UNPUBLISHED Ligation sequencing gDNA - automated Hamilton NGS STAR 96 (SQK-LSK110-XL)
- Home
- Documentation
- UNPUBLISHED Ligation sequencing gDNA - automated Hamilton NGS STAR 96 (SQK-LSK110-XL)
Protocol
UNPUBLISHED Ligation sequencing gDNA - automated Hamilton NGS STAR 96 (SQK-LSK110-XL) Vauto_gDNA_rev2
- This protocol uses genomic DNA
- Automation of library preparation
- Increased reproducibility and speed
- Reduces human error
- High sequencing output
- Library preparation time ~1.5 hours hands-on-time and ~3-4 hours automation time
- Fragmentation is optional
- No PCR required
- Suitable for processing multiple samples simultaneously
- Compatible with R10.3 flow cells
For Research Use Only
FOR RESEARCH USE ONLY
Contents
Introduction to the protocol
Automated library preparation
- 4. DNA repair and end-prep
- 5. Adapter ligation and clean-up
- 6. Priming and loading multiple flow cells on a PromethION
Secuenciación y análisis de datos
- 7. Data acquisition and basecalling
- 8. Reutilización y devoluciones de las celdas de flujo
- 9. Análisis posterior
Troubleshooting
Overview
- This protocol uses genomic DNA
- Automation of library preparation
- Increased reproducibility and speed
- Reduces human error
- High sequencing output
- Library preparation time ~1.5 hours hands-on-time and ~3-4 hours automation time
- Fragmentation is optional
- No PCR required
- Suitable for processing multiple samples simultaneously
- Compatible with R10.3 flow cells
For Research Use Only
1. Overview of the protocol
ATENCIÓN
This protocol is currently UNPUBLISHED
UNPUBLISHED protocols are for internal development use ONLY, and should NEVER be shared with external users and customers.
In case of doubt, please contact Product Management.
Ligation Sequencing Kit XL features
This kit is recommended for users who:
- want to optimise their sequencing experiment for throughput
- would like to utilise upstream processes such as size selection, whole genome amplification, or enrich for long reads
IMPORTANTE
Optional fragmentation and size selection
By default, the protocol contains no DNA fragmentation step, however in some cases it may be advantageous to fragment your sample. For example, when working with lower amounts of input gDNA (100 ng – 500 ng), fragmentation will increase the number of DNA molecules and therefore increase throughput. Instructions are available in the DNA Fragmentation section of Extraction methods.
Additionally, we offer several options for size-selecting your DNA sample to enrich for long fragments - instructions are available in the Size Selection section of Extraction methods.
Introduction to the automated Ligation Sequencing protocol for gDNA
This protocol describes how to carry out sequencing of a DNA sample using the Ligation Sequencing Kit XL (SQK-LSK110-XL). It is highly recommended that a Lambda control experiment is completed first to become familiar with the technology.
Steps in the sequencing workflow:
Prepare for your experiment You will need to:
- Extract your DNA and check its length, quantity and purity. The quality checks performed during the protocol are essential in ensuring experimental success.
- Ensure you have your sequencing kit, the correct equipment, primed liquid-handling robot and third-party reagents
- Download the software for acquiring and analysing your data
- Check your flow cells to ensure they have enough pores for a good sequencing run
__Library preparation__ You will need to:
- Repair the DNA and prepare the DNA ends for adapter attachment
- Attach sequencing adapters supplied in the kit to the DNA ends
- Prime the flow cell and load your DNA library into the flow cell
Sequencing and analysis You will need to:
- Start a sequencing run using the MinKNOW software which will collect raw data from the device and convert it into basecalled reads
- Start the EPI2ME software and select a workflow to further analysis (this step is optional)
IMPORTANTE
Compatibility of this protocol
This protocol should only be used in combination with:
- Ligation Sequencing Kit XL (SQK-LSK110-XL)
- Control Expansion (EXP-CTL001)
- Flow Cell Wash Kit (EXP-WSH004)
- Sequencing Auxiliary Vials (EXP-AUX002)
- FLO-PRO002 (R9.4.1) flow cells
- FLO-PRO111 (R10.3) flow cells
2. Equipment and consumables
Material
- 1 µg (o 100-200 fmol) de ADNg
- 1.5-3 µg (or 150-300 fmol) high molecular weight genomic DNA for R10.3 flow cells
- Ligation Sequencing Kit XL (SQK-LSK110-XL)
Consumibles
- Agencourt AMPure XP beads (Beckman Coulter, A63881)
- NEBNext® Companion Module for Oxford Nanopore Technologies® Ligation Sequencing (NEB E7180S or E7180L) (módulo de acompañamiento NEBNext de secuenciación por ligación para Oxford Nanopore Technologies®) Como alternativa, se pueden utilizar los siguientes productos de NEBNext®:
- NEBNext FFPE Repair Mix (NEB M6630) (mezcla de reparación de ADN)
- NEBNext Ultra II End Repair/dA-tailing Module (NEB E7546) (Módulo de reparación de extremos/Adición de dA)
- NEBNext Quick Ligation Module (NEB E6056) (Módulo de ligación rápida)
- Agua sin nucleasas (p. ej., ThermoFisher AM9937)
- Etanol al 80 % recién preparado con agua sin nucleasas
- Hamilton 50 µl CO-RE tips with filter (Cat# 235948)
- Hamilton 300 µl CO-RE tips with filter (Cat# 235903)
- Hamilton 1000 µl CO-RE tips with filter (Cat# 235905)
- Hamilton 60 ml Reagent Reservoir, Self-Standing with Lid (Cat# 56694-01)
- Hamilton PCR ComfortLid (Cat# 814300)
- Bio-Rad Hard-Shell® 96-Well PCR Plates (Cat# HSP9601)
- Roche Diagnotics MagNA Pure LC Medium Reagent Tubs 20 (Cat# 03004058001)
- Sarstedt Inc Screw Cap Micro tube 2 ml, PP 1000/case (e.g. FisherScientific, Cat# NC0418367)
- Thermo Scientific™ Abgene™ 96 Well 0.8 ml Polypropylene Deepwell Storage Plate (Thermo Scientific™, cat # AB0859)
- Qubit dsDNA HS Assay Kit (Invitrogen Q32851) (kit de ensayo ADNbc alta sensibilidad)
- Tubos de ensayo Qubit™ (Invitrogen Q32856)
Instrumental
- Cubeta con hielo
- Mezclador vórtex
- Microplate centrifuge, e.g. Fisherbrand™ Mini Plate Spinner Centrifuge (Fisher Scientific, 11766427)
- Hamilton NGS STAR 96 (NGS STAR with Multi-Probe Head 96)
- Hamilton On-Deck Thermal Cycler (ODTC)
Equipo opcional
- Bioanalizador Agilent (o equivalente)
- Qubit fluorometer plate reader (or equivalent for QC check)
For this protocol, you will need 1 µg (or 100-200 fmol) high molecular weight genomic DNA for every sample to be barcoded if using R9.4.1 flow cells or 1.5-3 µg (or 150-300 fmol) for R10.3 flow cells.
Although 1 µg (or 100-200 fmol) gDNA is recommended, users can start with lower input quantities (down to 100 ng) if performing DNA fragmentation to increase the number of DNA molecules in the sample, or if amplifying the sample by PCR.
Input DNA
How to QC your input DNA
It is important to use a plate reader to ensure the input DNA meets the quantity and quality requirements. Using too little or too much DNA, or DNA of poor quality (e.g. highly fragmented or containing RNA or chemical contaminants) can affect your library preparation.
For instructions on how to perform quality control of your DNA sample, please read the Input DNA/RNA QC protocol.
Input file worklist
A worklist input Excel file is required prior to running the protocol on the Hamilton NGS STAR 96. These contain information regarding the appropriate number of samples and well identifiers.
Example:
Source_SampleID | Source_Well | Target_Well |
|---|---|---|
| Sample_01 | A1 | A1 |
| Sample_02 | B1 | B2 |
| Sample_03 | C1 | C1 |
Hamilton NGS STAR 96
This method has been tested and validated using the Hamilton NGS STAR 96 (with 8 channels and MPH96) including an on deck thermal cycler (ODTC). An option to not use the ODTC is available in the method. The protocol may require some fine tuning for the NGS STAR 96 setup and the temperature/humidity of the customer laboratory.
Please contact your Hamilton representative for further details.
Deck layout 
- ODTC: On-Deck Thermal Cycler Module
- MIDI plates: Abgene™ 96 Well 0.8mL Polypropylene Deepwell Storage Plate
- Troughs: Hamilton 20 ml Reagent Reservoirs
- HSP plates: Bio-Rad Hard-Shell® 96-Well PCR Plates
- Hamilton ComfortLid: Hamilton PCR ComfortLid
NEBNext® Companion Module para secuenciación por ligación, de Oxford Nanopore Technologies®
A los clientes que no conozcan la secuenciación por nanoporos, se les recomienda que adquieran el módulo de acompañamiento NEBNext® Ligation Sequencing para Oxford Nanopore Technologies® (NEB E7180S o E7180L), que contiene todos los reactivos NEB necesarios para usar con el kit Ligation Sequencing Kit.
Nótese que en los protocolos con amplicones no es necesario usar ni la mezcla de reparación de ADN NEBNext FFPE DNA Repair Mix, ni el tampón de reparación de ADN NEBNext FFPE DNA Repair Buffer.
Consumables quantities:
| Consumables | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Hamilton 50 µl CO-RE tips with filter | 216 | 432 | 768 |
| Hamilton 300 µl CO-RE tips with filter | 451 | 694 | 1176 |
| Hamilton 1000 µl CO-RE tips with filter | 64 | 64 | 64 |
| Hamilton 60 ml Reagent Reservoir, Self-Standing with Lid | 3 | 3 | 3 |
| Hamilton PCR ComfortLid | 1 | 1 | 1 |
| Bio-Rad Hard-Shell® 96-Well PCR Plate | 2 | 2 | 2 |
| Roche Diagnostics MagNA Pure LC Medium Reagent Tubs 20 | 2 | 2 | 2 |
| Sarstedt Inc Screw Cap Micro Tube 2ml | 2 | 4 | 5 |
| Abgene™ 96 Well 0.8mL Polypropylene Deepwell Storage Plate | 2 | 2 | 2 |
| Kits | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Ligation Sequencing Kit XL (SQK-LSK110-XL) | 1 kit | 1 kit | 2 kits |
| NEBNext Companion Module for Oxford Nanopore Technologies Ligation Sequencing (Cat# E7180S) | 2 kits | 3 kits | 5 kits |
| Alternatively: | - | - | - |
| NEBNext FFPE DNA Repair Mix (Cat# M6630L) | 1 kit | 1 kit | 2 kits |
| NEBNext Ultra II End Repair/dA-Tailing Module (Cat# E7546L) | 1 kit | 2 kits | 3 kits |
| NEBNext Quick Ligation Module (Cat# E6056L) | 1 kit | 2 kits | 3 kits |
Note: These are the number of kits required for one run through for the selected number of samples.
Ligation Sequencing Kit XL (SQK-LSK110-XL) contents
| Name | Acronym | Cap colour | Number of vials | Fill volume per vial (µl) |
|---|---|---|---|---|
| Adapter Mix F | AMX F | Green | 1 | 320 |
| Sequencing Buffer | SBII | 10 ml bottle | 1 | 5,000 |
| Ligation Buffer | LNB | White | 1 | 1,500 |
| S Fragment Buffer | SFB | 10 ml bottle | 4 | 7,500 |
| L Fragment Buffer | LFB | 10 ml bottle | 4 | 7,500 |
| DNA CS | DCS | Yellow | 1 | 100 |
| Elution Buffer | EB | 10 ml bottle | 1 | 10,000 |
| Loading beads II | LBII | Pink | 2 | 1,500 |
| Loading solution | LS | White cap, pink sticker | 2 | 1,500 |
| Flush buffer XL | FB | 20 ml bottle | 4 | 15,500 |
| Flush tether | FLT | White cap, purple sticker | 1 | 1,600 |
3. Computer requirements and software
PromethION 24/48 IT requirements
The PromethION device contains all the hardware required to control up to 24 (for the P24 model) or 48 (for the P48 model) sequencing experiments and acquire the data. The device is further enhanced with high performance GPU technology for real-time basecalling. Read more in the PromethION IT Requirements document.
PromethION 2 Solo IT requirements
The PromethION 2 (P2) Solo is a device which directly connects into a GridION Mk1 or a stand-alone computer that meets the miminum specifications for real-time data streaming and analysis. Up to two PromethION flow cells can be can be run and each is independently addressable, meaning experiments can be run concurrently or individually. For information on the computer IT requirements, please see the PromethION 2 Solo IT requirements document.
Software for nanopore sequencing
MinKNOW
The MinKNOW software controls the nanopore sequencing device, collects sequencing data and basecalls in real time. You will be using MinKNOW for every sequencing experiment to sequence, basecall and demultiplex if your samples were barcoded.
For instructions on how to run the MinKNOW software, please refer to the MinKNOW protocol.
EPI2ME (optional)
The EPI2ME cloud-based platform performs further analysis of basecalled data, for example alignment to the Lambda genome, barcoding, or taxonomic classification. You will use the EPI2ME platform only if you would like further analysis of your data post-basecalling.
For instructions on how to create an EPI2ME account and install the EPI2ME Desktop Agent, please refer to the EPI2ME Platform protocol.
Verificar la celda de flujo
Antes de empezar el experimento de secuenciación, recomendamos verificar el número de poros disponibles, presentes en la celda de flujo. La comprobación deberá realizarse en los primeros tres meses desde su adquisición, si se trata de celdas de flujo MinION, GridION o PromethION, y en las primeras cuatro semanas tras la compra de celdas de flujo Flongle. Oxford Nanopore Technologies sustituirá cualquier celda de flujo con un número de poros inferior al indicado en la tabla siguiente, siempre y cuando el resultado se notifique dentro de los dos días siguientes a la comprobación y se hayan seguido las instrucciones de almacenamiento. Para verificar la celda de flujo, siga las instrucciones del documento Flow Cell Check.
| Celda de flujo | Número mínimo de poros activos cubierto por la garantía |
|---|---|
| Flongle | 50 |
| MinION/GridION | 800 |
| PromethION | 5000 |
4. DNA repair and end-prep
Material
- 1.5-3 µg (or 150-300 fmol) high molecular weight genomic DNA for R10.3 flow cells
Consumibles
- Agua sin nucleasas (p. ej., ThermoFisher AM9937)
- NEBNext FFPE DNA Repair Mix (NEB M6630)
- NEBNext Ultra II End repair/dA-tailing Module (NEB E7546)
- Agencourt AMPure XP beads (Beckman Coulter™, A63881)
- Etanol al 80 % recién preparado con agua sin nucleasas
- Thermo Scientific™ Abgene™ 96 Well 0.8 ml Polypropylene Deepwell Storage Plate (Thermo Scientific™, cat # AB0859)
- Sarstedt Inc Screw Cap Micro tube 2 ml, PP 1000/case (e.g. FisherScientific, Cat# NC0418367)
- Roche Diagnotics MagNA Pure LC Medium Reagent Tubs 20 (Cat# 03004058001)
- Bio-Rad Hard-Shell® 96-Well PCR Plates (Cat# HSP9601)
- Hamilton PCR ComfortLid (Cat# 814300)
- Hamilton 60 ml Reagent Reservoir, Self-Standing with Lid (Cat# 56694-01)
- Hamilton 1000 µl CO-RE tips with filter (Cat# 235905)
- Hamilton 300 µl CO-RE tips with filter (Cat# 235903)
- Hamilton 50 µl CO-RE tips with filter (Cat# 235948)
- Qubit dsDNA HS Assay Kit (Invitrogen Q32851) (kit de ensayo ADNbc alta sensibilidad)
- Tubos de ensayo Qubit™ (Invitrogen Q32856)
Instrumental
- Cubeta con hielo
- P1000 pipette and tips
- Pipeta y puntas P200
- Pipeta y puntas P100
- Pipeta y puntas P10
- Microplate centrifuge, e.g. Fisherbrand™ Mini Plate Spinner Centrifuge (Fisher Scientific, 11766427)
- Mezclador vórtex
- Qubit fluorometer (or equivalent)
Consumables and equipment quantities:
| Consumable/equipment | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Hamilton 50 µl CO-RE tips with filter | 96 | 192 | 384 |
| Hamilton 300 µl CO-RE tips with filter | 201 | 298 | 490 |
| Hamilton 1000 µl CO-RE tips with filter | 32 | 32 | 32 |
| Hamilton 60 ml Reagent Reservoir, Self-Standing with Lid | 2 (1 EtOH & H20) | 2 (1 EtOH & H20) | 2 (1 EtOH & H20) |
| Hamilton PCR ComfortLid | 1 | 1 | 1 |
| Bio-Rad Hard-Shell® 96-Well PCR Plate | 1 (1 input sample & 1 end prepped sample) | 1 (1 input sample & 1 end prepped sample) | 1 (1 input sample & 1 end prepped sample) |
| Hamilton 20 ml Reagent Reservoirs | 1 | 1 | 1 |
| Sarstedt Inc Screw Cap Micro Tube 2 ml | 1 | 2 | 2 |
| Abgene™ 96 Well 0.8 ml Polypropylene Deepwell Storage Plate | 1 | 1 | 1 |
Reagents quantities:
| Reagents | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| 80% ethanol | 16.5 ml | 28 ml | 51 ml |
| AMPure XP Beads | 3.7 ml | 5.4 ml | 8.9 ml |
| NEBNext Companion Module for Oxford Nanopore Technologies Ligation Sequencing (Cat# E7180S) regarding the reagents below | 2 tubes | 3 tubes | 5 tubes |
| Alternatively: | - | - | - |
| NEBNext FFPE DNA Repair Buffer | 1 tube | 1 tube | 1 tube |
| NEBNext FFPE DNA Repair Mix | 1 tube | 1 tube | 2 tubes |
| Ultra II End-prep Reaction Buffer | 1 tube | 2 tubes | 3 tubes |
| Ultra II End-prep Enzyme Mix | 1 tube | 1 tube | 2 tubes |
Note: Dead volumes are included.
IMPORTANTE
Optional fragmentation and size selection
By default, the protocol contains no DNA fragmentation step, however in some cases it may be advantageous to fragment your sample. For example, when working with lower amounts of input gDNA (100 ng – 500 ng), fragmentation will increase the number of DNA molecules and therefore increase throughput. Instructions are available in the DNA Fragmentation section of Extraction methods.
Additionally, we offer several options for size-selecting your DNA sample to enrich for long fragments - instructions are available in the Size Selection section of Extraction methods.
Preparar los reactivos NEBNext FFPE DNA Repair Mix y NEBNext Ultra II End Repair / dA-tailing Module siguiendo las instrucciones del fabricante y poner en hielo.
Para obtener un rendimiento óptimo, NEB recomienda lo siguiente:
Descongelar todos los reactivos en hielo.
Golpear suavemente los tubos de reactivos con el índice o invertirlos, para asegurarse de que estén bien mezclados.
Nota: No mezclar en vórtex las mezclas FFPE DNA Repair Mix, ni Ultra II End Prep Enzyme Mix.Centrifugar los tubos antes de abrirlos.
Los tampones Ultra II End Prep Buffer y FFPE DNA Repair Buffer pueden tener un poco de precipitado. Dejar que la mezcla alcance la temperatura ambiente y mezclar pipeteando varias veces para romper el precipitado; para solubilizarlo, agitar el tubo en vórtex durante 30 s.
Nota: Es importante mezclar bien los tampones mediante vórtex.El tampón FFPE DNA Repair Buffer puede tener un matiz amarillo; no importa si está así; se puede utilizar.
Prepare each DNA sample per well with nuclease-free water in the input plate.
- Per sample, transfer 1 μg (or 100-200 fmol) of genomic DNA into a well of the input plate
- Adjust the volume to 48 μl with nuclease-free water
- Mix thoroughly by pipetting
- Spin down briefly in a microfuge
Quantify 1 µl of each eluted sample using a Qubit fluorometer plate reader off deck.
Switch on the Hamilton NGS STAR 96 robot and open 'Hamilton Run Control' on the computer by clicking the icon:
![]()
Click 'File' and 'Open' to choose the method to run on the liquid handling robot.
Click 'Process01: DNA repair and end-prep' to start.
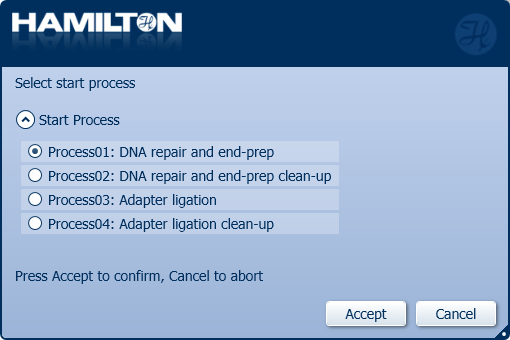
Click 'Process02: DNA repair and end-prep clean-up' to stop the automated library preparation and quantify the samples before the adapter ligation step.
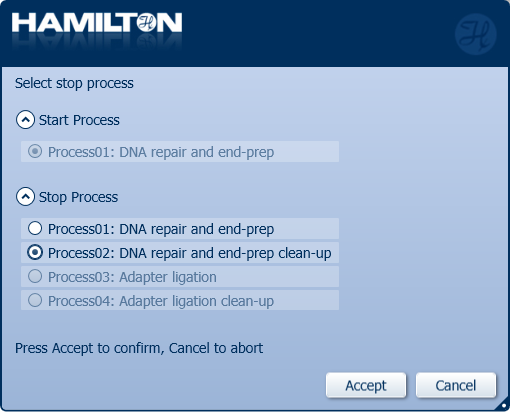
IMPORTANTE
It is mandatory for users to have an MPH module installed and we recommend the use of an ODTC module.
Select whether an ODTC module is available to use in the run and select Yes to use the MPH (96 Head) module.

Click 'Browse' to choose the Input File Worklist for the specific number of samples in the run and click 'OK'.
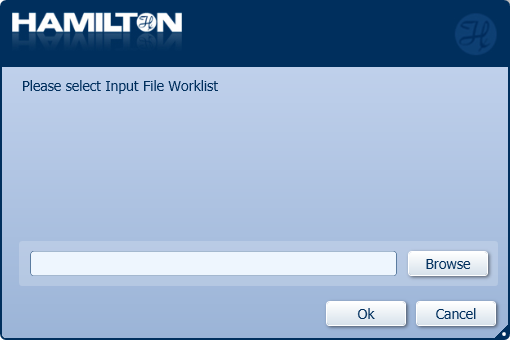
An input file worklist for the number of samples in the run must be generated before the run. Example of an input file worklist:
Source_SampleID | Source_Well | Target_Well |
|---|---|---|
| Sample_01 | A1 | A1 |
| Sample_02 | B1 | B1 |
| Sample_03 | C1 | C1 |
Prepare the End Prep Mastermix with the following reagents according to the Hamilton user interface. Click either 'Yes' or 'No' to continue.
Note: It is user preference whether to print and save the instructions.
Reagent volumes for all sample numbers:
| Reagent | Volume X24 samples | Volume X48 samples | Volume X96 samples |
|---|---|---|---|
| NEBNext FFPE DNA Repair Buffer | 106.6 µl | 213.3 µl | 414.8 µl |
| NEBNext FFPE DNA Repair Mix | 60.9 µl | 121.8 µl | 237 µl |
| Ultra II End-prep Reaction Buffer | 106.6 µl | 213.3 µl | 414.8 µl |
| Ultra II End-prep Enzyme Mix | 91.4 µl | 182.8 µl | 355.6 µl |
| Total | 365.6 µl | 731.2 µl | 1422.2 µl |
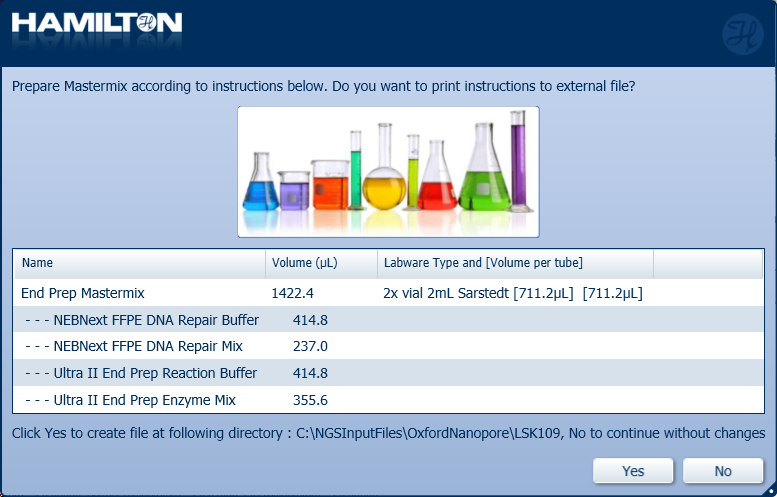
Insert the ComfortLid position as displayed on screen. Click 'Ok' to continue.
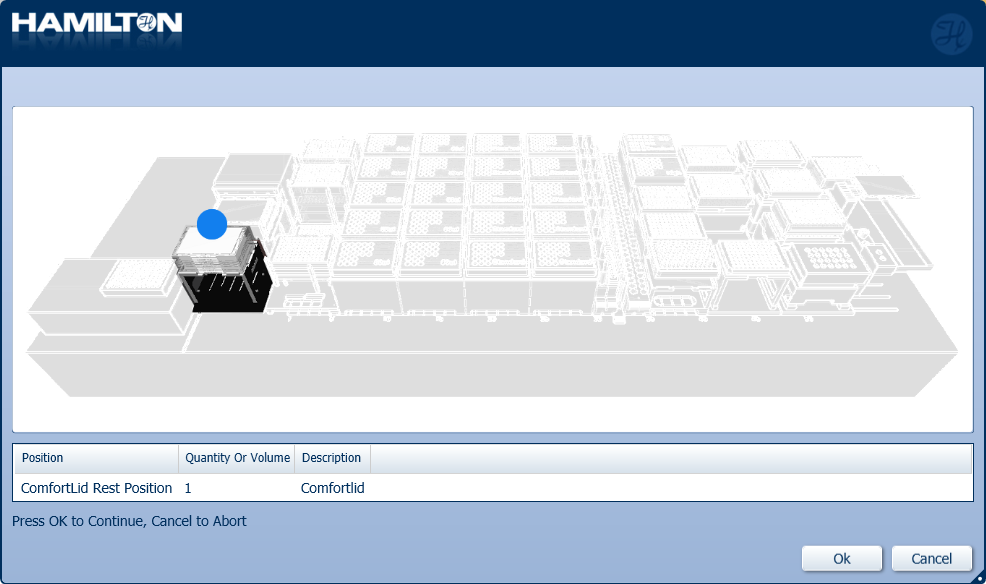
Insert plates to their corresponding positions. Click 'Ok' to continue.
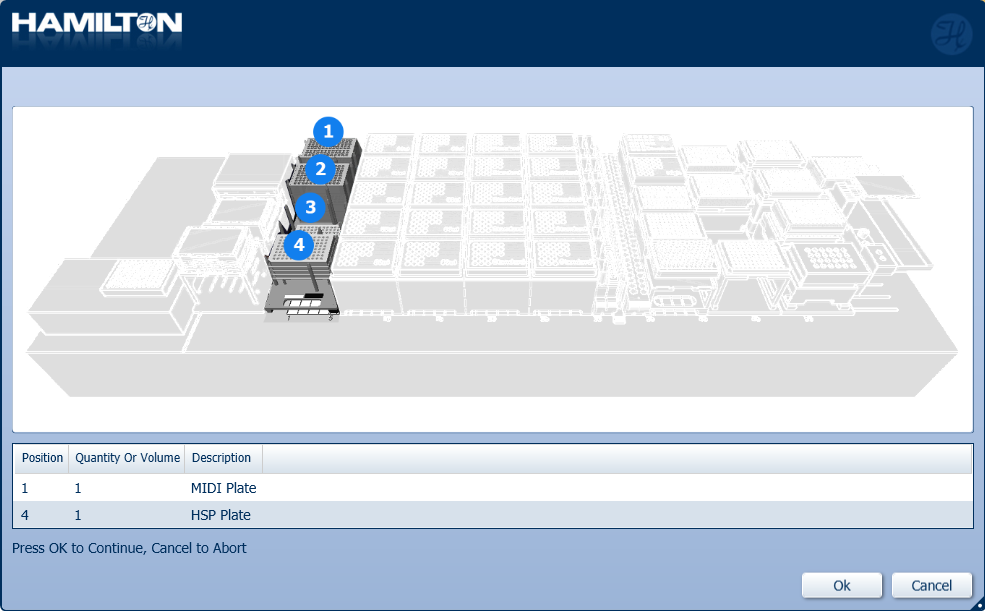
Load a full deck of 50 µl tips into the positions on screen. Click 'Ok' to continue.
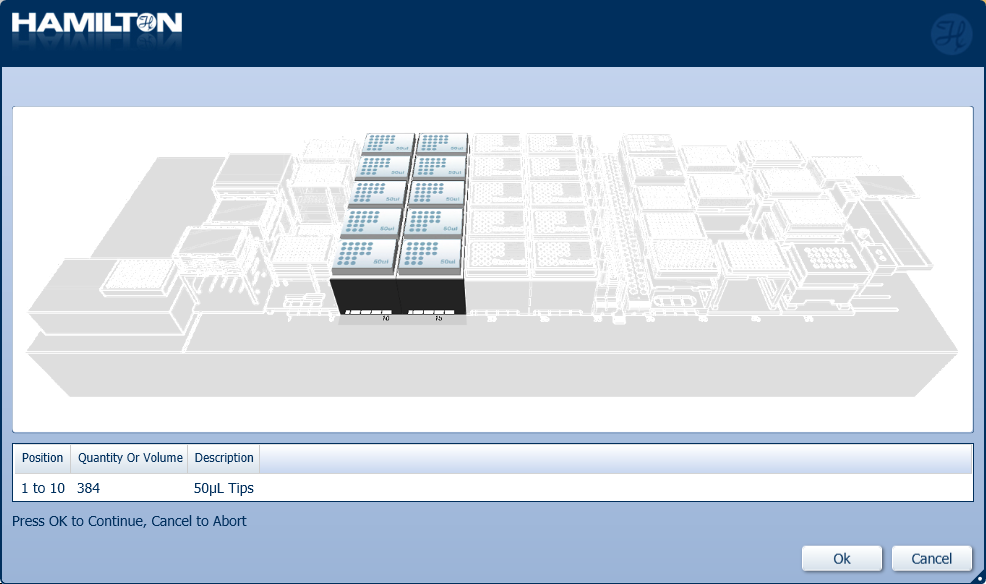
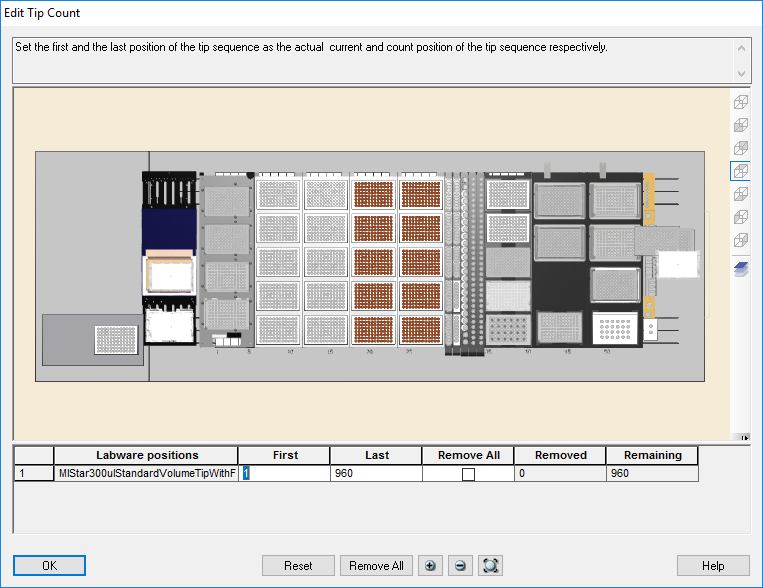
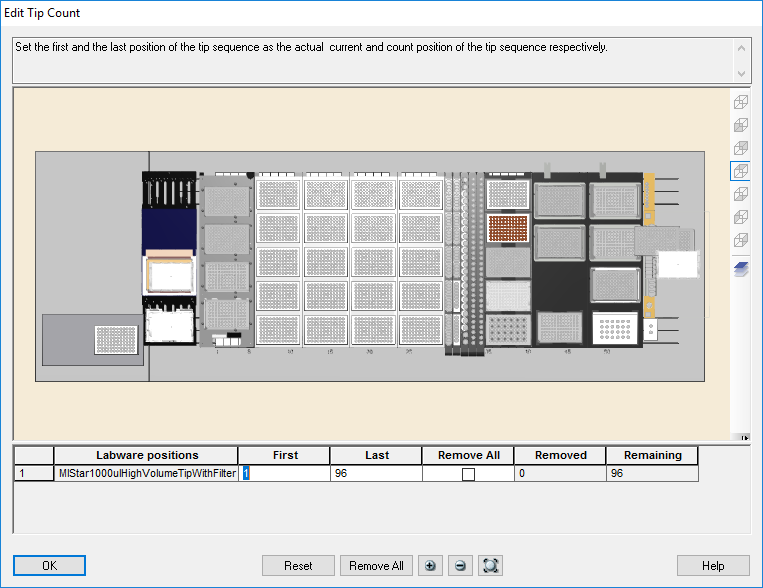
Highlight the 50 µl tips available to use on the 'Edit Tip Count' window and click 'Ok' to continue.
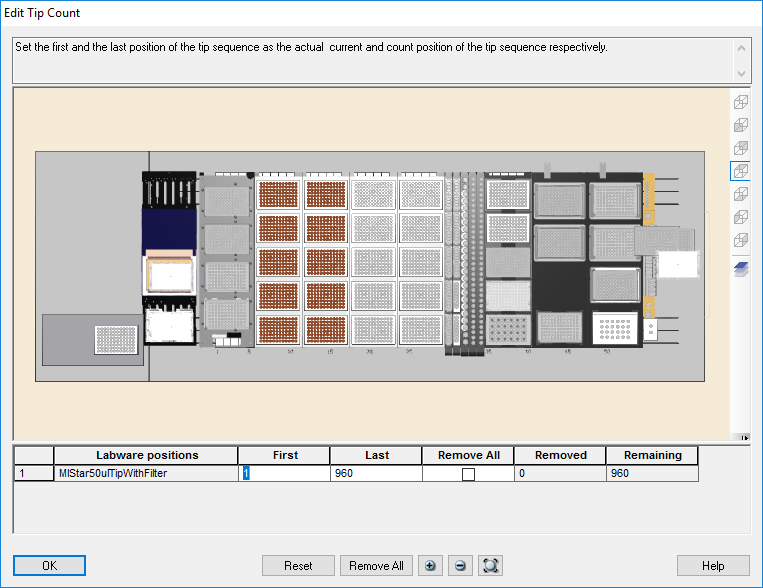
Load a full deck of 300 µl tips in the positions on screen. Click 'Ok' to continue.
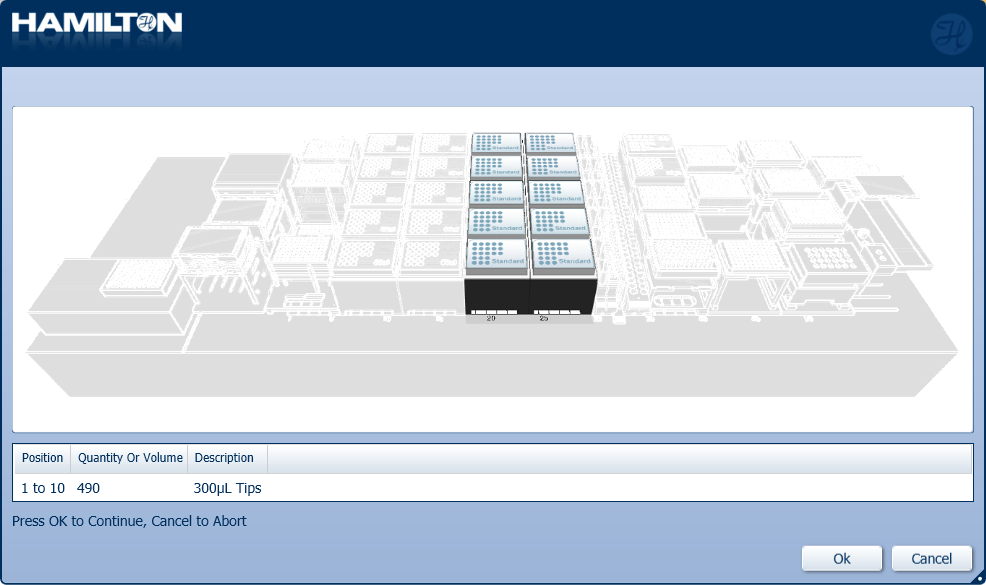
Highlight the 300 µl tips available to use on the 'Edit Tip Count' window and click 'Ok' to continue.
Freshly prepare 80% ethanol in nuclease-free water in a trough.
| Reagents | Volume X24 samples | Volume X48 samples | Volume X96 samples |
|---|---|---|---|
| 80% ethanol | 16.5 ml | 28 ml | 51 ml |
Insert the trough of 80% ethanol in the position on screen and click 'Ok' to continue.
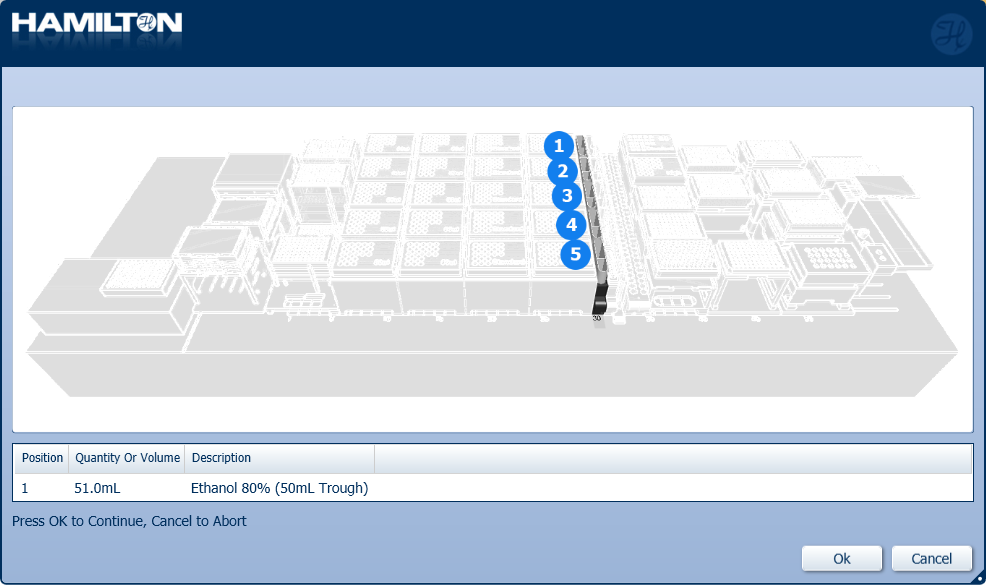
CONSEJO
If the consumables used for troughs are not barcoded, click 'Exclude' on all the selected troughs inserted in the robot and click 'Execute' to continue.

Prepare the AMPure XP beads by vortexing and load the 20 ml trough with the volume required:
| Reagents | Volume X24 samples | Volume X48 samples | Volume X96 samples |
|---|---|---|---|
| Beads | 3.7 ml | 5.4 ml | 8.9 ml |
IMPORTANTE
Ensure the AMPure XP beads are well mixed before use by vortexing.
Insert the trough of AMPure XP beads and nuclease-free water in their positions on screen. Click 'Ok' to continue.
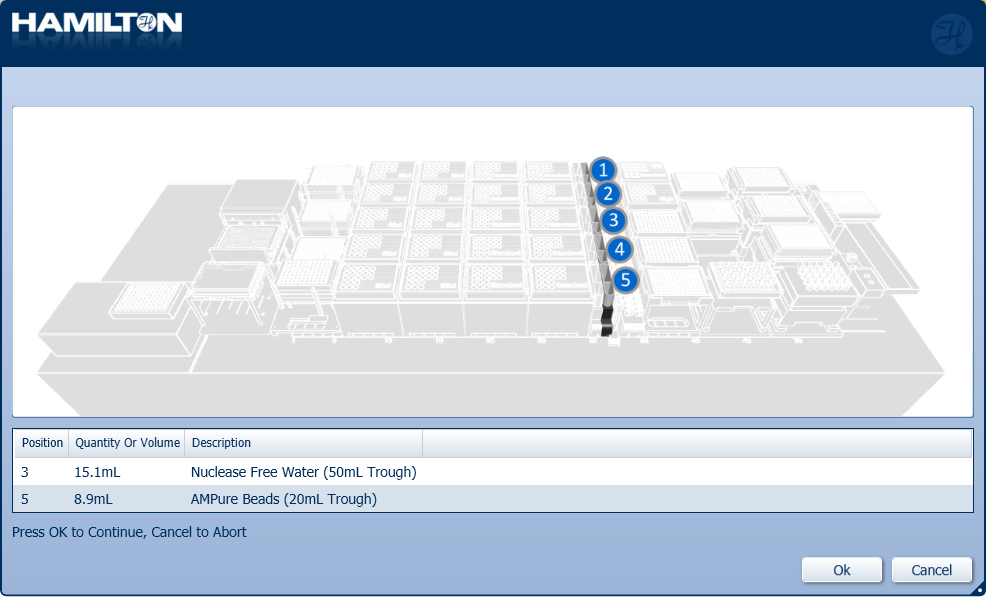
CONSEJO
If the consumables used for troughs are not barcoded, click 'Exclude' on all the selected troughs inserted in the robot and click 'Execute' to continue.

Load 1000 µl tips and insert the input plate of DNA samples into the position on screen. Click 'Ok' to continue.
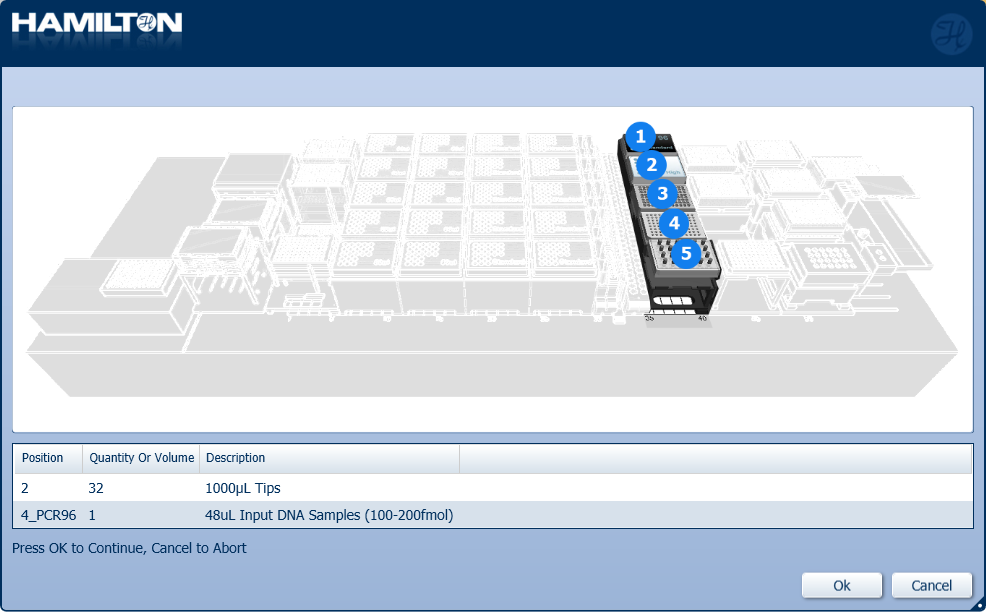
Highlight the 1000 µl tips available to use on the 'Edit Tip Count' window and click 'Ok' to continue.
ATENCIÓN
Ensure the mastermix is well mixed and homogenous before loading the 2 ml Sarstedt tubes. Mixing in the robot is not effective.
Mix and insert the prepared End Prep Mastermix into the positions on screen.
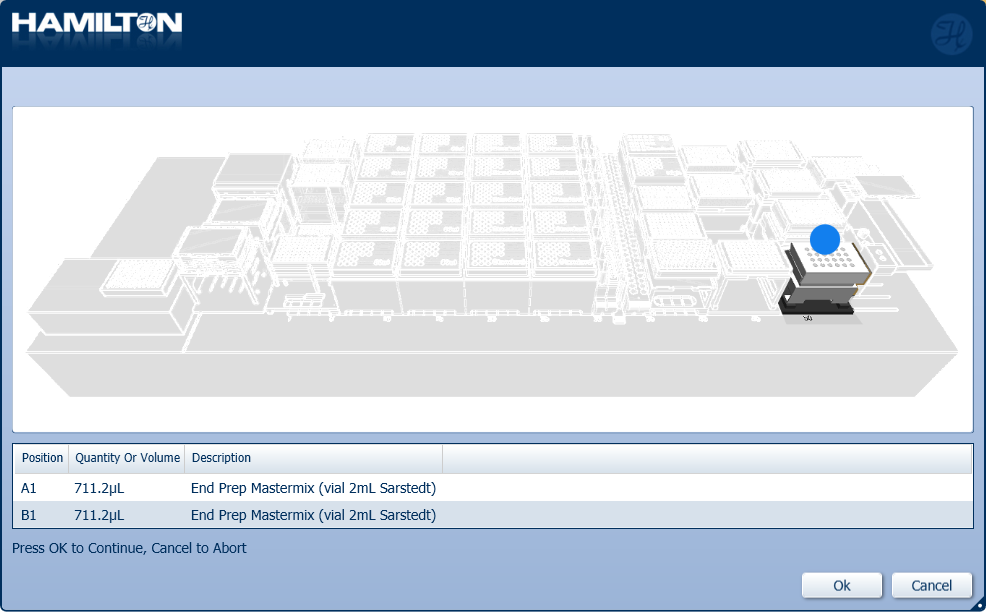
Click 'Ok' to start the DNA repair and end-prep automation process.
Once the automation process has finished, there will be an on screen prompt to unload the plate. Click 'Ok' to continue.

Quantify 1 µl of each eluted sample using a Qubit fluorometer plate reader off deck.
FIN DEL PROCESO
Take forward the repaired and end repaired DNA into the adapter ligation and clean-up step.
5. Adapter ligation and clean-up
Material
- Adapter Mix F (AMX F)
- Ligation Buffer (LNB) (tampón de ligación) del kit Ligation Sequencing Kit
- Long Fragment Buffer (LFB) (tampón para fragmentos largos)
- Short Fragment Buffer (SFB)
- Elution Buffer (EB) (tampón de elución) del kit de Oxford Nanopore
Consumibles
- Agencourt AMPure XP beads (Beckman Coulter™, A63881)
- NEBNext Quick Ligation Module (NEB E6056) (Módulo de ligación rápida)
- Thermo Scientific™ Abgene™ 96 Well 0.8 ml Polypropylene Deepwell Storage Plate (Thermo Scientific™, cat # AB0859)
- Sarstedt Inc Screw Cap Micro tube 2 ml, PP 1000/case (e.g. FisherScientific, Cat# NC0418367)
- Roche Diagnotics MagNA Pure LC Medium Reagent Tubs 20 (Cat# 03004058001)
- Bio-Rad Hard-Shell® 96-Well PCR Plates (Cat# HSP9601)
- Hamilton 60 ml Reagent Reservoir, Self-Standing with Lid (Cat# 56694-01)
- Hamilton 1000 µl CO-RE tips with filter (Cat# 235905)
- Hamilton 300 µl CO-RE tips with filter (Cat# 235903)
- Hamilton 50 µl CO-RE tips with filter (Cat# 235948)
Instrumental
- P1000 pipette and tips
- Pipeta y puntas P200
- Pipeta y puntas P100
- Pipeta y puntas P10
- Vortex mixer
- Microplate centrifuge, e.g. Fisherbrand™ Mini Plate Spinner Centrifuge (Fisher Scientific, 11766427)
Consumables and equipment quantities:
| Consumable/equipment | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Hamilton 50 µl CO-RE tips with filter | 120 | 240 | 384 |
| Hamilton 300 µl CO-RE tips with filter | 250 | 396 | 686 |
| Hamilton 1000 µl CO-RE tips with filter | 32 | 32 | 32 |
| Hamilton 60 ml Reagent Reservoir, Self-Standing with Lid | 2 (1 L/SFB & 1 EB) | 2 (1 L/SFB & 1 EB) | 2 (1 L/SFB & 1 EB) |
| Bio-Rad Hard-Shell® 96-Well PCR Plate | 1 | 1 | 1 |
| Hamilton 20 ml Reagent Reservoirs | 1 | 1 | 1 |
| Sarstedt Inc Screw Cap Micro Tube 2 ml | 1 | 2 | 3 |
| Abgene™ 96 Well 0.8 ml Polypropylene Deepwell Storage Plate | 1 | 1 | 1 |
Reagents quantities:
| Reagents | X24 samples | X48 samples | X96 samples |
|---|---|---|---|
| Adapter Mix F (AMX-F) | 0.5 tube | 1 tube | 2 tubes |
| Ligation Buffer (LNB) | 0.5 tube | 1 tube | 2 tubes |
| Elution Buffer (EB) | 0.5 bottle | 1 bottle | 2 bottles |
| Long Fragment Buffer (LFB) | 2 bottles | 4 bottles | 8 bottles |
| Short Fragment Buffer (SFB) | 2 bottles | 4 bottles | 8 bottles |
| AMPure XP Beads | 3.1 ml | 4.3 ml | 6.6 ml |
| NEBNext Companion Module for Oxford Nanopore Technologies Ligation Sequencing (Cat# E7180S) regarding the reagent below: | 2 tubes | 3 tubes | 5 tubes |
| Alternatively: | - | - | - |
| Quick T4 DNA Ligase | 1 tube | 2 tubes | 3 tubes |
Note: Dead volumes are included.
IMPORTANTE
Although the recommended 3rd party ligase is supplied with its own buffer, the ligation efficiency of Adapter Mix F (AMX-F) is higher when using Ligation Buffer supplied within the Ligation Sequencing Kit.
Spin down and store the Quick T4 Ligase on ice until use.
Spin down and combine all the required tubes of Adapter Mix F (AMX-F) required, and place on ice.
Thaw the Ligation Buffer (LNB) at room temperature, spin down and combine all the required tubes. Place on ice immediately after thawing and mixing.
Thaw a bottle of Elution Buffer (EB) at room temperature, mix by vortexing and place on ice.
IMPORTANTE
La fase de lavados tras la ligación de los adaptadores está diseñada para enriquecer los fragmentos de ADN de >3 kb o para purificar todos los fragmentos por igual, según el tampón que se utilice -Long Fragment Buffer (LFB) o Short Fragment Buffer (SFB).
Para enriquecer fragmentos de ADN de 3 kb o mayores, utilizar el tampón para fragmentos largos, Long Fragment Buffer (LFB).
Para conservar fragmentos de ADN de todos los tamaños, utilizar el tampón para fragmentos cortos, Short Fragment Buffer (SFB).
To enrich for DNA fragments of 3 kb or longer, thaw the Long Fragment Buffer (LFB) at room temperature, mix by vortexing and combine all the required bottles before storing on ice.
To retain DNA fragments of all sizes, thaw the Short Fragment Buffer (SFB) at room temperature, mix by vortexing and combine all the required bottles before storing on ice.
Click 'Process03: Adapter ligation' to start.

Click 'Process04: Adapter ligation and clean-up' to stop the automated library preparation and quantify the samples before sequencing.

IMPORTANTE
It is mandatory for the MPH module to be installed on the liquid handling robot. Select 'Yes' to use the MPH (96 Head) module.

Click 'Browse' to choose the Input File Worklist used during DNA repair and end-prep.

Prepare the Adapter Ligation Mastermix with the following reagents in 2 ml Sarstedt tubes according to the Hamilton user interface. Click either 'Yes' or 'No' to continue.
Note: It is user preference whether to print and save the instructions.
Reagent volumes for all sample numbers:
| Reagent | Volume X24 samples | Volume X48 samples | Volume X96 samples |
|---|---|---|---|
| Adapter Mix F (AMX-F) | 140.5 µl | 281 µl | 559.5 µl |
| Ligation Buffer (LNB) | 702.5 µl | 1405 µl | 2797.5 µl |
| Quick T4 DNA Ligase | 281 µl | 562 µl | 1119 µl |

ATENCIÓN
Ensure the mastermix is well mixed and homogenous before loading the 2 ml Sarstedt tubes. Mixing in the robot is not effective.
Insert plates to their corresponding positions on screen. Click 'Ok' to continue.

Load a full deck of 50 µl tips into the positions on screen. Click 'Ok' to continue.

Highlight the 50 µl tips available to use on the 'Edit Tip Count' window. Click 'Ok' to continue.

Load a full deck of 300 µl tips in the positions on screen. Click 'Ok' to continue.

Highlight the 300 µl tips available to use on the 'Edit Tip Count' window. Click 'Ok' to continue.

Prepare the AMPure XP beads by vortexing and load the 20 ml trough with the volume required:
| Reagents | Volume X24 samples | Volume X48 samples | Volume X96 samples |
|---|---|---|---|
| Beads | 3.1 ml | 4.3 ml | 6.6 ml |
IMPORTANTE
Ensure the AMPure XP beads are well mixed before use by vortexing.
Prepare troughs of L/SFB and EB in troughs and insert in the positions on screen with the trough of AMPure Beads. Click 'Ok' to continue.
| Reagent | Volume X24 samples | Volume X48 samples | Volume X96 samples |
|---|---|---|---|
| Long/Short Fragment Buffer | 2 bottles | 4 bottles | 8 bottles |
| Elution Buffer | 0.5 bottle | 1 bottle | 2 bottles |

CONSEJO
If the troughs are not barcoded, click 'Exclude' on all the selected troughs inserted in the robot and click 'Execute' to continue.

Insert 1000 µl tips and the Clean End Prep Plate to the correct positions on screen. Click 'Ok' to continue.

Highlight the 1000 µl tips available to use on the 'Edit Tip Count' window. Click 'Ok' to continue.

Insert the prepared Adapter Ligation Mastermix into the positions on screen. Click 'Ok' to continue.

ATENCIÓN
Ensure the mastermix is well mixed and homogenous before loading the 2 ml Sarstedt tubes. Mixing in the robot is not effective.
Once the automation process has finished, there will be an on screen prompt to unload the plate. Click 'Ok' to continue.

Quantify 1 µl of each eluted sample using a Qubit fluorometer plate reader off deck.
FIN DEL PROCESO
Seal the plate once the library is prepared and store on ice until ready to load onto the flow cell.
We do not recommend running the liquid handling robot overnight as the plate must be sealed and stored on ice as soon as library preparation is finished.
CONSEJO
Recomendaciones de guardado de la biblioteca
Se recomienda guardar las bibliotecas en tubos Eppendorf DNA LoBind a 4 ⁰C, durante periodos de tiempo cortos o en caso de uso repetido, por ejemplo, para recargar celdas de flujo entre lavados. Para uso individual y para conservar a largo plazo por periodos de más de 3 meses, se recomienda guardar las bibliotecas a -80 ⁰C en tubos Eppendorf DNA LoBind.
IMPORTANTE
We recommend loading 5-50 fmol of the final prepared library onto a flow cell.
Loading more than the maximal recommended amount of DNA can have a detrimental effect on output as higher quantities of DNA results in a larger number of ligated DNA ends with loaded motor protein. This depletes fuel in the Sequencing Buffer, regardless of whether or not the DNA fragments are being sequenced. This leads to fuel depletion and speed drop-off early in the sequencing run. Dilute the libraries in Elution Buffer if required.
If you are using the Flongle for sample prep development, we recommend loading 3-20 fmol instead.
MEDIDA OPCIONAL
If quantities allow, the libraries may be diluted in Elution Buffer (EB) for splitting across multiple flow cells.
Additional buffer for doing this can be found in the Sequencing Auxiliary Vials expansion (EXP-AUX002), available to purchase separately. This expansion also contains additional vials of Sequencing Buffer (SBII) and Loading Beads (LBII), required for loading the libraries onto flow cells.
6. Priming and loading multiple flow cells on a PromethION
Material
- Flush Buffer (FB)
- Flush Tether (FLT)
- Sequencing Buffer II (SBII)
- Loading Beads II (LBII)
- Loading Solution (LS)
Consumibles
- Celda de flujo PromethION
- 1.5 ml Eppendorf DNA LoBind tubes
- 2 ml Eppendorf DNA LoBind tubes
Instrumental
- PromethION 2 Solo device
- Dispositivo PromethION 24/48
- P1000 pipette and tips
- Pipeta y puntas P200
- Pipeta y puntas P20
Using the Loading Solution
We recommend using the Loading Beads II (LBII) for loading your library onto the flow cell for most sequencing experiments. However, if you have previously used water to load your library, you must use Loading Solution (LS) instead of water. Note: some customers have noticed that viscous libraries can be loaded more easily when not using Loading Beads II.
Thaw the Sequencing Buffer II (SBII), Loading Beads II (LBII), Flush Tether (FLT) and on tube of Flush Buffer (FB) at room temperature before mixing the reagents by vortexing and spin down at room temperature.
IMPORTANTE
Scale up reagent volumes as needed.
Ensure to prepare enough reagents for the total number of flow cells being processed and to take into account extra volume required for pipetting errors.
CONSEJO
Each vial provides enough reagent for the preparation of 12 samples. Thaw the appropriate number of vials of each reagent.
Prepare the flow cell priming mix in a suitable vial for the number of flow cells to flush. Once combined, mix well by briefly vortexing.
| Reagent | Volume per flow cell |
|---|---|
| Flush Tether (FLT) | 30 µl |
| Flush Buffer (FB) | 1,170 µl |
IMPORTANTE
Una vez sacadas de la nevera, esperar 20 minutos antes de insertar las celdas de flujo en el dispositivo y así darles tiempo a que estén a temperatura ambiente. En entornos húmedos se puede formar condensación. Inspeccione las clavijas doradas del conector, situadas en la parte superior e inferior de la celda de flujo, en busca de condensación y si la hubiera, límpiela con una toallita sin pelusa. Procure que la almohadilla térmica (color negro) esté enganchada en la parte posterior.
For PromethION 2 Solo, load the flow cell(s) as follows:
Place the flow cell flat on the metal plate.
Slide the flow cell into the docking port until the gold pins or green board cannot be seen.
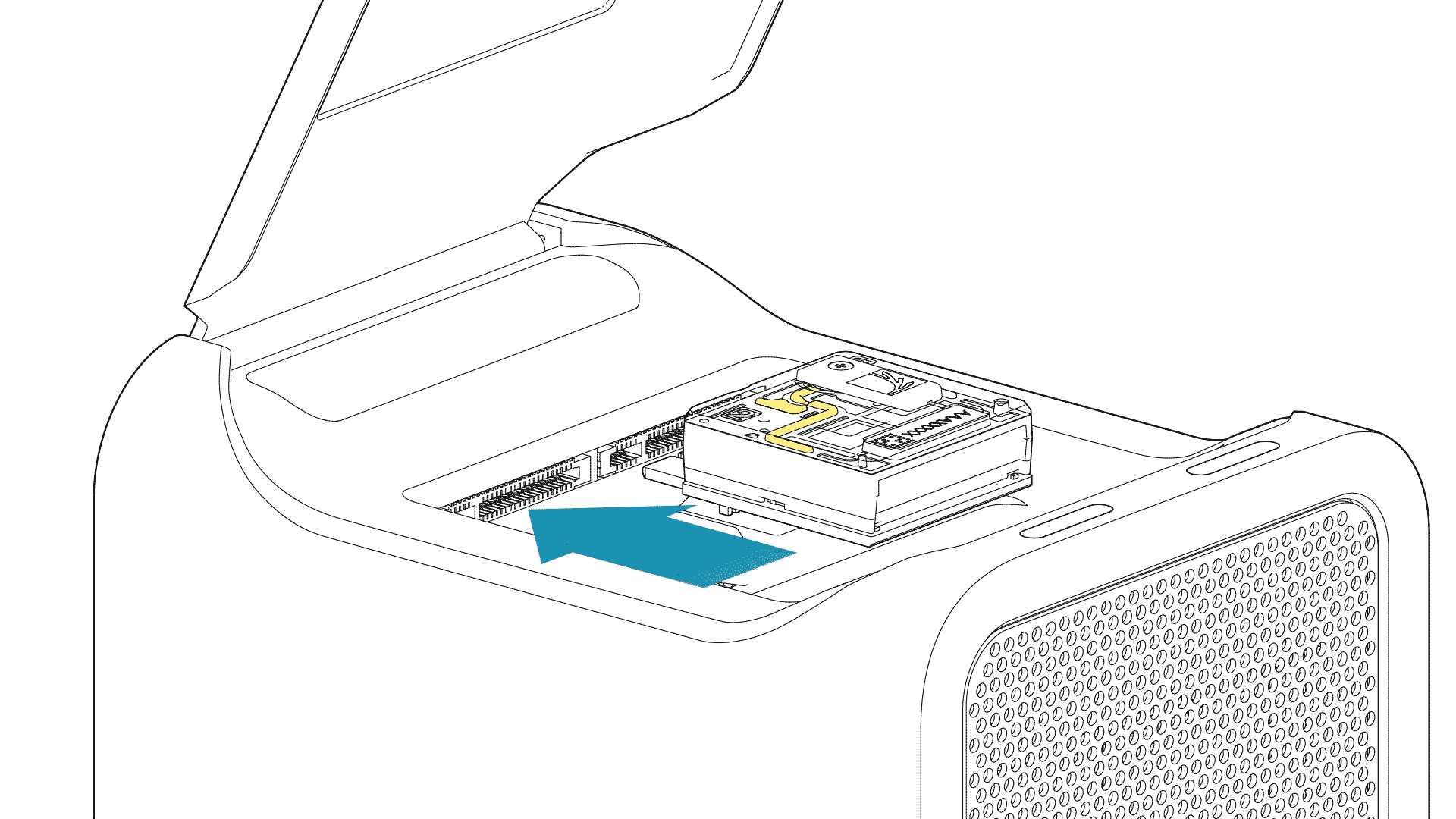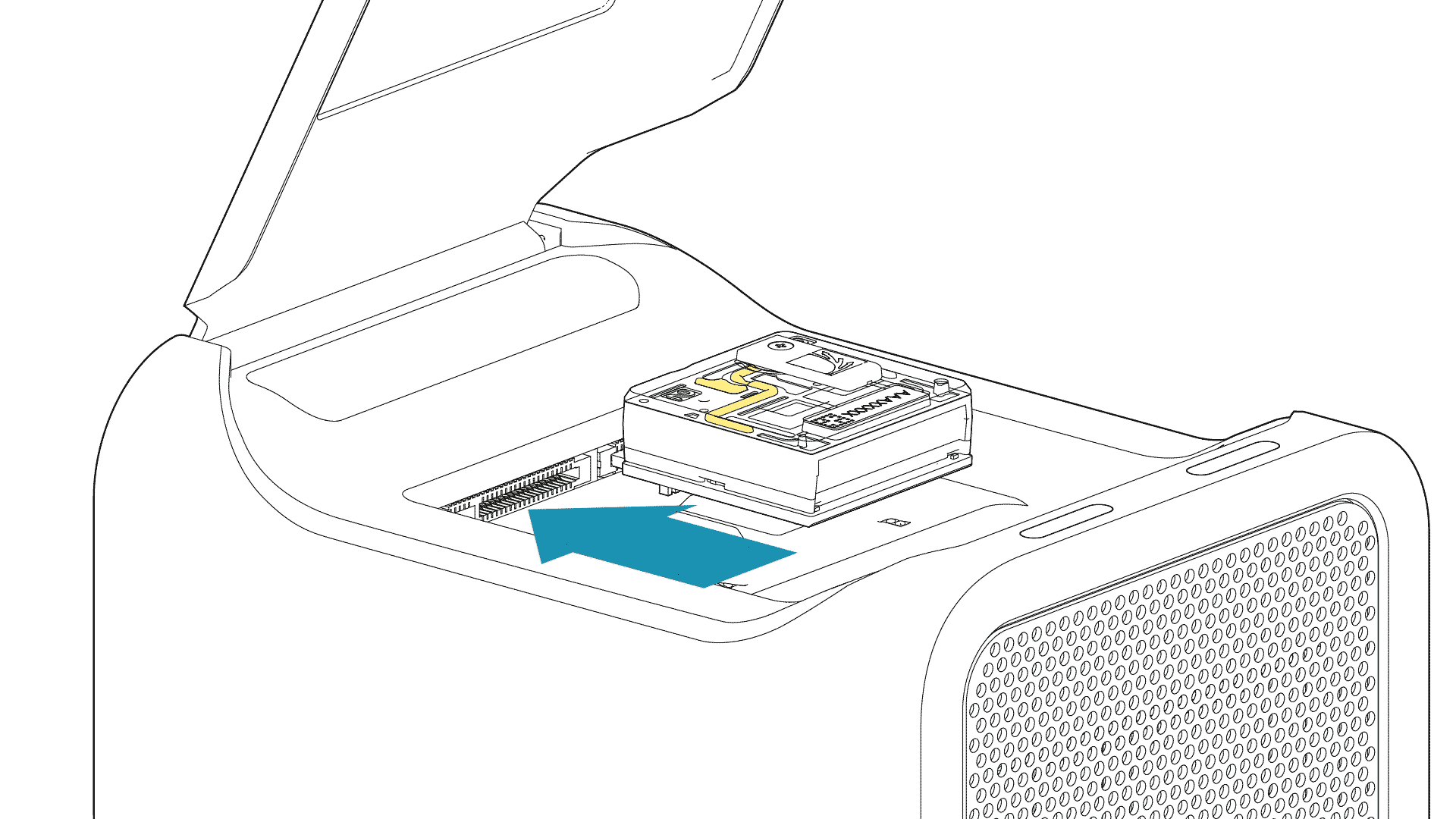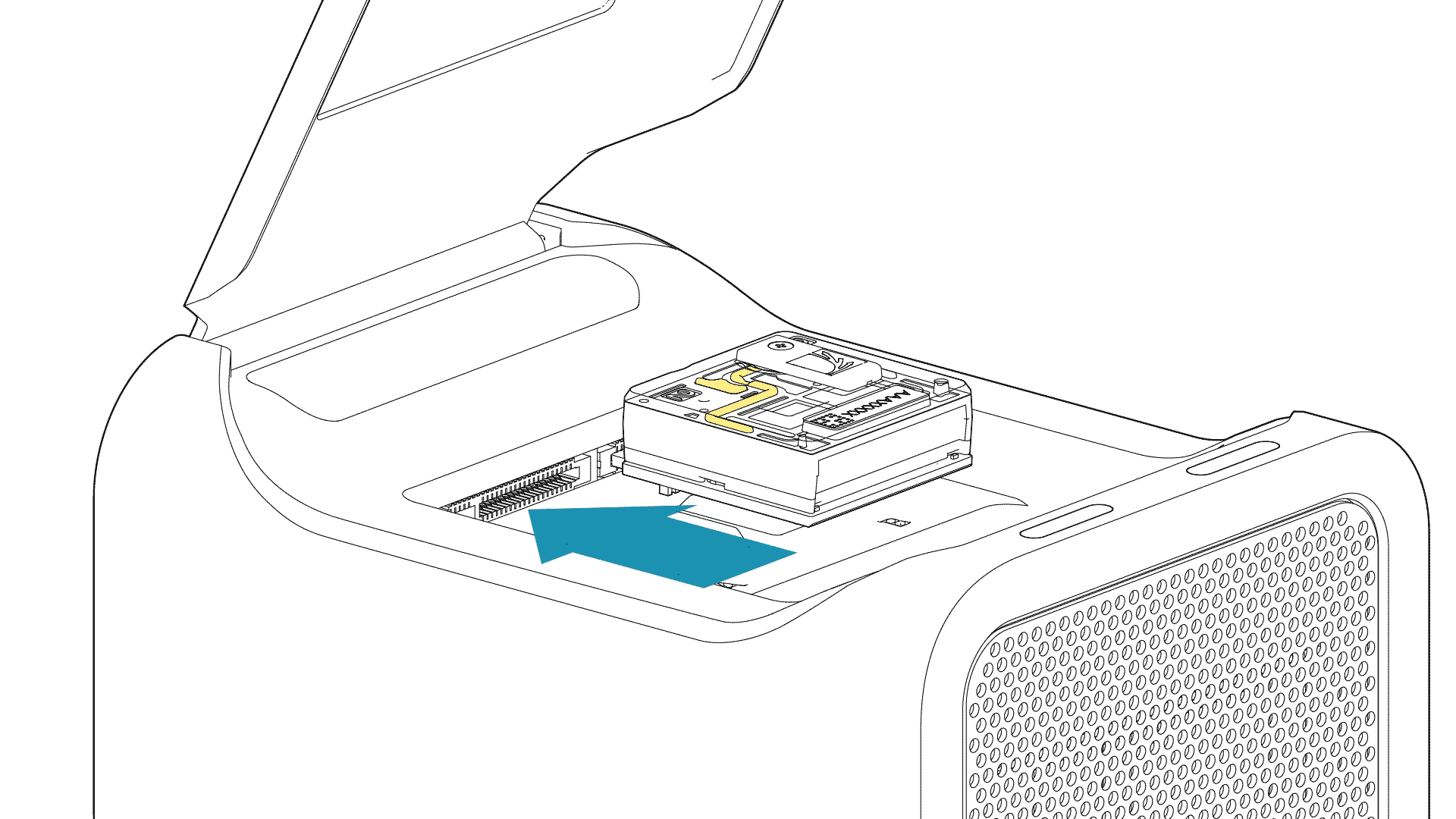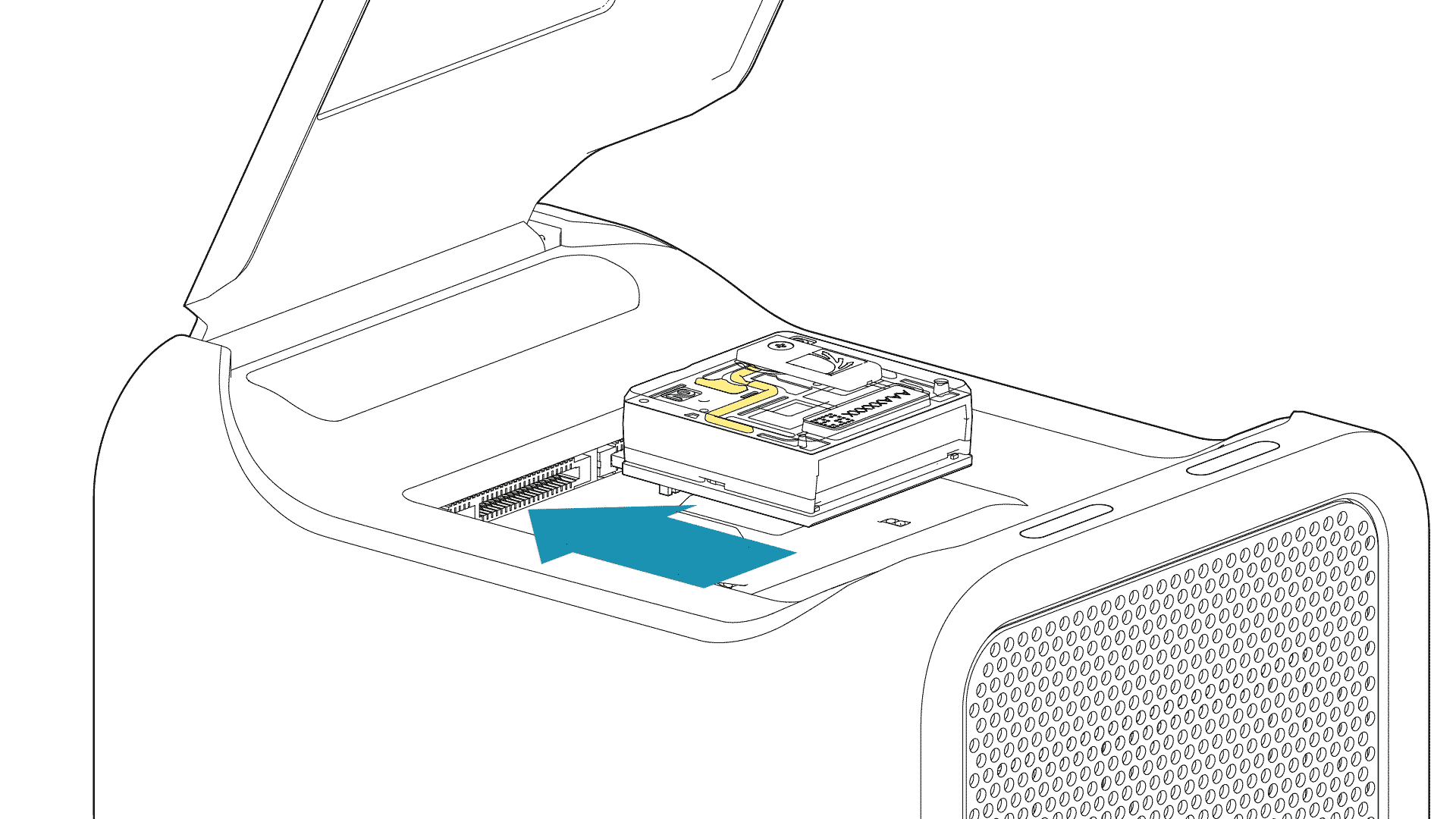
For the PromethION 24/48, load the flow cell(s) into the docking ports:
- Line up the flow cell with the connector horizontally and vertically before smoothly inserting into position.
- Press down firmly onto the flow cell and ensure the latch engages and clicks into place.
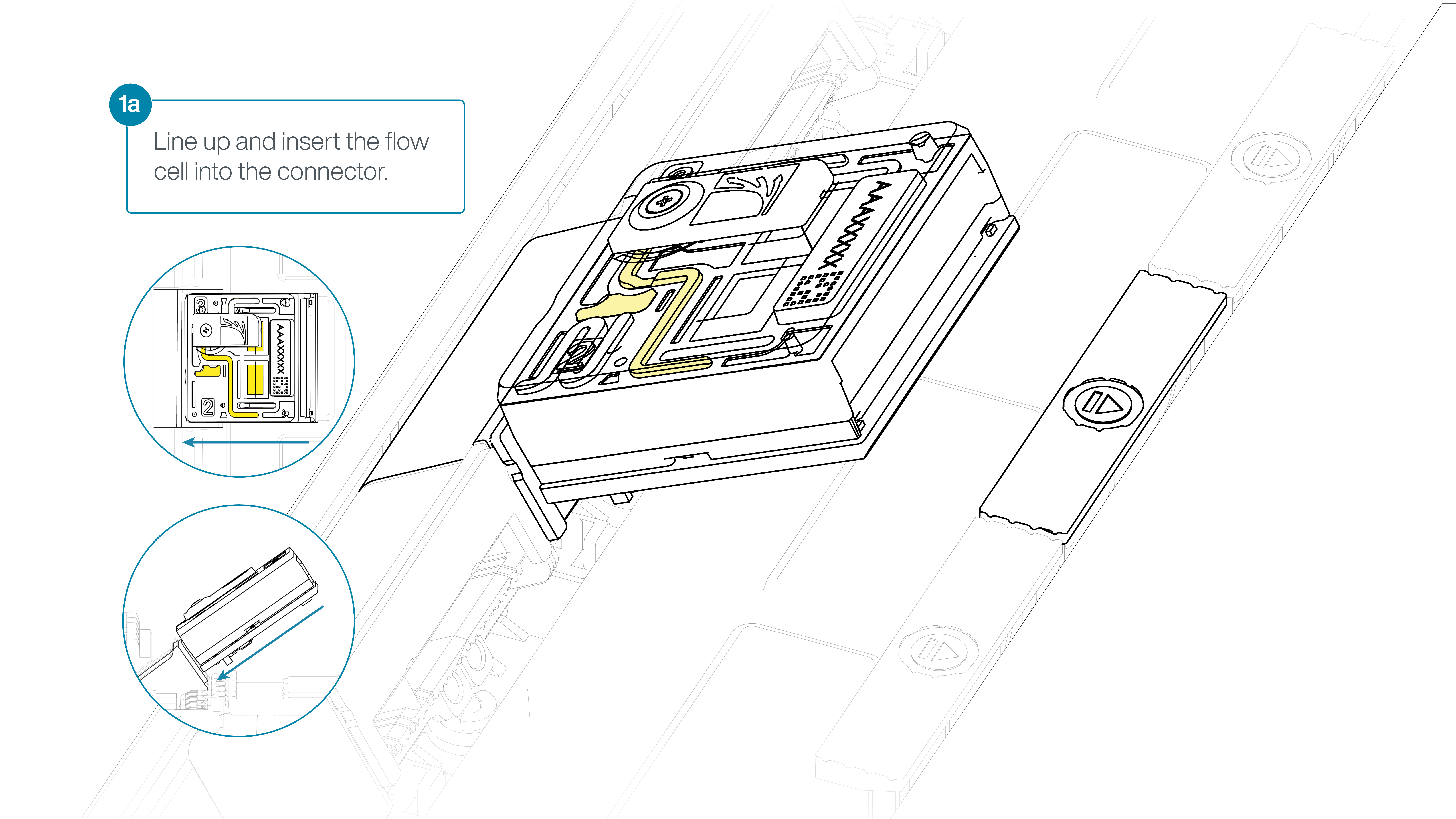
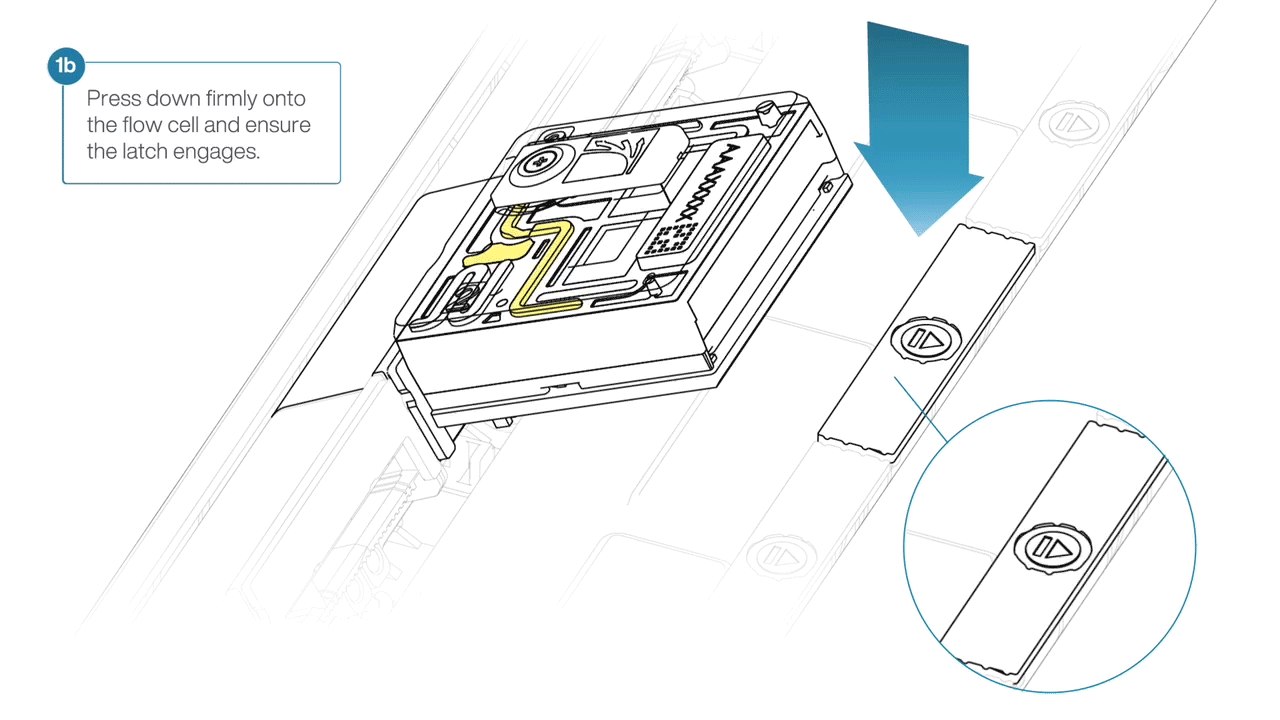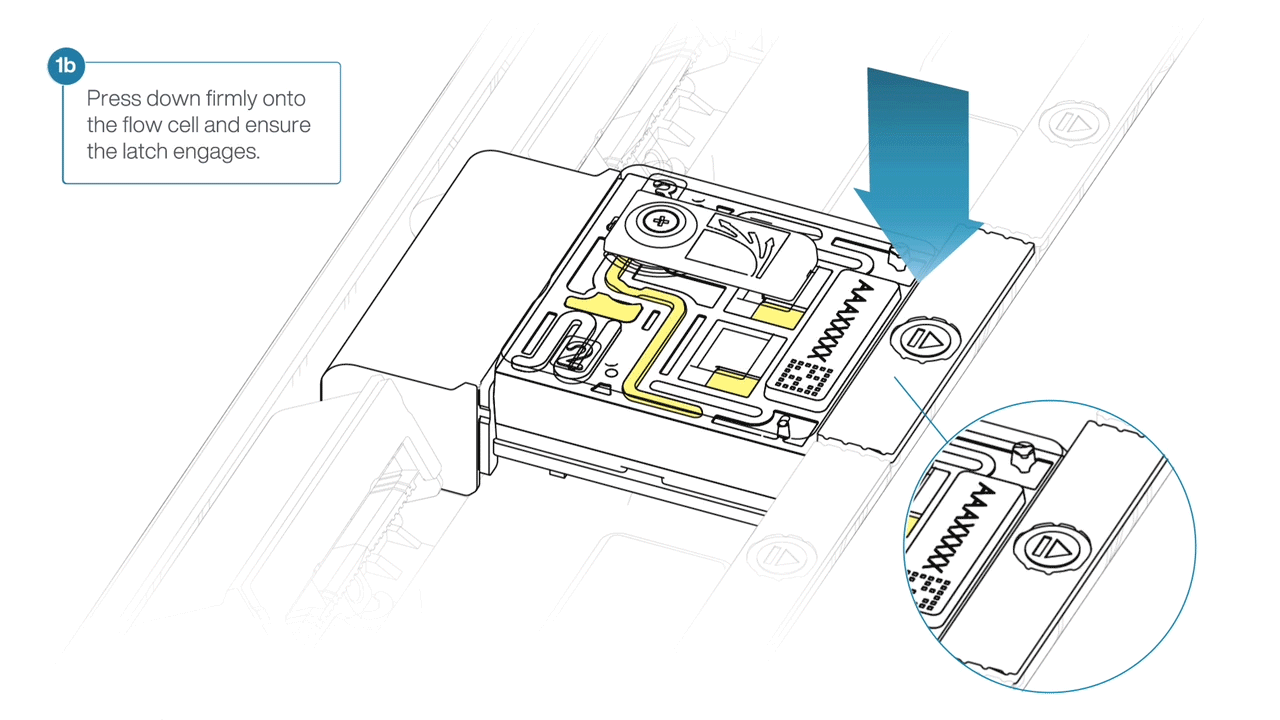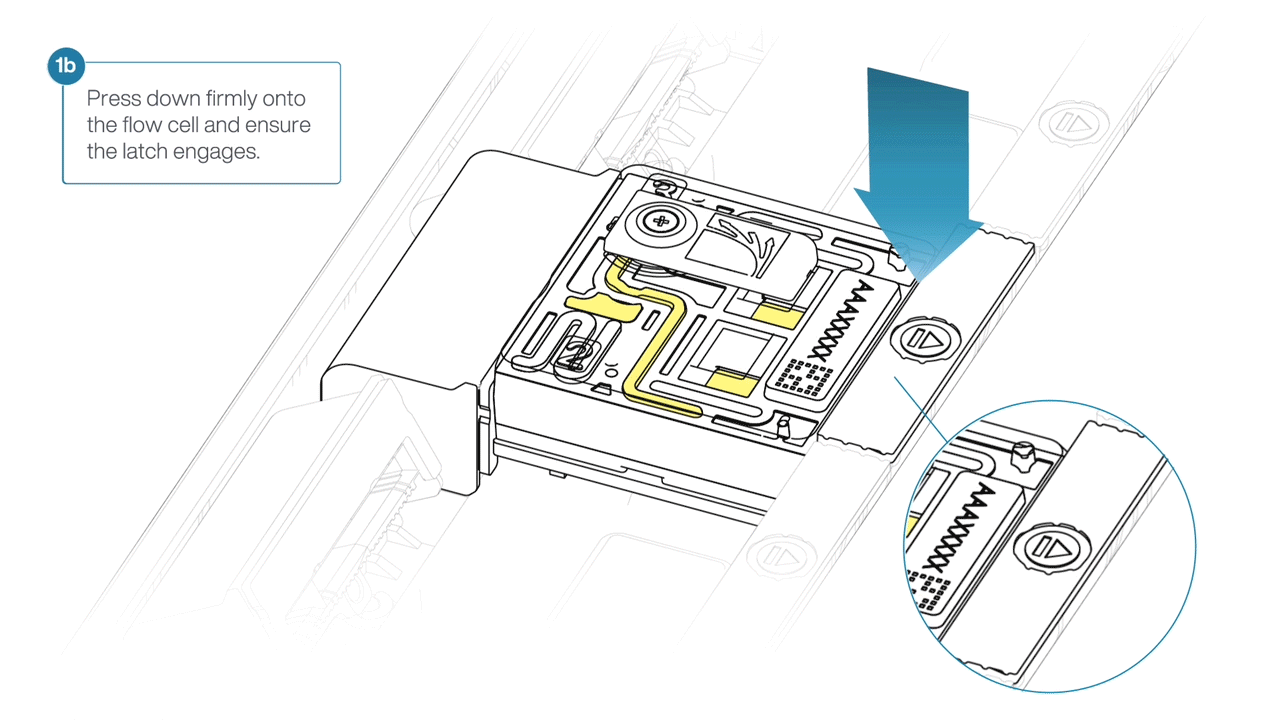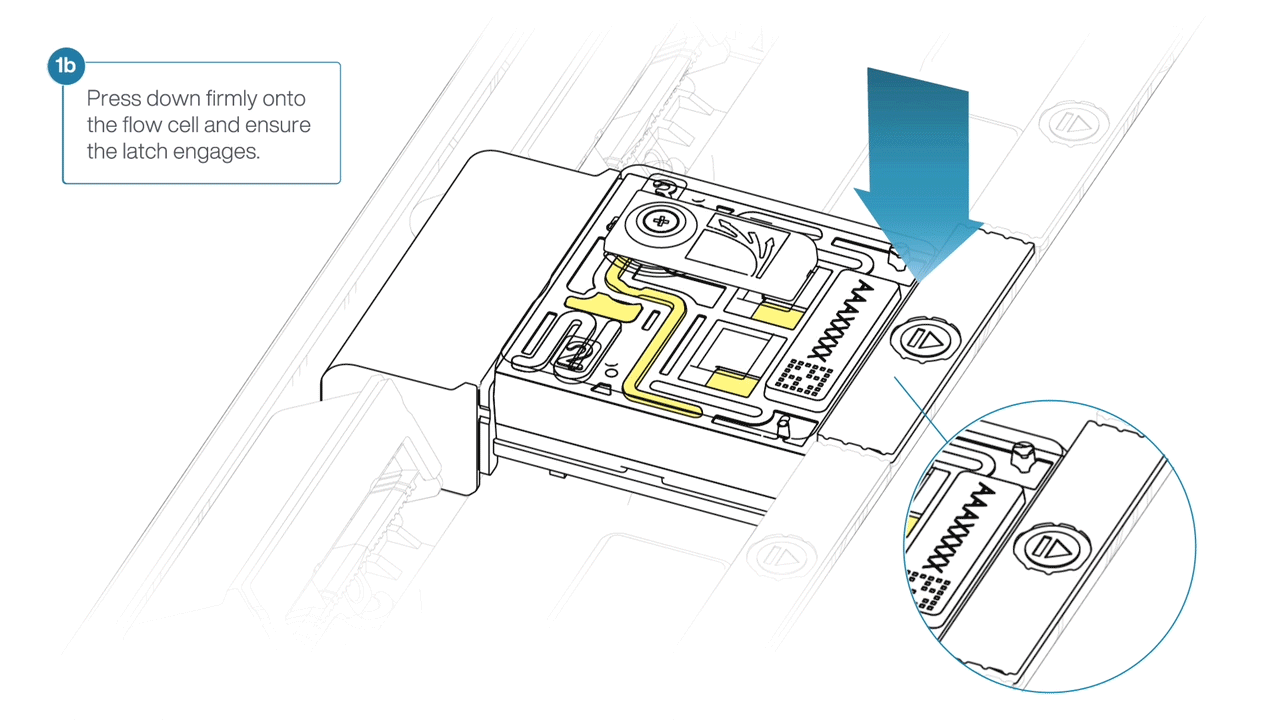
IMPORTANTE
Insertion of the flow cells at the wrong angle can cause damage to the pins on the PromethION and affect your sequencing results. If you find the pins on a PromethION position are damaged, please contact support@nanoporetech.com for assistance.

If not already completed, perform a flow cell check on all flow cells.
Please refer to the Flow Cell Check protocol for further information.
Slide the inlet port cover clockwise to open.
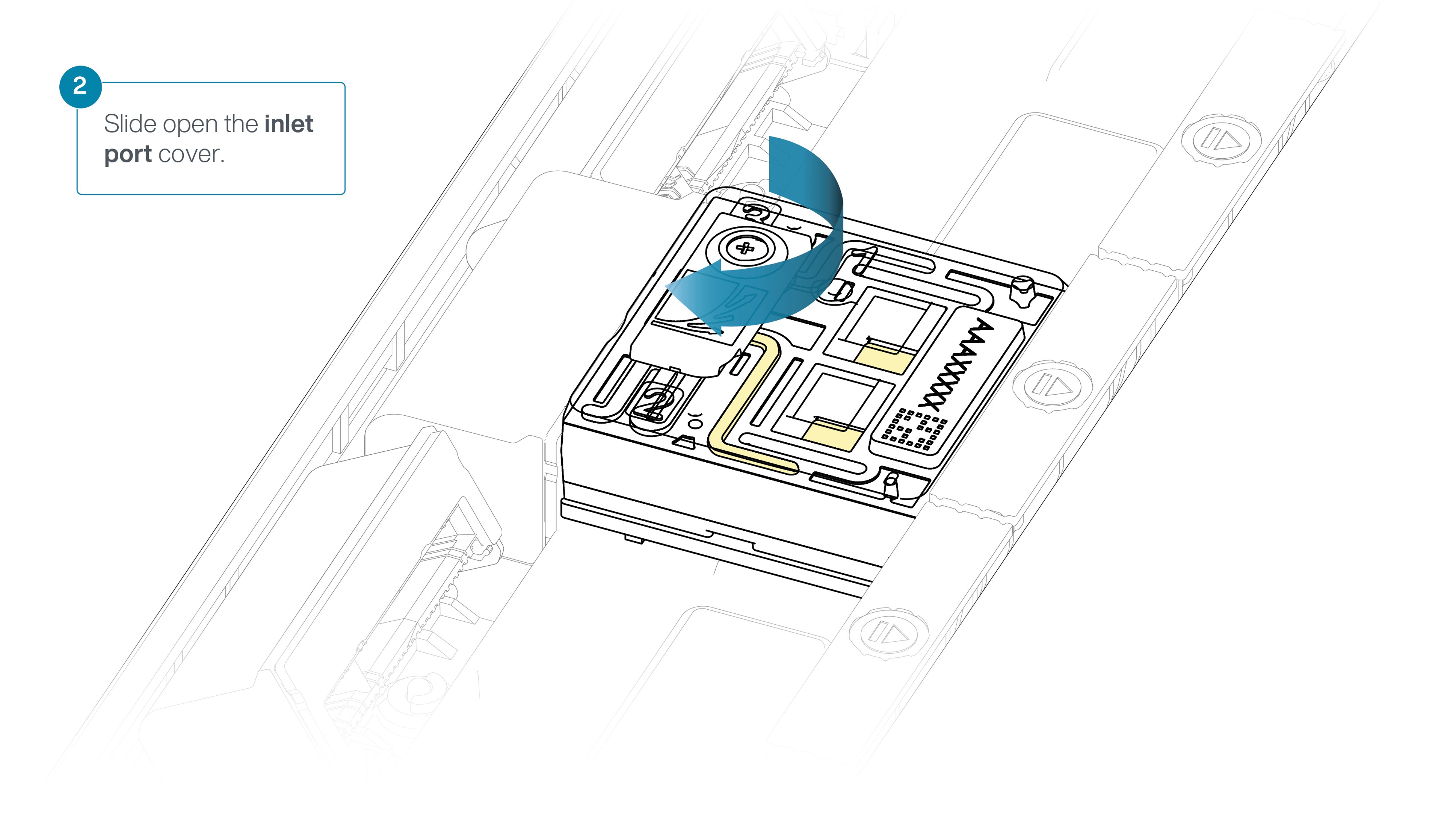
IMPORTANTE
Tenga cuidado a la hora de extraer el tampón. No retire más de 20-30 μl y asegúrese de que el tampón cubra la matriz de poros en todo momento. La introducción de burbujas de aire en la matriz puede dañar los poros de manera irreversible.
After opening the inlet port, draw back a small volume to remove any air bubbles:
- Set a P1000 pipette tip to 200 µl.
- Insert the tip into the inlet port.
- Turn the wheel until the dial shows 220-230 µl, or until you see a small volume of buffer entering the pipette tip.
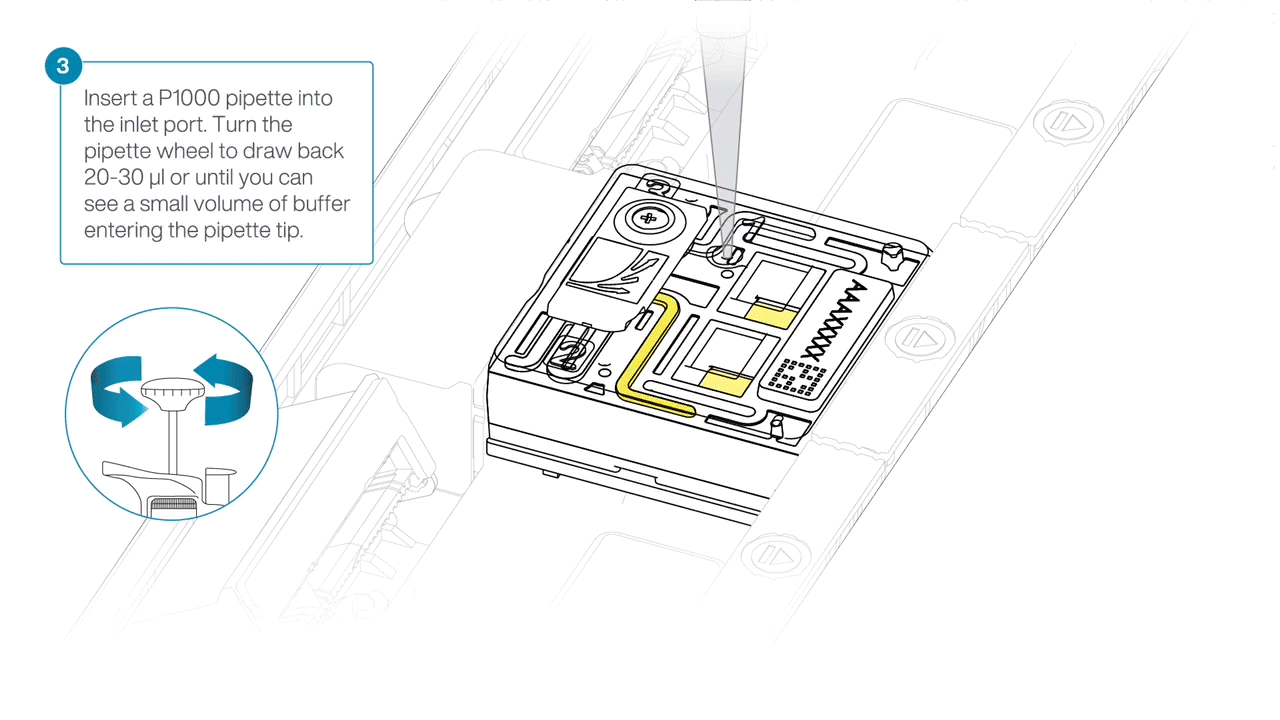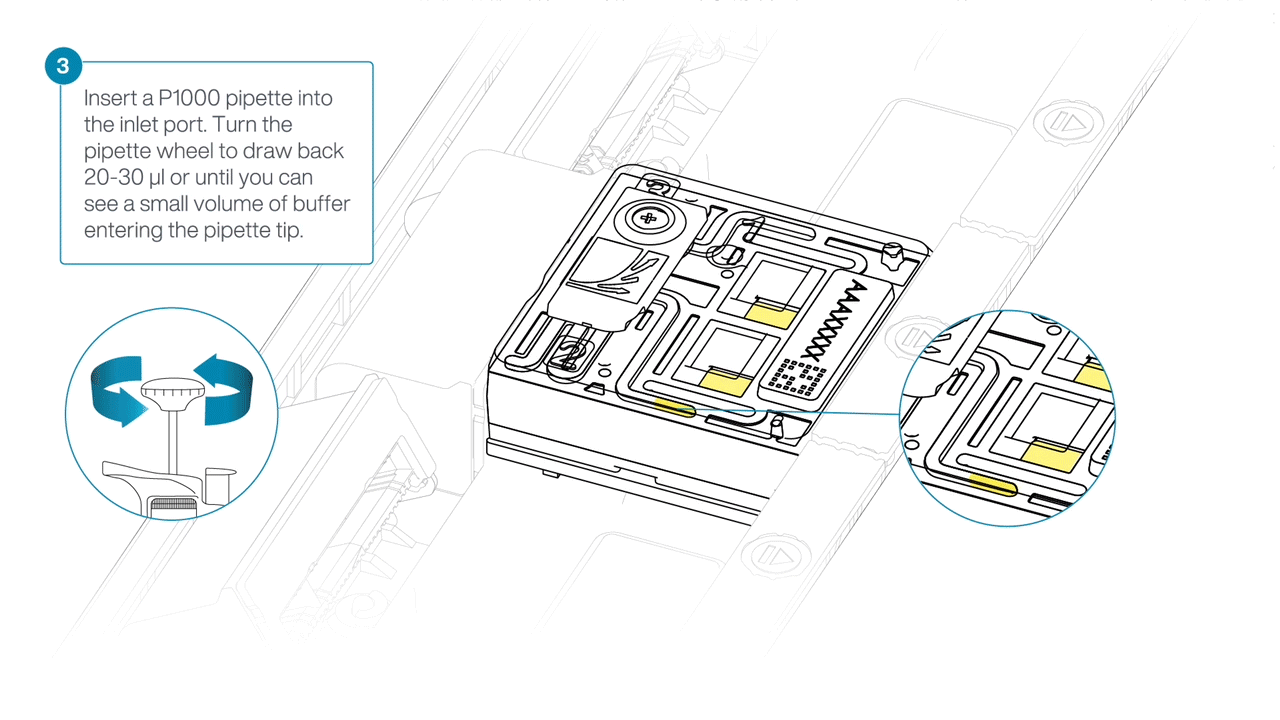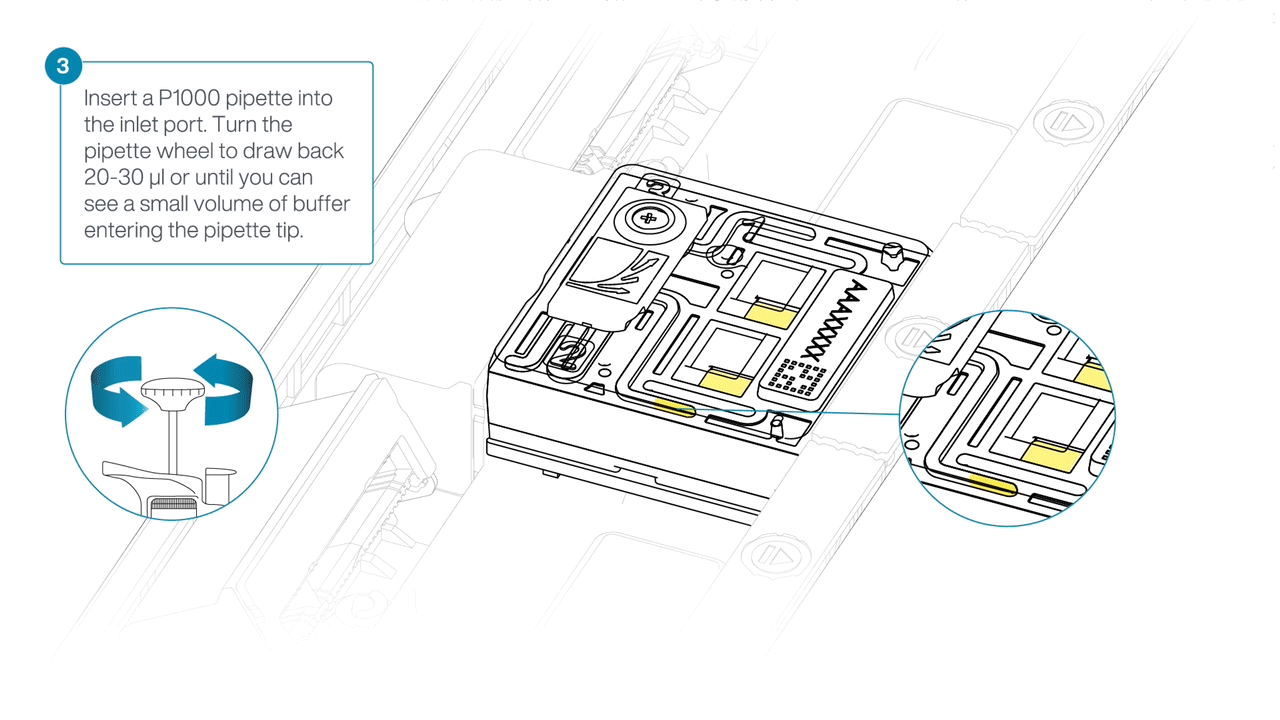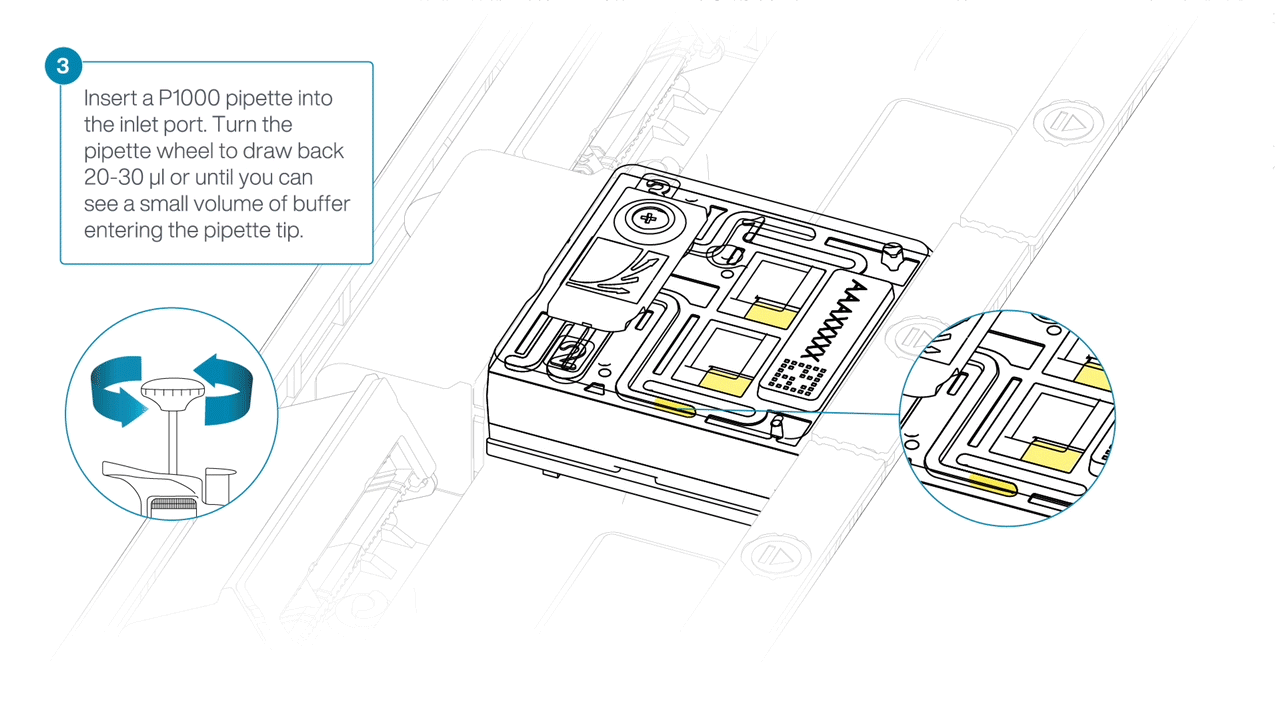
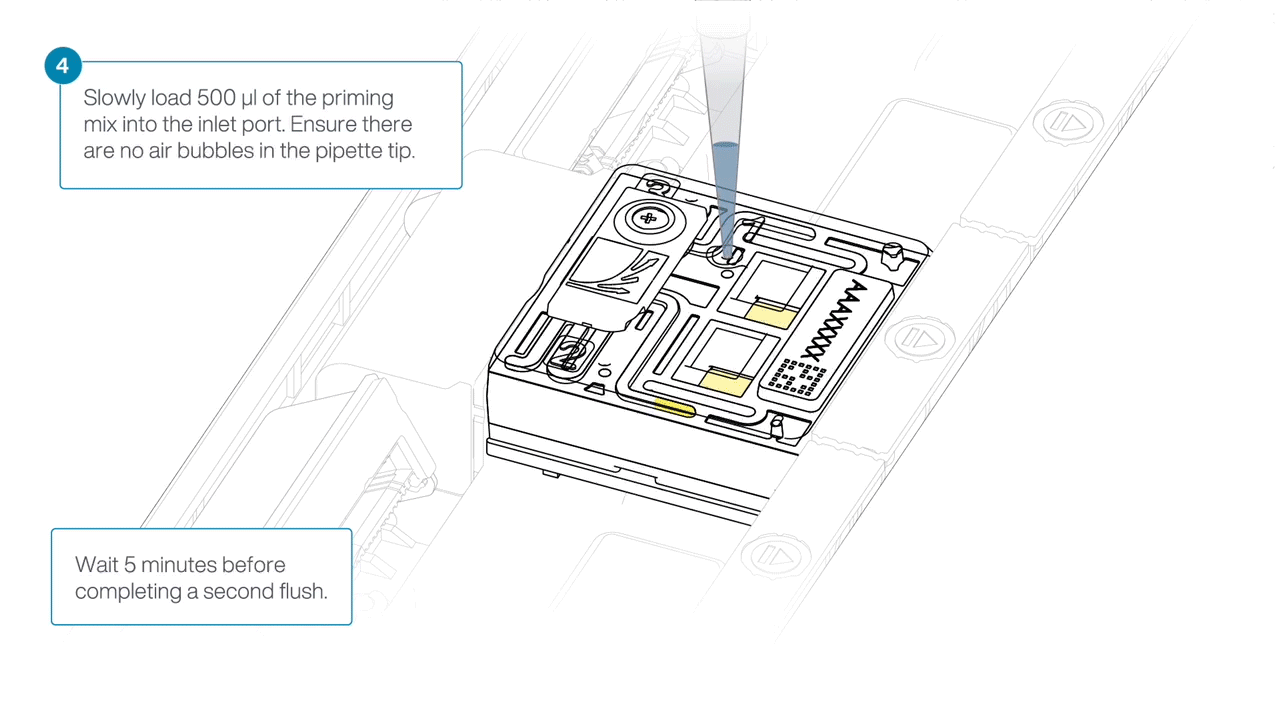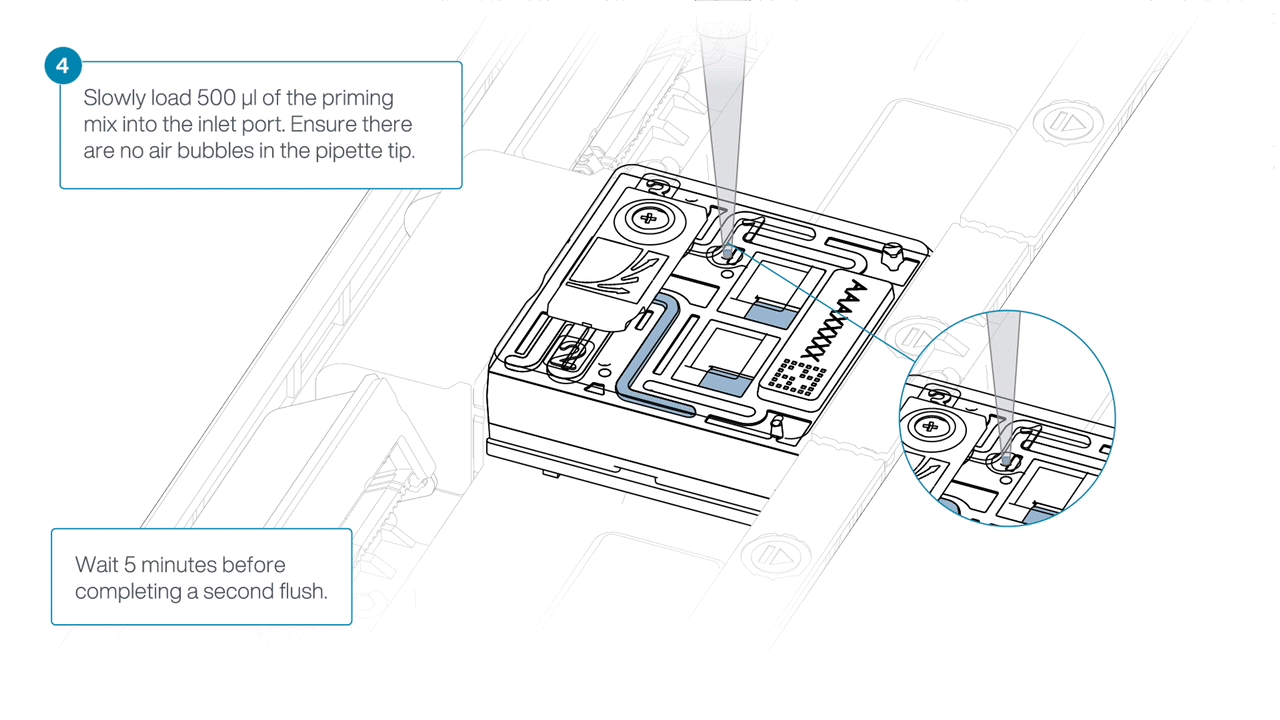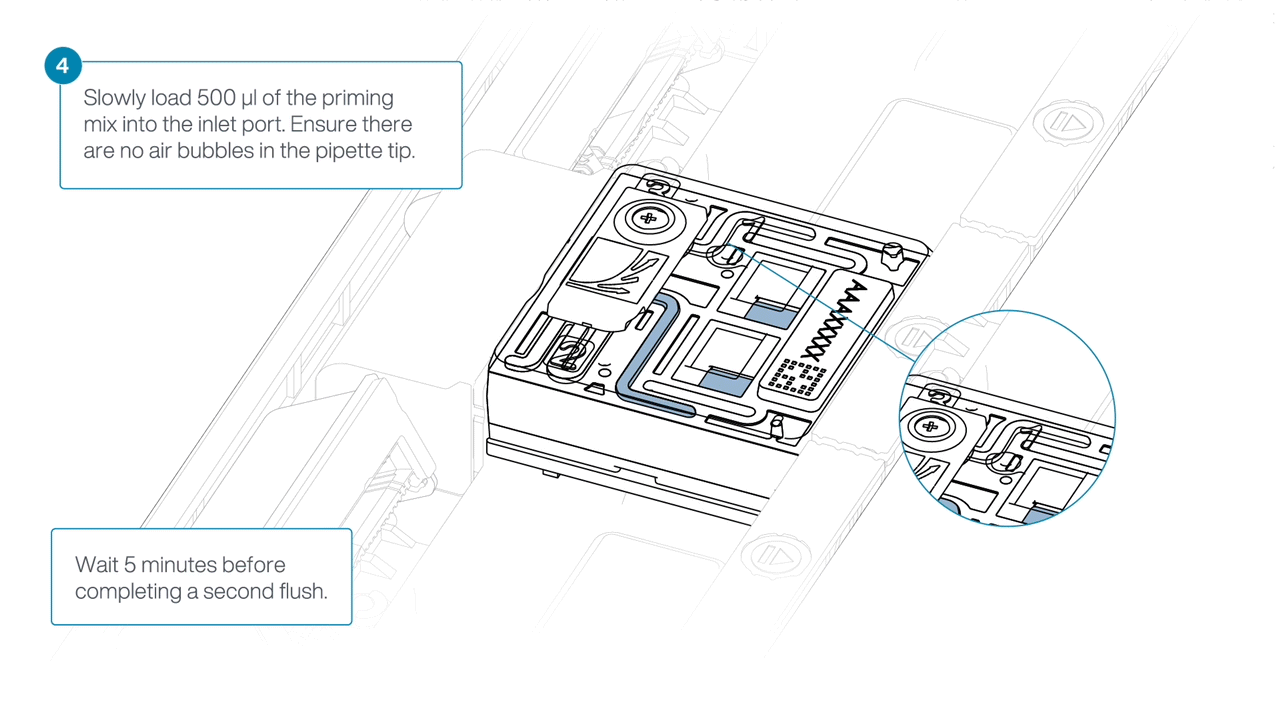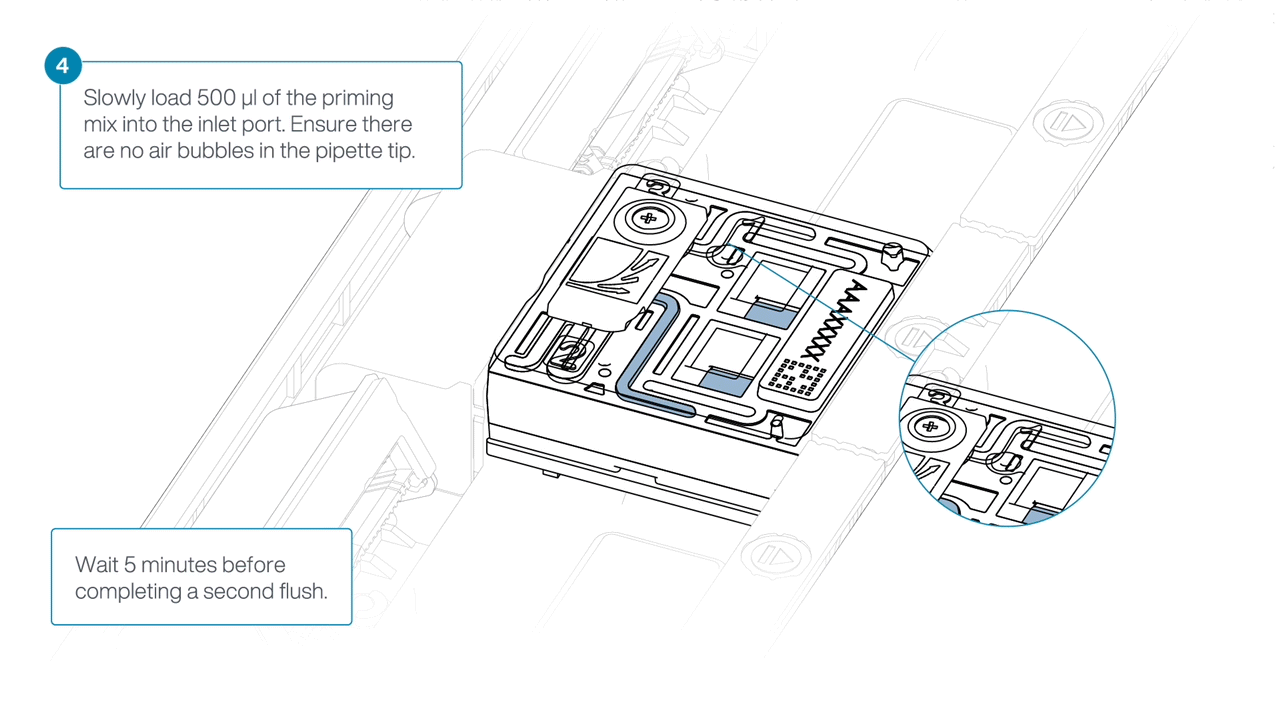
Load 500 µl of the priming mix into the flow cell via the inlet port, avoiding the introduction of air bubbles. Wait five minutes. During this time, prepare the library for loading using the next steps in the protocol.
Thoroughly mix the contents of the Sequencing buffer II (SBII) and Loading Beads II (LBII) tubes by vortexing.
IMPORTANTE
The Loading Beads II (LBII) tube contains a suspension of beads. These beads settle very quickly. It is vital that they are mixed immediately before use.
In a separate tube for each library, prepare for loading by adding the following reagents:
| Reagent | Volume |
|---|---|
| SBII | 75 µl |
| LB | 51 µl |
| DNA library | 24 µl |
| Total | 150 µl |
MEDIDA OPCIONAL
The Ligation Sequencing Kit XL kit is designed for users running multiple samples/flow cells. When handling multiple DNA preparations, the Sequencing Buffer (SBII) and Loading Beads (LBII) can be combined in a master mix:
Mix the Sequencing Buffer (SBII) and Loading Beads (LBII) as described above, scaling up the final volume for the appropriate number of samples and adding up to 20% excess of each reagent. Mix the master mix by pipetting immediately before adding to the DNA samples. Pipette 126 µl of the master mix into each DNA sample-containing tube. Mix the sample by pipetting.
Complete the flow cell priming by slowly loading 500 µl of the priming mix into the inlet port.
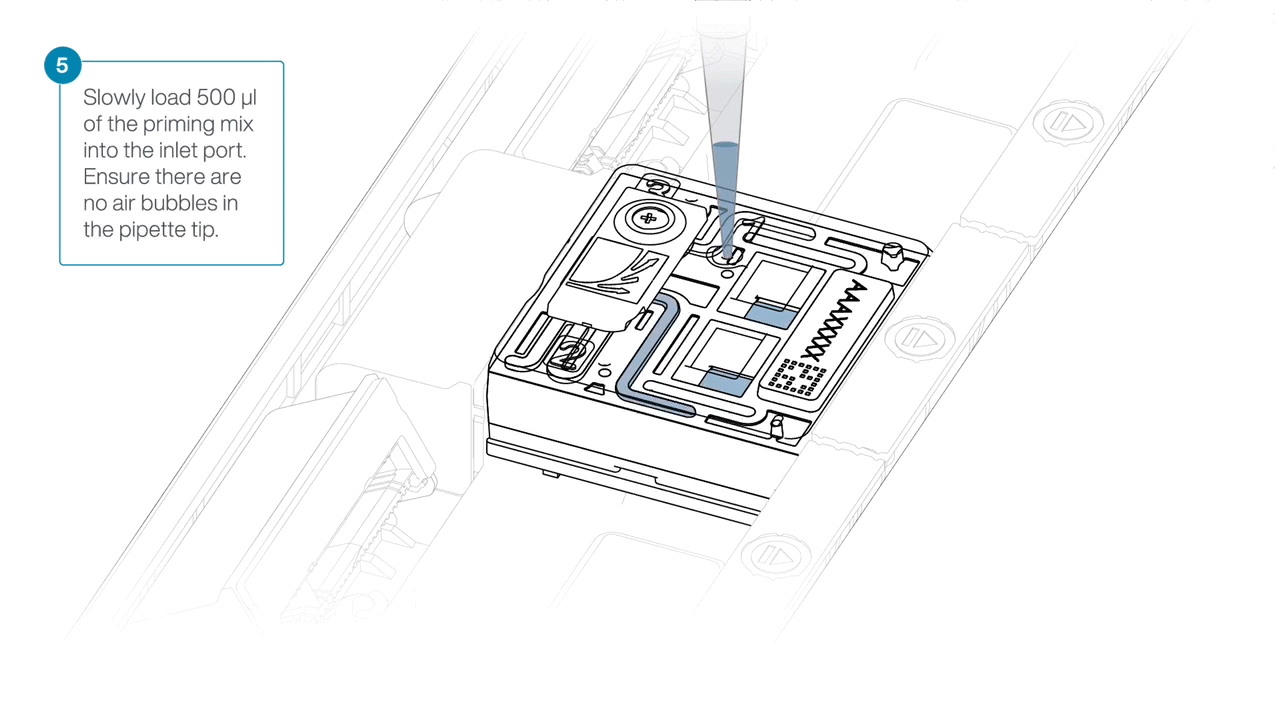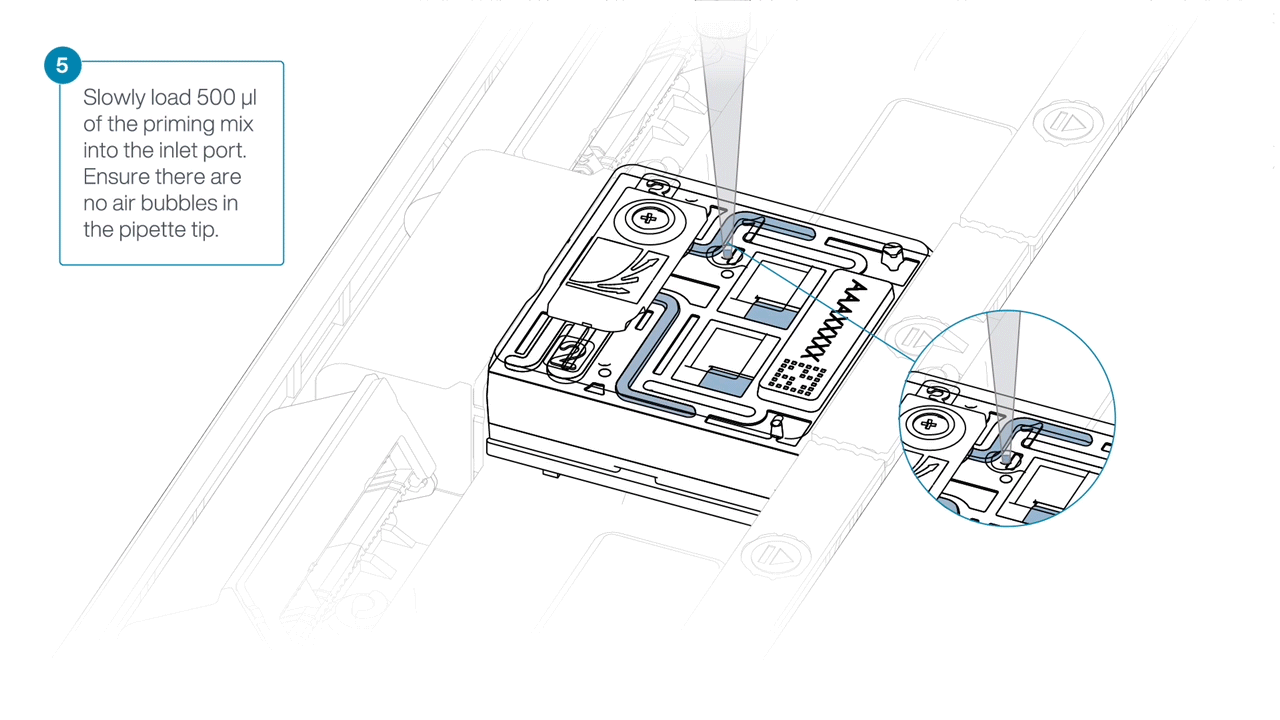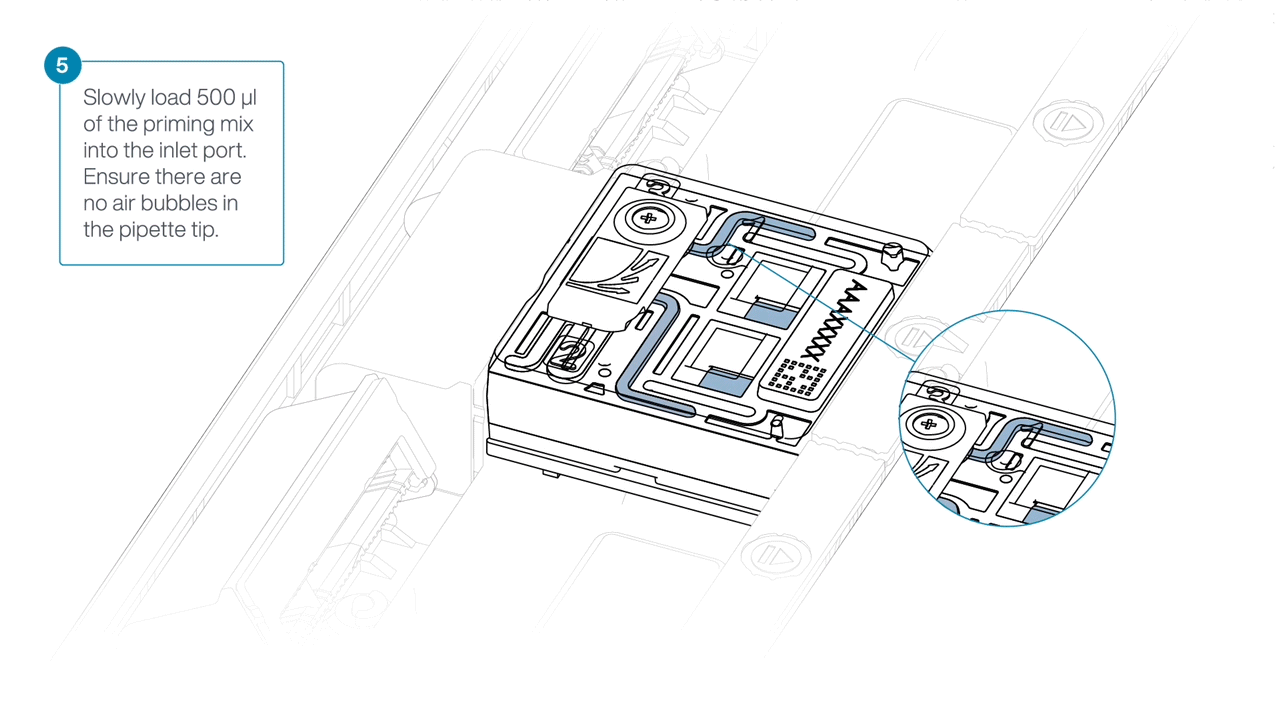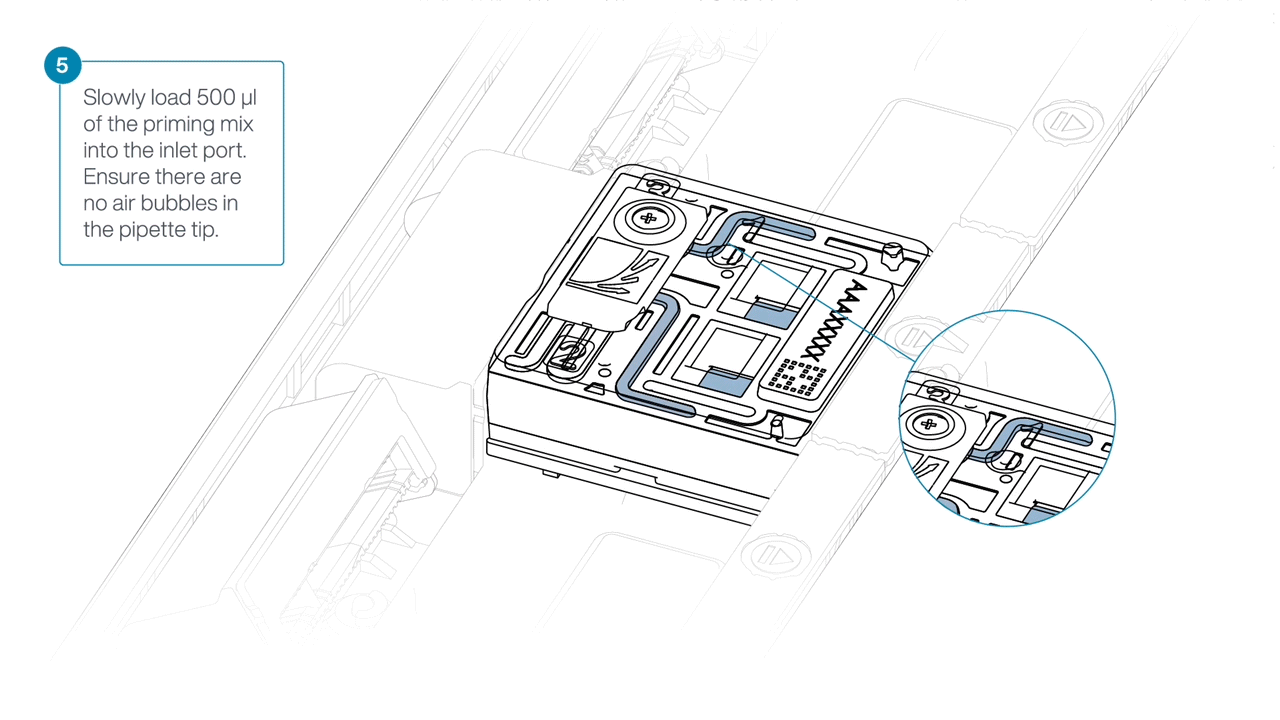
Mezclar la biblioteca pipeteando suavemente, justo antes de cargar.
Using a P1000, insert the pipette tip into the inlet port and add 150 µl of library.
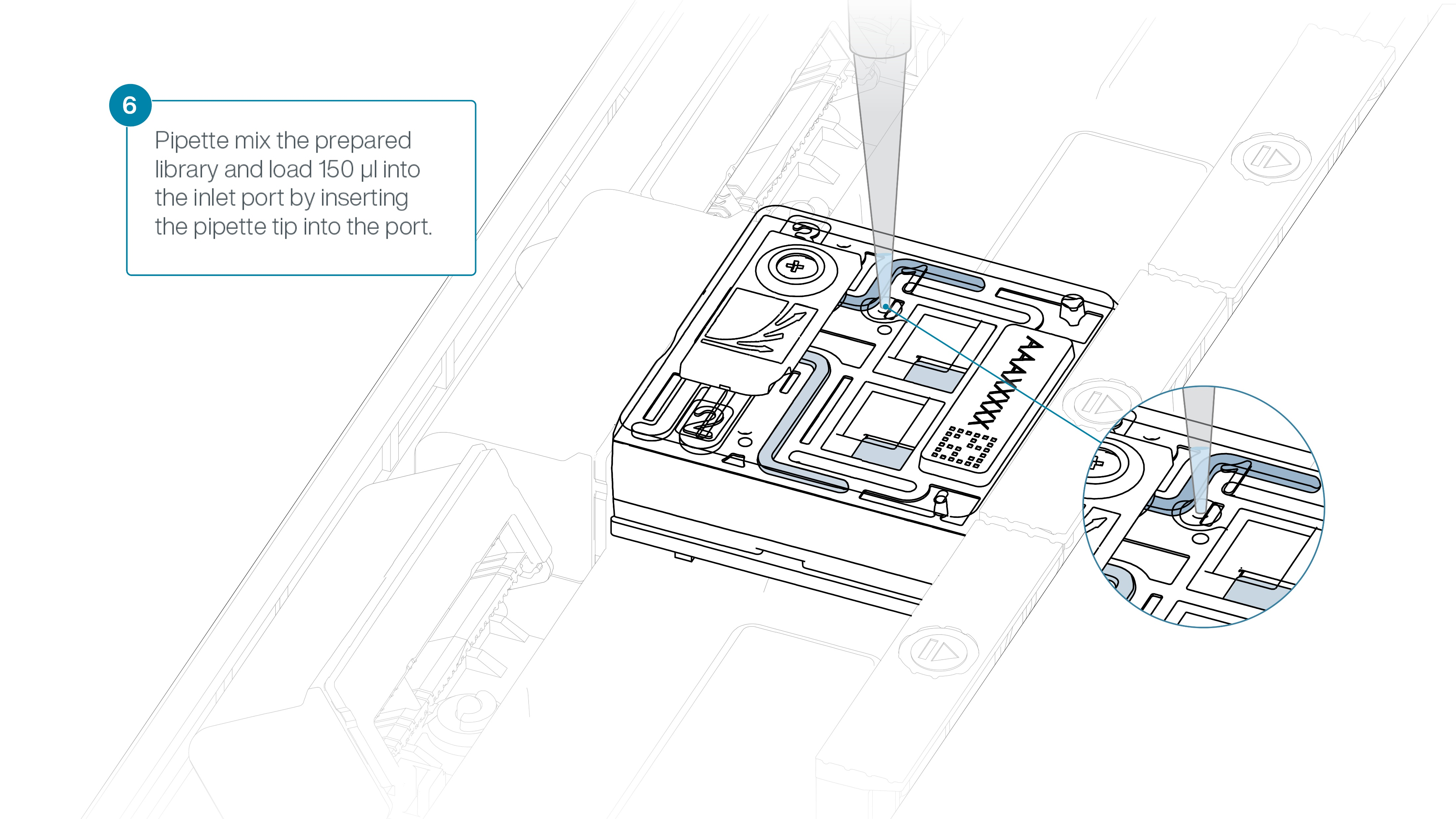
Close the valve to seal the inlet port and close the PromethION lid when ready.
Wait a minimum of 10 minutes after loading the flow cells onto the PromethION before initiating any experiments. This will help to increase the sequencing output.
For multiple flow cell washing, use the same experiment name and identifying sample IDs for all runs to enable all flow cells to be paused simultaneously.
7. Data acquisition and basecalling
How to start sequencing
Once you have loaded your flow cell, the sequencing run can be started on MinKNOW, our sequencing software that controls the device, data acquisition and real-time basecalling. For more detailed information on setting up and using MinKNOW, please see the MinKNOW protocol.
MinKNOW can be used and set up to sequence in multiple ways:
- On a computer either direcly or remotely connected to a sequencing device.
- Directly on a GridION, MinION Mk1C or PromethION 24/48 sequencing device.
For more information on using MinKNOW on a sequencing device, please see the device user manuals:
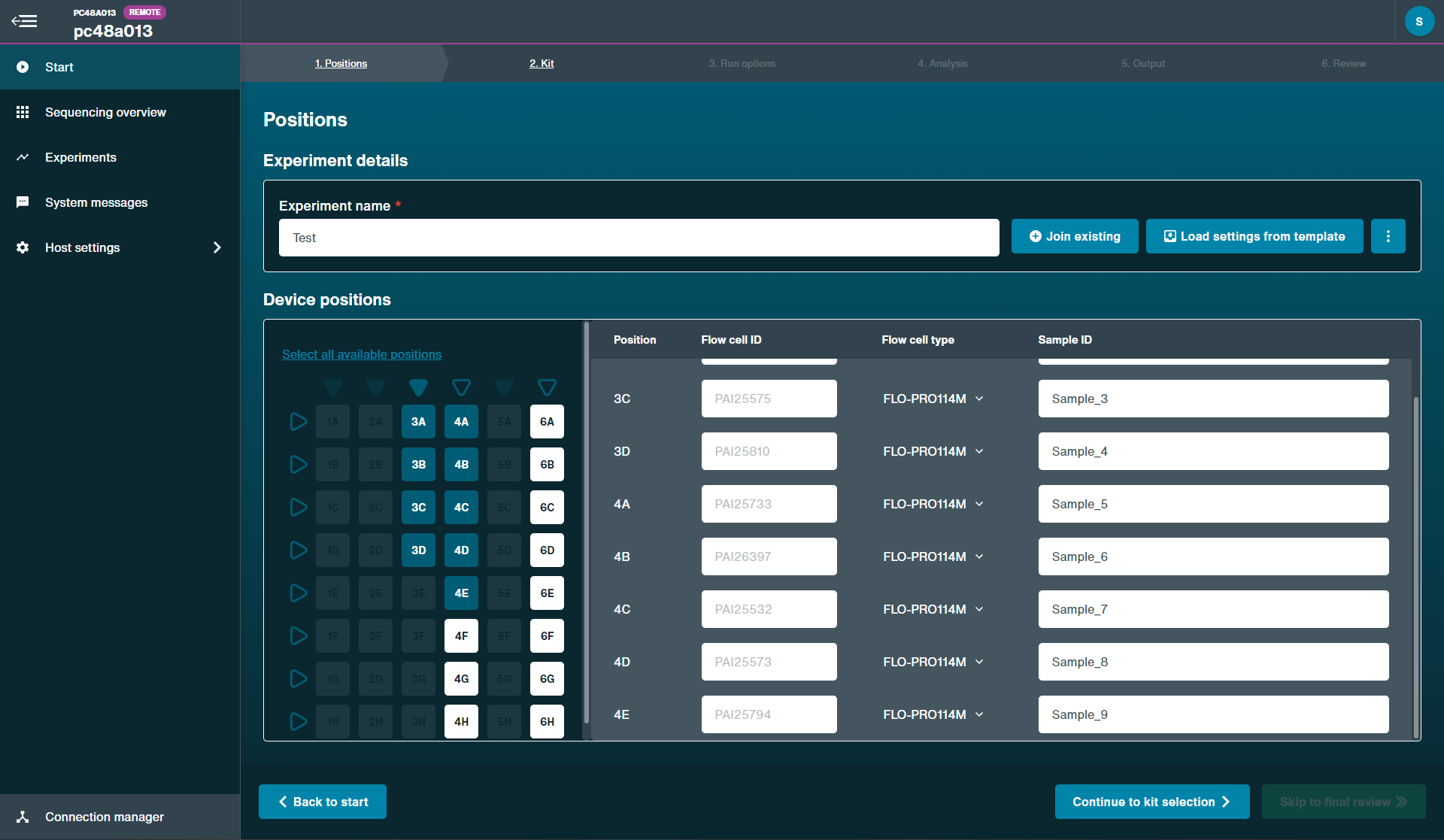
To start a sequencing run on MinKNOW:
1. Navigate to the start page and click Start sequencing.
2. Fill in your experiment details, such as name and flow cell position and sample ID.
3. Select the sequencing kit used in the library preparation on the Kit page.
4. Configure the sequencing and output parameters for your sequencing run or keep to the default settings on the Run configuration tab.
Note: If basecalling was turned off when a sequencing run was set up, basecalling can be performed post-run on MinKNOW. For more information, please see the MinKNOW protocol.
5. Click Start on the Review page to start the sequencing run.
Análisis de datos tras la secuenciación
Una vez la secuenciación ha finalizado, la celda de flujo se puede reutilizar o devolver, como se indica en la sección Reutilización y devoluciones de celdas de flujo.
Tras la secuenciación y la identificación de bases, se puede proceder a analizar los datos. Si desea más información sobre las opciones de identificación de bases y de análisis posterior, consulte el documento Data Analysis.
En la sección Downstream analysis/Análisis posterior, le presentamos otras opciones para analizar los datos.
8. Reutilización y devoluciones de las celdas de flujo
Material
- Flow Cell Wash Kit (EXP-WSH004) (kit de lavado de celda de flujo)
Si al terminar el experimento desea volver a usar la celda de flujo, siga las instrucciones del protocolo Flow Cell Wash Kit y guarde la celda de flujo lavada a 2-8 ⁰C.
El protocolo Flow Cell Wash Kit está disponible en la comunidad Nanopore.
CONSEJO
Una vez terminado el experimento, recomendamos lavar la celda de flujo cuanto antes. Si no es posible, se puede dejar en el dispositivo y lavar al día siguiente.
Otra posibilidad es seguir el procedimiento de devolución para lavar la celda de flujo y enviarla a Oxford Nanopore.
Aquí puede encontrar las instrucciones para devolver celdas de flujo.
Nota: Antes de proceder a su devolución, las celdas de flujo deben lavarse con agua desionizada.
IMPORTANTE
Ante cualquier duda o pregunta acerca del experimento de secuenciación, consulte la guía de resolución de problemas, Troubleshooting Guide, que se encuentra en la versión en línea de este protocolo.
9. Análisis posterior
Análisis posterior a la identificación de bases
Existen varias opciones para completar el análisis de los datos de bases identificadas:
1. Plataforma EPI2ME
La plataforma EPI2ME es un servicio de análisis de datos, alojado en la red, desarrollado por Metrichor Ltd., filial de Oxford Nanopore Technologies. EPI2ME ofrece una serie de procesos de análisis, p. ej., de identificación metagenómica, identificación de especies a partir de un código (barcoding), alineación e identificación de variantes estructurales. El análisis no requiere equipo ni capacidad computacional extra y proporciona un informe fácil de interpretar con los resultados. Para obtener información sobre cómo realizar un proceso de análisis en EPI2ME, siga las instrucciones del protocolo, empezando por la sección "Starting data analysis".
2. EPI2ME Labs, tutoriales y procesos de trabajo
Para realizar un análisis de datos más exhaustivo, Oxford Nanopore Technologies ofrece una serie de tutoriales y procesos de trabajo de bioinformática, disponibles en EPI2ME Labs, que encontrará en la sección del mismo nombre de la comunidad Nanopore. La plataforma proporciona un espacio donde los proyectos que depositan en GitHub nuestros equipos de Investigación y Aplicaciones, se pueden exponer con textos descriptivos, código bioinformático funcional y datos de ejemplo.
3. Herramientas de análisis para la investigación
El departamento de Investigación de Oxford Nanopore Technologies ha creado una serie de herramientas de análisis que están disponibles en el repositorio Oxford Nanopore de GitHub. Las herramientas están diseñadas para usuarios avanzados y contienen instrucciones sobre cómo instalar y ejecutar el programa. Estas herramientas están públicamente disponibles y cuentan con un mantenimiento mínimo.
4. Herramientas de análisis desarrolladas por la comunidad
Si no se proporciona en ninguno de los recursos anteriores un método de análisis que responda a las necesidades de investigación requeridas, puede consultar la sección Bioinformatics del centro de recursos Resource Centre. Varios miembros de la comunidad Nanopore han desarrollado sus propias herramientas y segmentaciones para analizar datos de secuenciación por nanoporos. La mayoría de ellos está disponible en GitHub. Nótese que Oxford Nanopore Technologies no desarrolla ni mantiene esas herramientas y no garantiza que sean compatibles con la última configuración de química/software.
10. Issues during automation of library preparation
Please contact your automation vendor FAS and/or Nanopore FAS if you have any issues.
11. Issues during the sequencing run
A continuación hay una lista de los problemas más frecuentes, con algunas posibles causas y soluciones propuestas.
También disponemos de una página de preguntas frecuentes, FAQ, en la sección Support de la comunidad Nanopore.
Si ha probado las soluciones propuestas y continúa teniendo problemas, póngase en contacto con el departamento de asistencia técnica, bien por correo electrónico (support@nanoporetech.com) o a través del Live Chat de la comunidad Nanopore.
Menos poros al inicio de la secuenciación que después de verificar la celda de flujo
| Observación | Posible causa | Comentarios y acciones recomendadas |
|---|---|---|
| MinKNOW presentó al inicio de la secuenciación un número de poros inferior al indicado durante la comprobación de la celda de flujo | Se introdujo una burbuja de aire en la matriz de nanoporos | Tras comprobar el número de poros presente en la celda de flujo, es imprescindible quitar las burbujas que haya cerca del puerto de cebado. Si no se quitan, pueden desplazarse a la matriz de nanoporos y dañar de manera irreversible los nanoporos expuestos al aire. En este vídeo se muestran algunas buenas prácticas para evitar que esto ocurra. |
| MinKNOW presentó al inicio de la secuenciación un número de poros inferior al indicado durante la comprobación de la celda de flujo | La celda de flujo no está colocada correctamente | Detener el ciclo de secuenciación, quitar la celda de flujo del dispositivo e insertarla de nuevo. Comprobar que está firmemente asentada en el dispositivo y que ha alcanzado la temperatura deseada. Si procede, probar con una posición diferente del dispositivo (GriION/PromethION). |
| MinKNOW presentó al inicio de la secuenciación un número de poros inferior al indicado durante la comprobación de la celda de flujo | La presencia de contaminantes en la biblioteca ha dañado o bloqueado los poros | El número de poros resultante tras la comprobación de la celda de flujo se realiza usando el control de calidad de las moléculas de ADN presentes en el tampón de almacenamiento de la celda de flujo. Al inicio de la secuenciación, se utiliza la misma biblioteca para estimar el número de poros activos. Por este motivo, se estima que puede haber una variabilidad del 10 % en el número de poros detectados. Tener un número de poros considerablemente inferior al inicio de la secuenciación puede deberse a la presencia de contaminantes en la biblioteca que hayan dañado las membranas o bloqueado los poros. Para mejorar la pureza del material de entrada tal vez sea necesario usar métodos de purificación o extracción de ADN/ARN alternativos. Los efectos de los contaminantes están descritos en la página Contaminants. Se recomienda, probar con un método de extracción alternativo que no provoque el arrastre de contaminantes. |
Error en el script de MinKNOW
| Observación | Posible causa | Comentarios y acciones recomendadas |
|---|---|---|
| MinKNOW muestra el mensaje "Error en el script" | Reiniciar el ordenador y reiniciar MinKNOW. Si el problema continúa, reúna los archivos de registro MinKNOW log files y contacte con el servicio de asistencia técnica. Si no dispone de otro dispositivo de secuenciación, recomendamos que guarde la celda de flujo con la biblioteca cargada a 4 °C y contacte con el servicio de asistencia técnica para recibir recomendaciones de almacenamiento adicionales. |
Pore occupancy below 40%
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Pore occupancy <40% | Not enough library was loaded on the flow cell | Ensure you load the recommended amount of good quality library in the relevant library prep protocol onto your flow cell. Please quantify the library before loading and calculate mols using tools like the Promega Biomath Calculator, choosing "dsDNA: µg to pmol" |
| Pore occupancy close to 0 | The Ligation Sequencing Kit was used, and sequencing adapters did not ligate to the DNA | Make sure to use the NEBNext Quick Ligation Module (E6056) and Oxford Nanopore Technologies Ligation Buffer (LNB, provided in the sequencing kit) at the sequencing adapter ligation step, and use the correct amount of each reagent. A Lambda control library can be prepared to test the integrity of the third-party reagents. |
| Pore occupancy close to 0 | The Ligation Sequencing Kit was used, and ethanol was used instead of LFB or SFB at the wash step after sequencing adapter ligation | Ethanol can denature the motor protein on the sequencing adapters. Make sure the LFB or SFB buffer was used after ligation of sequencing adapters. |
| Pore occupancy close to 0 | No tether on the flow cell | Tethers are adding during flow cell priming (FLT/FCT tube). Make sure FLT/FCT was added to FB/FCF before priming. |
Longitud de lectura más corta de lo esperado
| Observación | Posible causa | Comentarios y acciones recomendadas |
|---|---|---|
| Longitud de lectura más corta de lo esperado | Fragmentación no deseada de la muestra de ADN | La longitud de lectura refleja la longitud del fragmento de la muestra de ADN. La muestra de ADN se puede fragmentar durante la extracción de la preparación de la biblioteca. 1. Consulte la sección de buenas prácticas de los métodos de extracción en la página Extraction Methods de la comunidad Nanopore. 2. Visualizar la distribución de la longitud de los fragmentos de las muestras de ADN en un gel de agarosa antes de proceder a la preparación de la biblioteca. 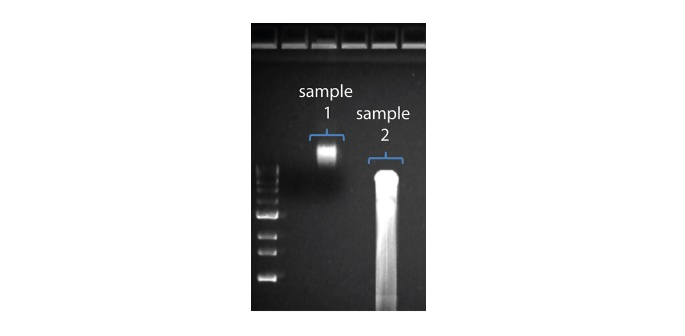 En la imagen superior, la muestra 1 contiene alto peso molecular, mientras que la muestra 2 se ha fragmentado. En la imagen superior, la muestra 1 contiene alto peso molecular, mientras que la muestra 2 se ha fragmentado.3. Durante la preparación de la biblioteca, evitar pipetear y agitar en vórtex cuando se mezclen los reactivos. Dar suaves golpes con el dedo o invertir el vial es suficiente. |
Gran proporción de poros no disponibles
| Observación | Posible causa | Comentarios y acciones recomendadas |
|---|---|---|
Gran proporción de poros no disponibles (se muestran en azul oscuro en el panel de canales y en el gráfico de actividad de poros) 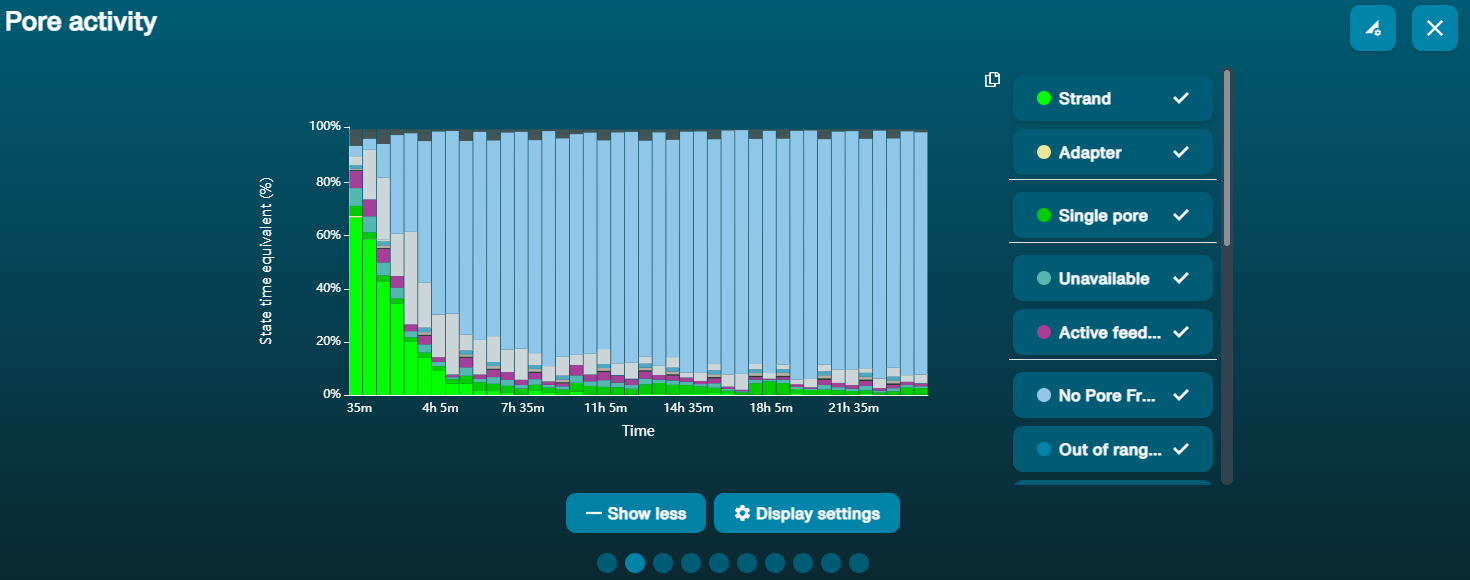 Conforme pasa el tiempo, el gráfico de actividad de poros de arriba muestra una proporción creciente de poros no disponibles. Conforme pasa el tiempo, el gráfico de actividad de poros de arriba muestra una proporción creciente de poros no disponibles. | Hay contaminantes presentes en la muestra | Algunos contaminantes se pueden eliminar de los poros mediante la función de desbloqueo incorporada en MinKNOW. Si funciona, el estado de los poros cambiará a "sequencing pores" (secuenciación de poros). Si la porción poros no disponibles se mantiene elevada o aumenta, pruebe una de las siguientes opciones: 1. Realizar un enjuague de nucleasa con el kit de lavado Flow Cell Wash Kit (EXP-WSH004) 2. Realizar varios ciclos de PCR para intentar diluir cualquier contaminante que pueda estar causando problemas. |
Gran proporción de poros inactivos
| Observación | Posible causa | Comentarios y acciones recomendadas |
|---|---|---|
| Gran proporción de poros inactivos/no disponibles (se muestran en azul claro en el panel de canales y en el gráfico de actividad de poros. Los poros o membranas están dañados de manera irreversible) | Se han introducido burbujas de aire en la celda de flujo | Las burbujas de aire introducidas durante el cebado de la celda y la carga de la biblioteca pueden dañar los poros de forma permanente. Para conocer las buenas prácticas de cebado y carga de la celda de flujo, ver el vídeo Priming and loading your flow cell |
| Gran proporción de poros inactivos/no disponibles | Ciertos compuestos copurificados con ADN | Compuestos conocidos, incluidos los polisacáridos, se asocian generalmente con el ADN genómico de las plantas. 1. Consulte la página Plant leaf DNA extraction method. 2. Limpiar usando el kit QIAGEN PowerClean Pro. 3. Realizar una amplificación del genoma completo con la muestra original de ADNg utilizando el kit QIAGEN REPLI-g. |
| Gran proporción de poros inactivos/no disponibles | Hay contaminantes presentes en la muestra | Los efectos de los contaminantes se muestran en la página Contaminants. Probar con un método de extracción alternativo que no provoque el arrastre de contaminantes. |
Reducción de la velocidad de secuenciación y del índice de calidad Qscore en una fase avanzada de la secuenciación
| Observación | Posible causa | Comentarios y acciones recomendadas |
|---|---|---|
| Reducción de la velocidad de secuenciación y el índice de calidad Qscore en una fase avanzada de la secuenciación | En la química del kit 9 (p. ej., SQK-LSK109), cuando la celda de flujo está sobrecargada con la biblioteca se observa un consumo rápido de combustible (consulte el protocolo correspondiente a su biblioteca de ADN para ver las recomendaciones) | Añadir más combustible a la celda de flujo, siguiendo las instrucciones en el protocolo de MinKNOW. En futuros experimentos, cargar cantidades menores de biblioteca en la celda de flujo. |
Fluctuación de la temperatura
| Observación | Posible causa | Comentarios y acciones recomendadas |
|---|---|---|
| Fluctuación de la temperatura | La celda de flujo ha perdido contacto con el dispositivo | Comprobar que una almohadilla térmica cubra la placa metálica de la parte posterior de la celda de flujo. Reinsertar la celda de flujo y presionar para asegurarse de que las clavijas del conector están bien conectadas al dispositivo. Si el problema continúa, contacte con el servicio de asistencia técnica. |
Error al intentar alcanzar la temperatura deseada
| Observación | Posible causa | Comentarios y acciones recomendadas |
|---|---|---|
| MinKNOW muestra el mensaje "Error al intentar alcanzar la temperatura deseada" | El dispositivo ha sido colocado en un lugar a una temperatura ambiente inferior a la media o en un lugar con escasa ventilación (lo que provoca el sobrecalientamiento de las celdas de flujo). | MinKNOW tiene un tiempo predeterminado para que las celdas de flujo alcancen la temperatura fijada. Una vez acabado el tiempo, aparece un mensaje de error, pero el experimento de secuenciación continua. Secuenciar a una temperatura incorrecta puede llevar a una disminución en el rendimiento y a generar un índice de calidad Qscore menor. Corrija la ubicación del dispositivo de secuenciación para asegurarse de que se encuentra a temperatura ambiente y con buena ventilación; a continuación, reinicie el proceso en MinKNOW. Para obtener más información sobre el control de temperatura de MinKNOW Mk 1B, consulte la sección de preguntas frecuentes, FAQ. |
Guppy – no input .fast5 was found or basecalled
| Observation | Possible cause | Comments and actions |
|---|---|---|
| No input .fast5 was found or basecalled | input_path did not point to the .fast5 file location | The --input_path has to be followed by the full file path to the .fast5 files to be basecalled, and the location has to be accessible either locally or remotely through SSH. |
| No input .fast5 was found or basecalled | The .fast5 files were in a subfolder at the input_path location | To allow Guppy to look into subfolders, add the --recursive flag to the command |
Guppy – no Pass or Fail folders were generated after basecalling
| Observation | Possible cause | Comments and actions |
|---|---|---|
| No Pass or Fail folders were generated after basecalling | The --qscore_filtering flag was not included in the command | The --qscore_filtering flag enables filtering of reads into Pass and Fail folders inside the output folder, based on their strand q-score. When performing live basecalling in MinKNOW, a q-score of 7 (corresponding to a basecall accuracy of ~80%) is used to separate reads into Pass and Fail folders. |
Guppy – unusually slow processing on a GPU computer
| Observation | Possible cause | Comments and actions |
|---|---|---|
| Unusually slow processing on a GPU computer | The --device flag wasn't included in the command | The --device flag specifies a GPU device to use for accelerate basecalling. If not included in the command, GPU will not be used. GPUs are counted from zero. An example is --device cuda:0 cuda:1, when 2 GPUs are specified to use by the Guppy command. |
