NCM 2022: SeqScreen-Nano: functional and taxonomic characterization of long read metagenomic data
- shared.published_on: December 7 2022
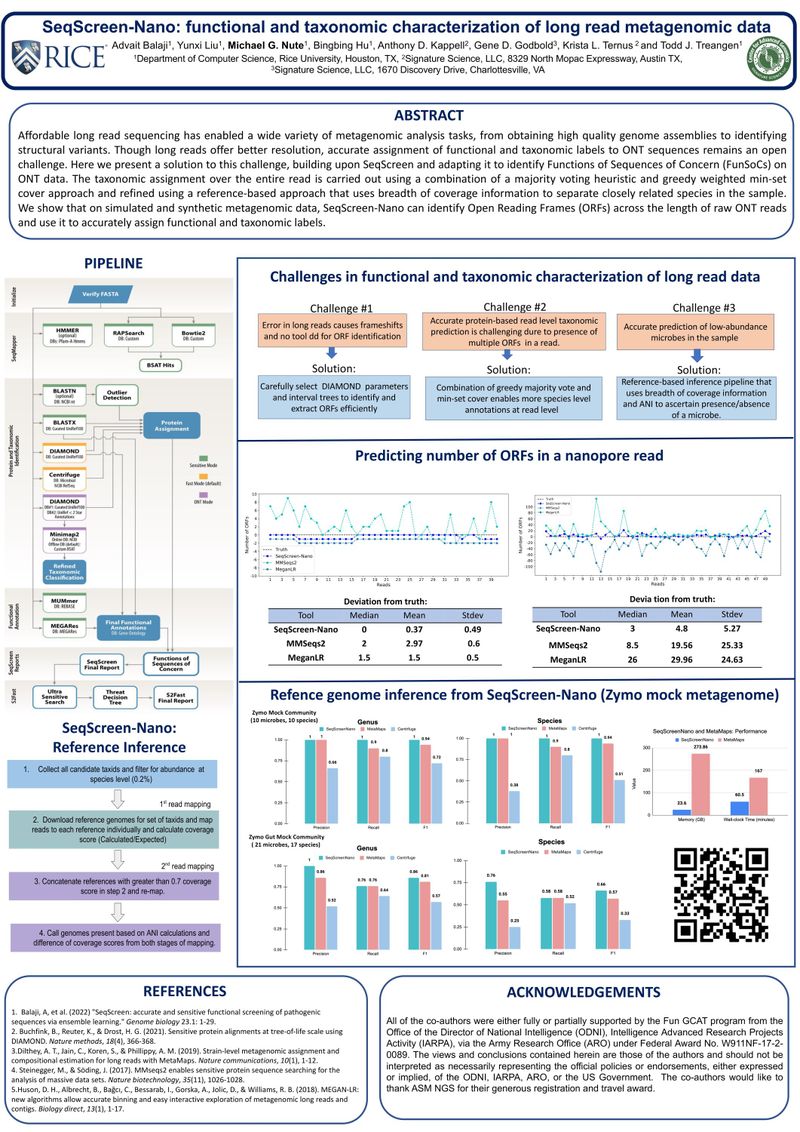
Affordable long read sequencing has enabled a wide variety of metagenomic analysis tasks, from obtaining high quality genome assemblies to identifying structural variants. Though long reads offer better resolution, accurate assignment of functional and taxonomic labels to Oxford Nanopore sequences remains an open challenge. Here we present a solution to this challenge, building upon SeqScreen and adapting it to identify Functions of Sequences of Concern (FunSoCs) on Oxford Nanopore data. The taxonomic assignment over the entire read is carried out using a combination of a majority voting heuristic and greedy weighted min-set cover approach and refined using a reference-based approach that uses breadth of coverage information to separate closely related species in the sample. We show that on simulated and synthetic metagenomic data, SeqScreen-Nano can identify Open Reading Frames (ORFs) across the length of raw Oxford Nanopore reads and use it to accurately assign functional and taxonomic labels.






