De novo Identification of DNA Modifications Enabled by Genome-Guided Nanopore Signal Processing
- shared.published_on: April 10 2017
- shared.source: BioRxiv
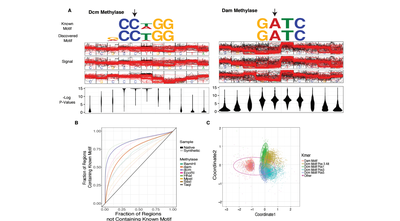
Advances in nanopore sequencing technology have enabled investigation of the full catalogue of covalent DNA modifications. We present the first algorithm for the identification of modified nucleotides without the need for prior training data along with the open source software implementation, nanoraw. Nanoraw accurately assigns contiguous raw nanopore signal to genomic positions, enabling novel data visualization, and increasing power and accuracy for the discovery of covalently modified bases in native DNA. Ground truth case studies utilizing synthetically methylated DNA show the capacity to identify three distinct methylation marks, 4mC, 5mC, and 6mA, in seven distinct sequence contexts without any changes to the algorithm. We demonstrate quantitative reproducibility simultaneously identifying 5mC and 6mA in native E. coli across biological replicates processed in different labs. Finally we propose a pipeline for the comprehensive discovery of DNA modifications in any genome without a priori knowledge of their chemical identities.