Mapping the yeast mitochondrial transcriptomes by direct RNA sequencing
- shared.published_on: June 4 2018
- shared.source: London Calling 2018
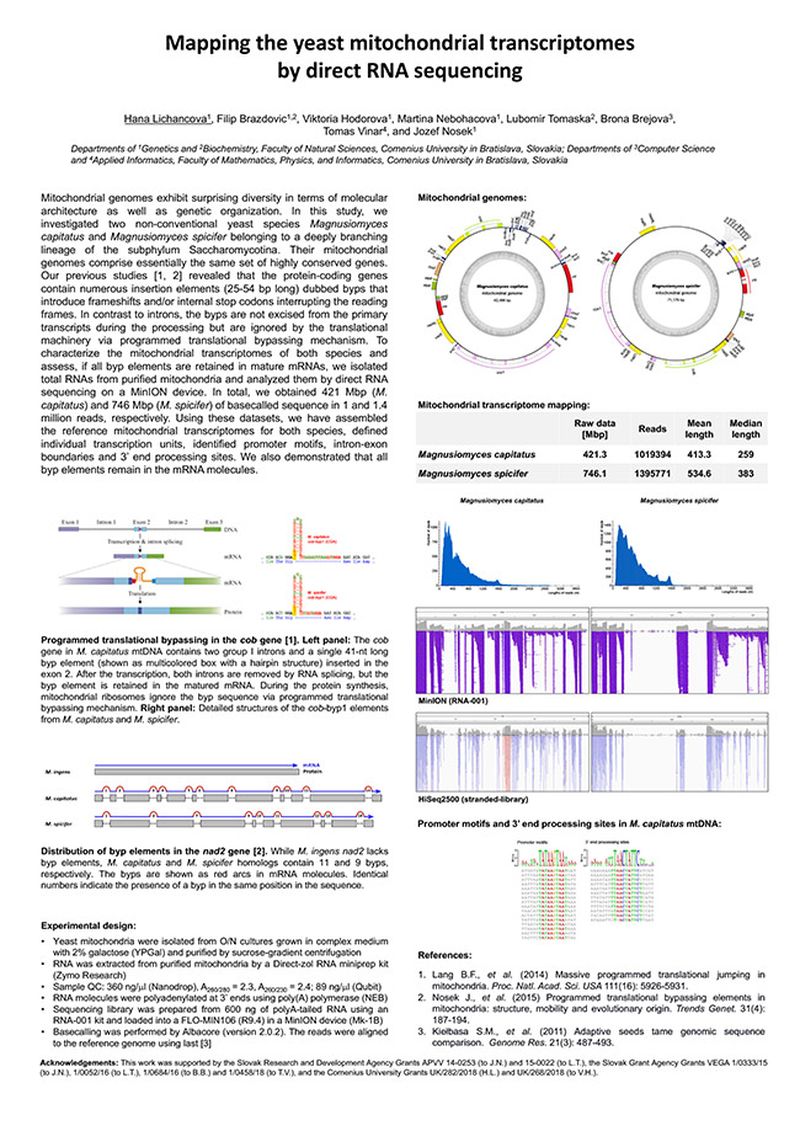
Mitochondrial genomes exhibit surprising diversity in terms of molecular architecture as well as genetic organization. In this study, we investigated two non-conventional yeast species Magnusiomyces capitatus and Magnusiomyces spicifer belonging to a deeply branching lineage of the subphylum Saccharomycotina. Their mitochondrial genomes comprise essentially the same set of highly conserved genes. Our previous studies [1, 2] revealed that the protein-coding genes contain numerous insertion elements (25-54 bp long) dubbed byps that introduce frameshifts and/or internal stop codons interrupting the reading frames. In contrast to introns, the byps are not excised from the primary transcripts during the processing but are ignored by the translational machinery via programmed translational bypassing mechanism. To characterize the mitochondrial transcriptomes of both species and assess, if all byp elements are retained in mature mRNAs, we isolated total RNAs from purified mitochondria and analyzed them by direct RNA sequencing on a MinION device. In total, we obtained 421 Mbp (M. capitatus) and 746 Mbp (M. spicifer) of basecalled sequence in 1 and 1.4 million reads, respectively. Using these datasets, we have assembled the reference mitochondrial transcriptomes for both species, defined individual transcription units, identified promoter motifs, intron-exon boundaries and 3’ end processing sites. We also demonstrated that all byp elements remain in the mRNA molecules.






