Science Unlocked: publication picks from April 2026
In this monthly series, we share a selection of recent publications that use Oxford Nanopore sequencing to unlock novel insights. Spanning tumour profiling, carrier screening, and pathogen surveillance, these studies showcase the advances in scientific research made possible by Oxford Nanopore sequencing.
Featured in this edition:
1. Nanopore-only T2T genome assembly
2. Accessible bioinformatics for resource-limited settings
3. Comprehensive tumour profiling in a single assay
4. Streamlined screening for inherited disorders
Bioinformatics
1. Telomere-to-telomere assembly using HERRO-corrected nanopore reads (Nature)
Telomere-to-telomere (T2T) assemblies are setting a new standard for reference-quality genomes, yet their generation has traditionally required multiple sequencing platforms, making the process costly and resource-intensive. However, Stanojević et al. have now developed HERRO (Haplotype-aware ERRor cOrrection), a deep learning framework that enables T2T assembly using only Oxford Nanopore ultra-long reads.
HERRO with the Verkko assembler produced error-corrected reads while maintaining haplotype integrity, enabling accurate, high-quality T2T assemblies using nanopore data alone. This approach simplifies workflows, reduces sequencing costs, enables scalability, and brings comprehensive, reference-quality genome assembly within reach for more projects.
‘HERRO achieves up to 100-fold improvement in read accuracy while preserving true biological differences between related genomic sequences’
Stanojević, D. et al.1
Keen to generate complete T2T assemblies to advance your research? Register your interest in our T2T product bundle.
Microbiology
2. Bridging the bioinformatics gap: tool selection for decentralised pathogen surveillance in Africa (Frontiers in Public Health)
Antimicrobial resistance (AMR) remains a pressing global challenge, particularly in low-resource settings. SeqAfrica, one of the first genomic AMR surveillance networks in Africa, identified a need for an analysis pipeline suitable for researchers with limited bioinformatics skills in low-resource settings. The pipeline needs to be capable of performing comprehensive quality control, genome assembly, species identification, multi-locus sequence typing, resistance gene detection, and phylogenetic analysis. The project trialled over 80 bioinformatics tools alongside the affordable MinION device.
EPI2ME, our cloud-integrated, graphical user interface (GUI)-based platform, emerged as a leading choice thanks to its simplicity, accessibility, and seamless integration into local workflows. EPI2ME is free for researchers to use on nanopore data and can be run locally or in the cloud. This work by Lacy-Roberts and Gibson et al. highlights how user-friendly solutions like EPI2ME can empower laboratories across Africa to generate actionable genomic insights — without the need for extensive bioinformatics expertise.
‘EPI2ME, which is an Oxford Nanopore Technologies tool, was favoured for its offline capability, ease of use, and compatibility with local infrastructure’
Lacy-Roberts, N. and Gibson, C.T. et al. 2
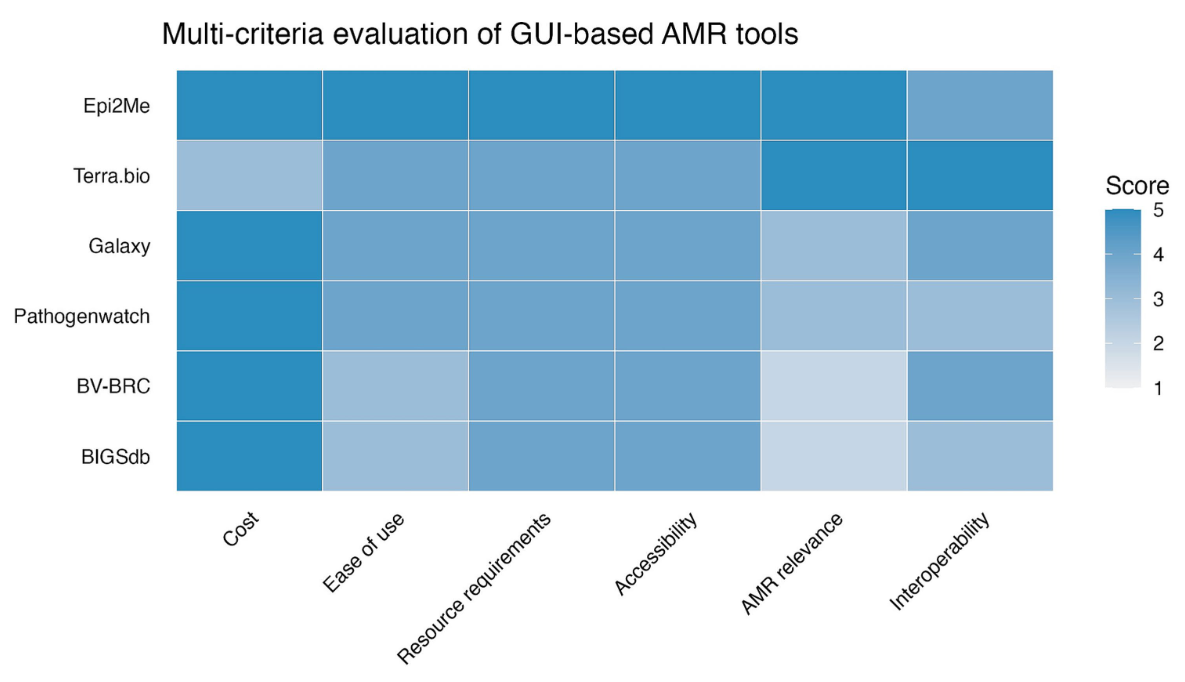
Figure 1: Multi-criteria evaluation of GUI-based AMR tools. Each criterion was scored on a five-point scale, where one indicated very poor performance, and five indicated excellent performance. Figure redistributed from Lacy-Roberts and Gibson et al. 20262 under Creative Commons Attribution License CC BY 4.0.
Find out more about data analysis using EPI2ME, or check out our nanopore-only microbial isolate sequencing solution (NO-MISS).
Cancer research
3. Whole-genome nanopore sequencing for rapid comprehensive classification of brain tumours (medRxiv)
Characterising central nervous system (CNS) tumours is challenging due to the range of genetic and epigenetic variants involved. Halldorsson et al. compared nanopore whole-genome sequencing (WGS) to clinical standard-of-care (SOC) tests on 90 CNS tumour research samples. Our technology successfully classified all tumours containing >15% tumour cell content, and identified complex deletions not covered by SOC tests. Nanopore sequencing provided methylation-based classification within one hour, which matched the final SOC diagnosis in 89% (80/90) cases. All diagnostically relevant copy number variations, single-nucleotide variants, and gene fusions aligned with SOC testing, and MGMT promoter methylation status matched in 94% of cases — a marker which could be therapeutically relevant for CNS tumours.
Compared with SOC tests, nanopore sequencing reduced hands-on time by 8–28 hours, saved an average of $400, and reduced the overall turnaround time from sample to result by five days. These findings show the potential for our technology to be used as a single, comprehensive method to characterise tumours and support timely therapeutic decisions in the future.
‘[Nanopore] WGS consolidates comprehensive molecular diagnostics into a single assay that is accurate, scalable, and clinically feasible’
Halldorsson, S. et al.3
See how you could multiply your insights with PromethION 24.
Human genetics
4. Performance evaluation of a PCR/Oxford Nanopore assay for carrier screening (The Journal of Molecular Diagnostics)
Carrier screening is critical for identifying individuals at risk of passing on genetic disorders, yet traditional approaches often require multiple workflows and can miss rare variants. Martin et al. evaluated a new approach for carrier screening of three common genetic disorders: cystic fibrosis (CF), spinal muscular atrophy (SMA), and fragile X syndrome (FXS). Using the AmplideX Nanopore Carrier Plus reagents, gene-specific PCR was performed, followed by barcoding and sequencing on MinION Flow Cells.
The results aligned well with traditional screening — 100% for SMA and FXS, and 97% for CF — while also detecting additional variants and low-level mosaicism. By combining multiple targets into a single assay, the method offers broader variant detection across complex genomic regions than current carrier screening methods. These findings highlight the potential for nanopore sequencing to deliver comprehensive, flexible solutions for genetic analysis in the future.
‘This single workflow enables accurate, reproducible screening for multiple disorders, with the ability to identify more CFTR variants than traditional genotyping panels’
Martin, K.E. et al. 4
Our flagship conference, London Calling, will take place 19–21 May 2026. In-person tickets have sold out, but you can still register for a free virtual ticket.
Want a sneak preview? Check out our blog on what’s coming.
Apply Oxford Nanopore sequencing to your own research questions and you'll never see sequencing the same way again. Explore the nanopore sequencing solution.
Oxford Nanopore Technologies products are not intended for use for health assessment or to diagnose, treat, mitigate, cure, or prevent any disease or condition.
Stanojević, D., Lin, D., Nurk, S., Florez de Sessions, P., and Šikić, M. Telomere-to-telomere assembly using HERRO-corrected simplex nanopore reads. Nature Online ahead of print (2026). DOI: https://doi.org/10.1038/s41586-026-10563-y
Lacy-Roberts, N. and Gibson, C.T. et al. Bridging the bioinformatics gap: tool selection for decentralised AMR genomic surveillance in Africa. Front. Public Health 14:1756324 (2026). DOI: https://doi.org/10.3389/fpubh.2026.1756324
Halldorsson, S. et al. Whole-genome long-read sequencing for rapid comprehensive molecular diagnostics of brain tumours. medRxiv 26351563 (2026). DOI: https://doi.org/10.64898/2026.04.23.26351563
Martin, K.E., Sábato, F., Lynch, J., Ferreira-Gonzalez, A., and Barrie, E.S. Performance evaluation of a PCR/nanopore assay for carrier screening for cystic fibrosis, spinal muscular atrophy, and fragile X syndrome. J. Mol. Diagn. (2026). DOI: https://doi.org/10.1016/j.jmoldx.2026.03.003






