl.
- Insertar la punta en el puerto de residuos 1.
- Aspirar lentamente hasta retirar el tampón.
Importante: dado que tanto el puerto de purgado como el puerto de muestra SpotON están cerrados, no debe salir ningún líquido de la zona de la matriz de sensores.
La celda de flujo debe guardarse a 2–8 °C.
Cuando se desee reutilizar, sacar de la nevera y dejar que se aclimate a temperatura ambiente durante aproximadamente 5 minutos.
Después de un lavado o tras haber estado guardada, recomendamos "Comprobar la celda de flujo" y verificar el número de poros disponibles que tiene.
Cargar la celda en el dispositivo con el Storage Buffer (S) e iniciar una comprobación de la misma, a fin de conocer el número de poros activos que tiene. Encontrará más información en la sección Comprobación de la celda de flujo del protocolo de MinKNOW.
Al terminar, acondicionar la celda de flujo y cargar la biblioteca antes de comenzar un nuevo experimento de secuenciación.







