SageHLS CATCH Workflow for Targeted Nanopore Sequencing
- shared.published_on: June 4 2018
- shared.source: London Calling 2018
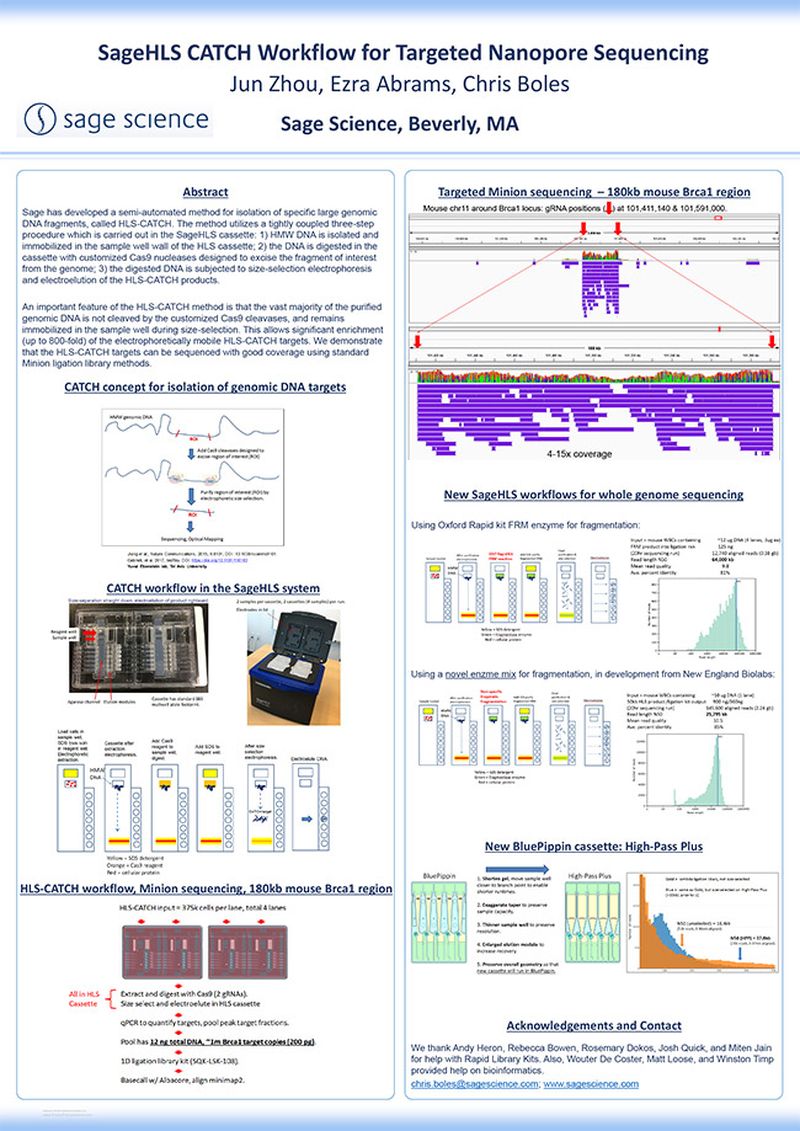
Sage has developed a semi-automated method for isolation of specific large genomic DNA fragments, called HLS-CATCH. The method utilizes a tightly coupled three-step procedure which is carried out in the SageHLS cassette: 1) HMW DNA is isolated and immobilized in the sample well wall of the HLS cassette; 2) the DNA is digested in the cassette with customized Cas9 nucleases designed to excise the fragment of interest from the genome; 3) the digested DNA is subjected to size-selection electrophoresis and electroelution of the HLS-CATCH products.
An important feature of the HLS-CATCH method is that the vast majority of the purified genomic DNA is not cleaved by the customized Cas9 cleavases, and remains immobilized in the sample well during size-selection. This allows significant enrichment (up to 800-fold) of the electrophoretically mobile HLS-CATCH targets. We demonstrate that the HLS-CATCH targets can be sequenced with good coverage using standard Minion ligation library methods.






