DNA methylation analysis in human tumour samples using nanopore sequencing
- shared.published_on: May 28 2022
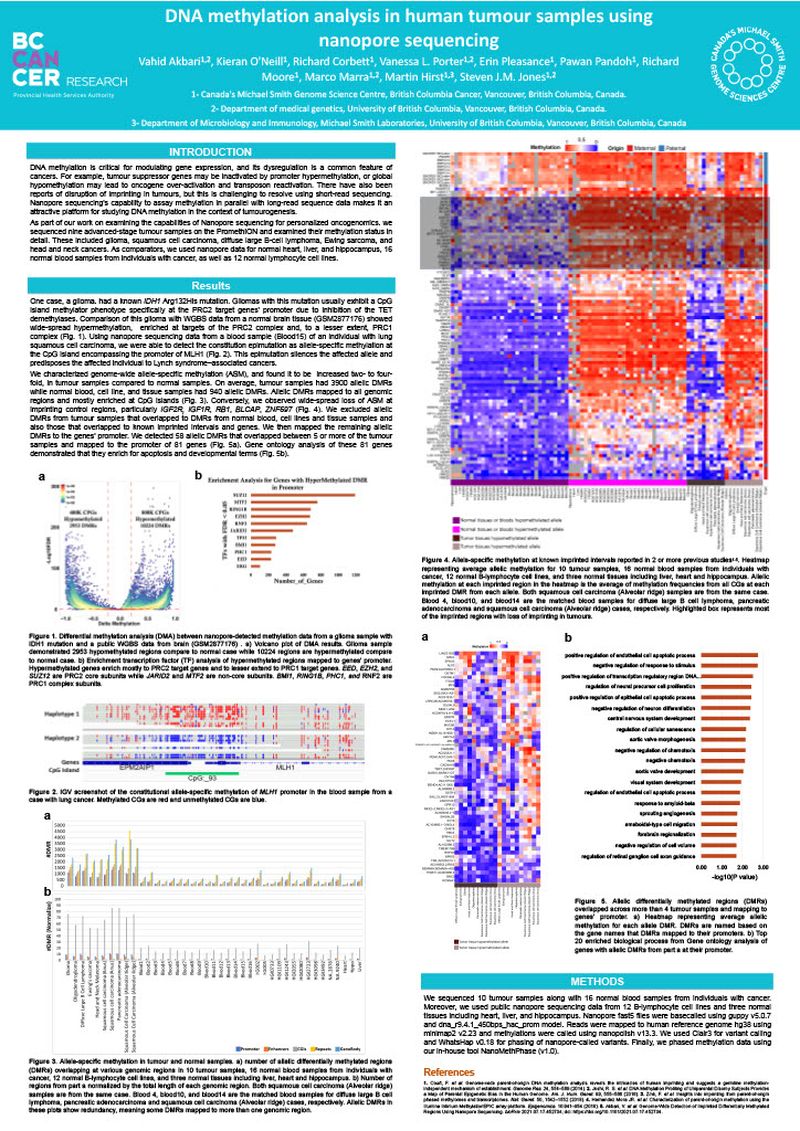
DNA methylation is critical for modulating gene expression, and its dysregulation is a common feature of cancers. For example, tumour suppressor genes may be inactivated by promoter hypermethylation, or global hypomethylation may lead to oncogene over-activation and transposon reactivation. There have also been reports of disruption of imprinting in tumours, but this is challenging to resolve using short-read sequencing. Nanopore sequencing’s capability to assay methylation in parallel with long-read sequence data makes it an attractive platform for studying DNA methylation in the context of tumourogenesis. As part of our work on examining the capabilities of Nanopore sequencing for personalized oncogenomics. We sequenced nine advanced-stage tumour samples on the PromethION and examined their methylation status in detail. These included glioma, squamous cell carcinoma, diffuse large B-cell lymphoma, Ewing sarcoma, and head and neck cancers. As comparators, we used nanopore data for normal heart, liver, and hippocampus, 16 normal blood samples from individuals with cancer, as well as 12 normal lymphocyte cell lines.






