Enrichment analysis of k-mer composition enables identification of telomeres
- shared.published_on: July 30 2019
- shared.source: ECCB 2019
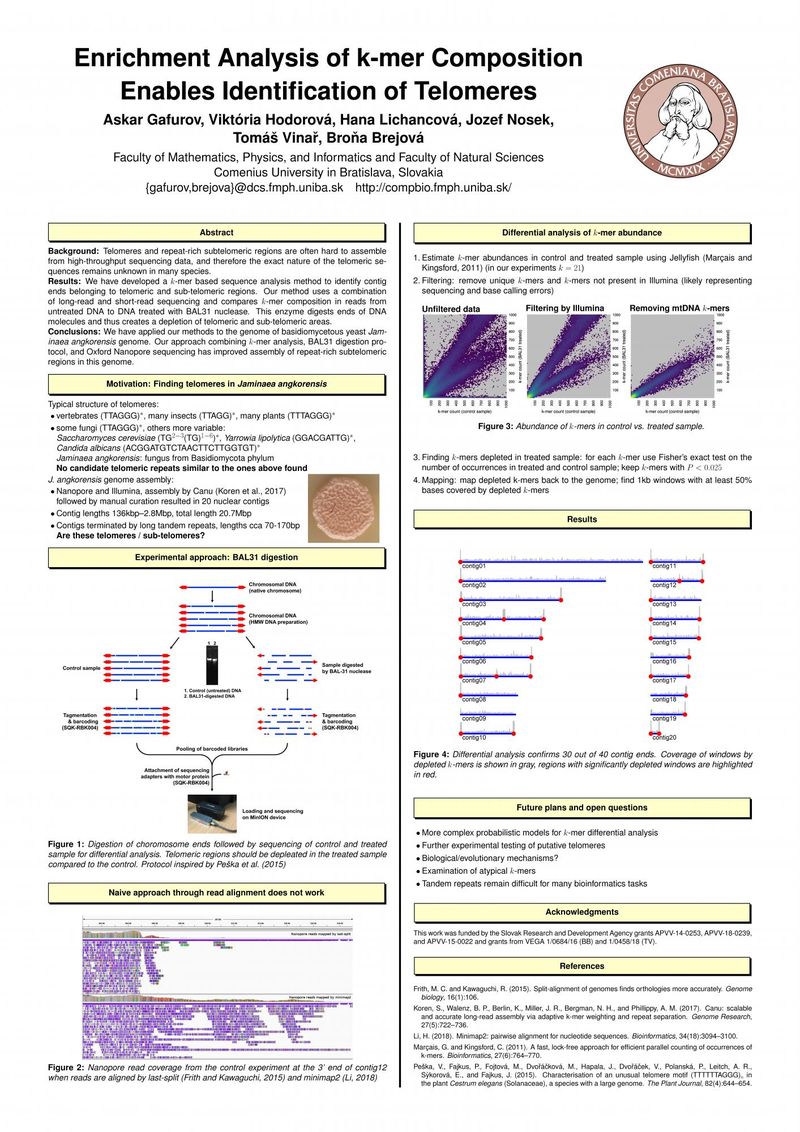
Background: Telomeres and repeat-rich subtelomeric regions are often hard to assemble from high-throughput sequencing data, and therefore the exact nature of the telomeric sequences remains unknown in many species.
Results: We have developed a k-mer based sequence analysis method to identify contig ends belonging to telomeric and sub-telomeric regions. Our method uses a combination of long-read and short-read sequencing and compares k-mer composition in reads from untreated DNA to DNA treated with BAL31 nuclease. This enzyme digests ends of DNA molecules and thus creates a depletion of telomeric and sub-telomeric areas.
Conclusions: We have applied our methods to the genome of basidiomycetous yeast Jaminaea angkorensis genome. Our approach combining k-mer analysis, BAL31 digestion protocol, and Oxford Nanopore sequencing has improved assembly of repeat-rich subtelomeric regions in this genome.






