Pathogen detection and beyond: understanding of microbial communities in the real world
- shared.published_on: September 4 2019
- shared.source: MPMI 2019
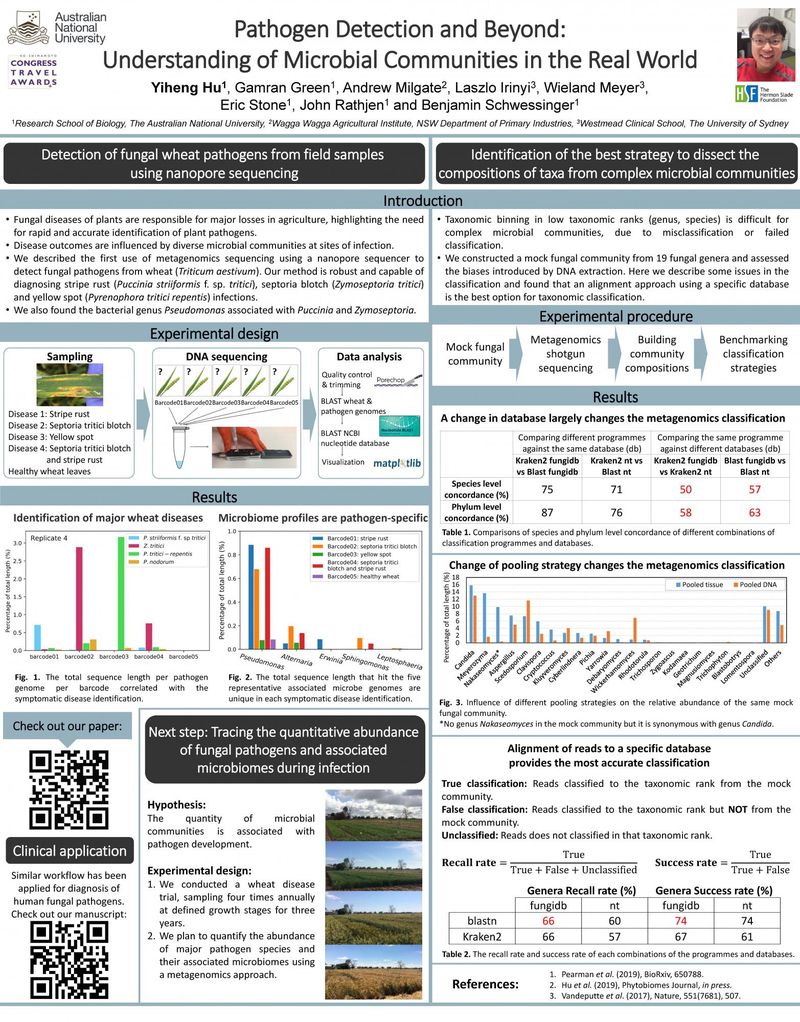
Detection of fungal wheat pathogens from field samples using nanopore sequencing
- Fungal diseases of plants are responsible for major losses in agriculture, highlighting the need for rapid and accurate identification of plant pathogens.
- Disease outcomes are influenced by diverse microbial communities at sites of infection
- We described the first use of metagenomics sequencing using a nanopore sequencer to detect fungal pathogens from wheat (Triticum aestivum). Our method is robust and capable of diagnosing stripe rust (Puccinia striiformis f. sp. tritici), septoria blotch (Zymoseptoria tritici) and yellow spot (Pyrenophora tritici repentis) infections
- We also found the bacterial genus Pseudomonas associated with Puccinia and Zymoseptoria
Identification of the best strategy to dissect the compositions of taxa from complex microbial communities
- Taxonomic binning in low taxonomic ranks (genus, species) is difficult for complex microbial communities, due to misclassification or failed classification
- We constructed a mock fungal community from 19 fungal genera and assessed the biases introduced by DNA extraction. Here we describe some issues in the classification and found that an alignment approach using a specific database is the best option for taxonomic classification






