NanoAmpli-Seq: A workflow for amplicon sequencing from mixed microbial communities on the nanopore sequencing platform
- shared.published_on: June 4 2018
- shared.source: London Calling 2018
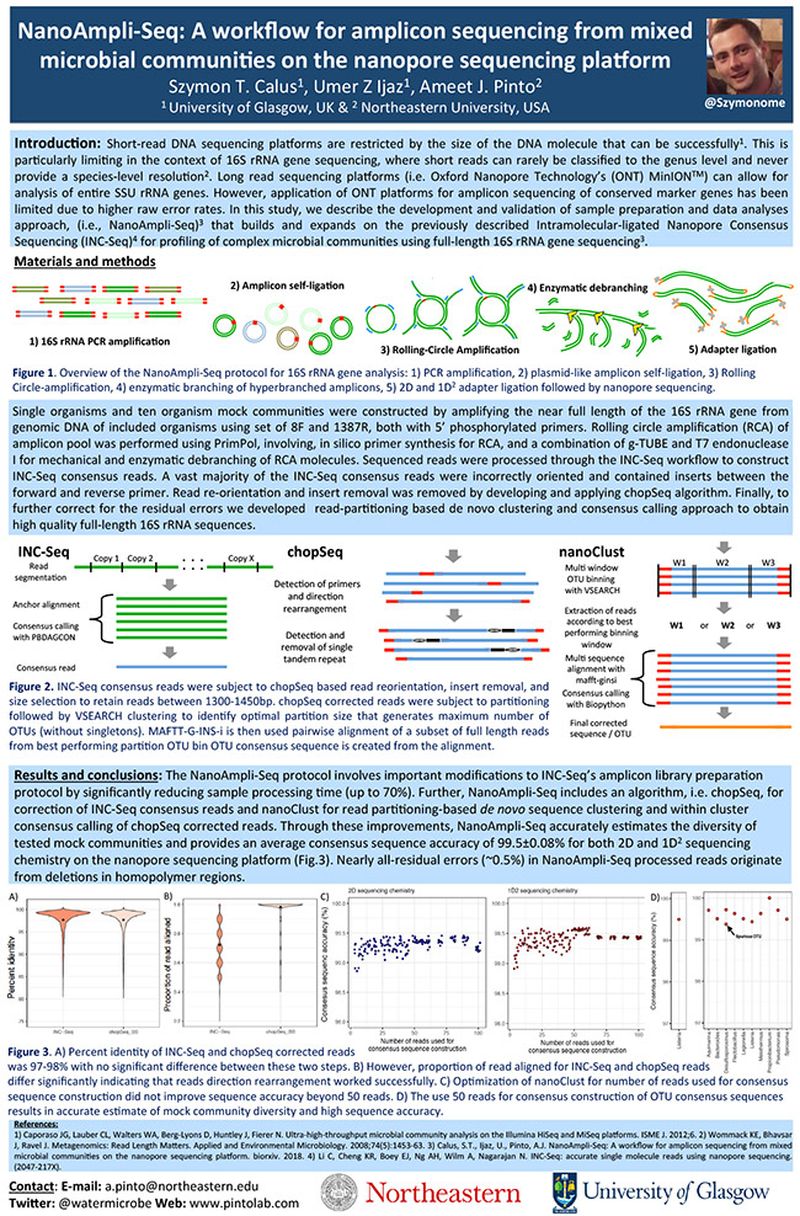
Short-‐read DNA sequencing platforms are restricted by the size of the DNA molecule that can be successfully1. This is particularly limiting in the context of 16S rRNA gene sequencing, where short reads can rarely be classified to the genus level and never provide a species-‐level resolution2. Long read sequencing platforms (i.e. Oxford Nanopore Technology’s (ONT) MinIONTM) can allow for analysis of entire SSU rRNA genes. However, application of ONT platforms for amplicon sequencing of conserved marker genes has been limited due to higher raw error rates. In this study, we describe the development and validation of sample preparation and data analyses approach, (i.e., NanoAmpli-Seq)3 that builds and expands on the previously described Intramolecular-‐ligated Nanopore Consensus Sequencing (INC-Seq)4 for profiling of complex microbial communities using full-‐length 16S rRNA gene sequencing3.






