Science Unlocked: publication picks from March 2026
In this monthly series, we share a selection of recent publications that use Oxford Nanopore sequencing to unlock novel insights. Spanning pathogen surveillance, genome assembly, and arthritis research, these studies showcase the advances in scientific research made possible by Oxford Nanopore sequencing.
Featured in this edition:
1. Uncovering hidden norovirus transmission in hospitals
2. Complete and accurate nanopore-only genome assemblies
3. Jumping genes may drive chromosomal instability in cancer
4. Assessing RNA quality for gene therapy applications
5. Transcriptomics highlights mechanisms of cell death in arthritis
Infectious disease
1. Identifying norovirus transmission in hospitals using whole-genome sequencing (Genome Medicine)
Norovirus gastroenteritis is a leading cause of death in children under five globally, but traditional strain identification methods such as Sanger sequencing can't resolve closely related outbreak strains. Oxford Nanopore sequencing could transform our understanding of how norovirus spreads within hospitals, and this research shows just how much detail has been out of reach. By sequencing over 700 stool samples, Grimes and Fink et al. generated near-complete viral genomes and paired them with medical record data to map transmission in unprecedented detail.
This high-resolution view revealed not only dominant and emerging strains, but also uncovered hidden transmission chains linking patients across wards, clinical spaces, and care teams. Nearly a third of closely related viral clusters pointed to likely in-hospital transmission events. It’s a powerful reminder that when you can see the genome end to end, you can start connecting the dots in real time, opening up the potential for earlier outbreak detection and more targeted infection control.
‘This work may advance viral strain tracking in hospitalised patients to uncover interruptible transmission patterns and inform best practices for preventing norovirus GII spread’
Grimes, L. and Fink, J. et al.1
Want to unlock even richer insights? Watch our webinar on metagenomics for pathogen surveillance.
2. Nanopore-only genome assembly of clinical Enterobacterales isolates is complete and accurate (Microbial Genomics)
What if you could assemble complete, high-quality bacterial genomes without needing to reach for short reads? In this study, Nagy et al. put nanopore-only sequencing to the test across 92 Enterobacterales isolates, and the results are hard to ignore. Using nanopore sequencing with super accuracy basecalling, they generated genome assemblies that were not only highly accurate, but often more structurally complete than hybrid approaches. With up to 95% of chromosomes fully circularised and over 90% of plasmids recovered, the approach captures the key elements for understanding antimicrobial resistance.
With Autocycler assembly and Medaka polishing, the workflow delivered near-error-free genomes with consistent detection of resistance and virulence genes that was indistinguishable from hybrid assemblies. This research is a compelling step towards simpler, more scalable isolate sequencing, where nanopore reads can give you the complete picture and help bring high-resolution pathogen surveillance within reach.
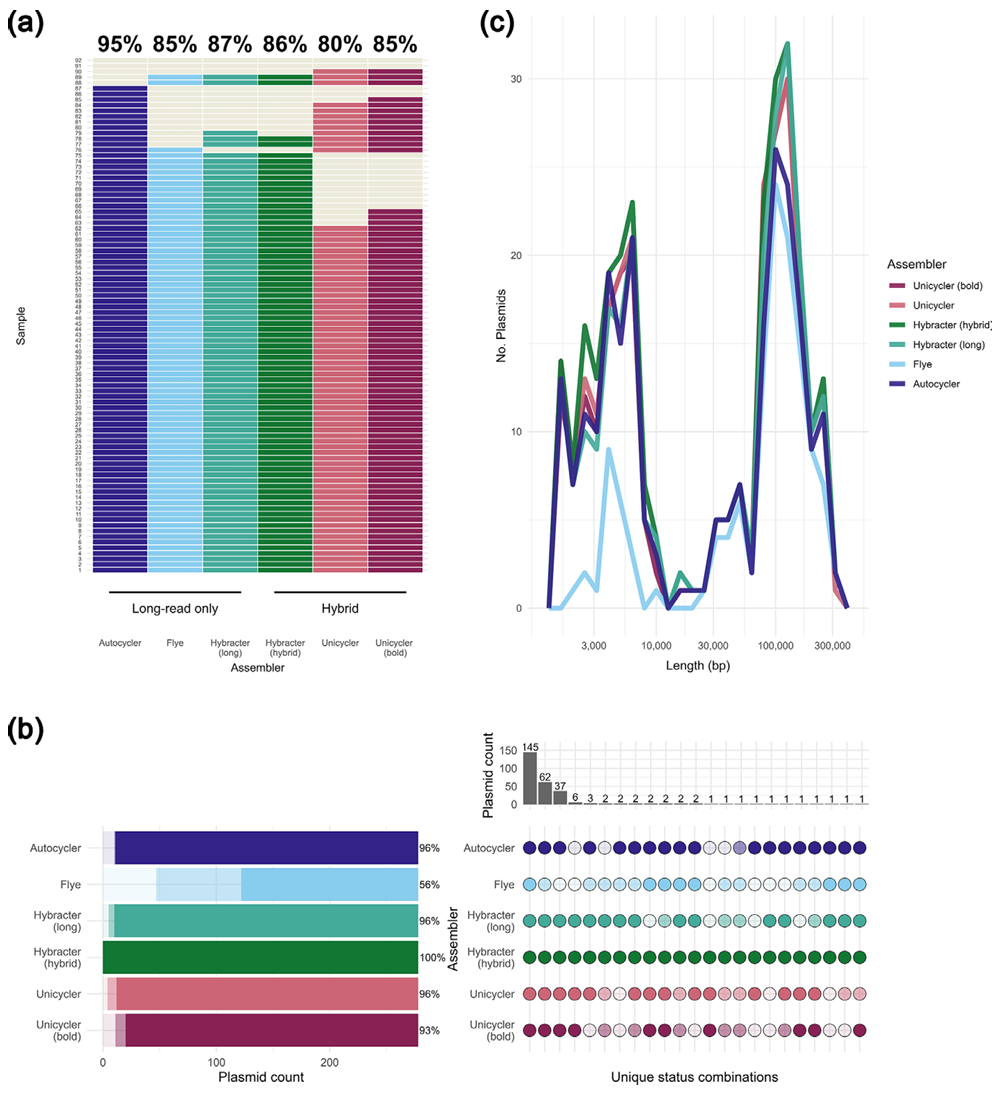
Figure 1: Structural completeness of 92 Enterobacterales genomes assembled using long-read-only (Oxford Nanopore) and hybrid (Oxford Nanopore and Illumina) approaches. (a) Proportion of fully circularised chromosomes achieved by each assembler. (b) Comparison of plasmid reconstruction across assemblers against a hybrid reference set, showing correctly assembled (dark), misassembled (medium), or missing (light) plasmids. (c) Length distribution of correctly reconstructed plasmids for each assembly method. Figure redistributed from Nagy et al. 20262 under Creative Commons Attribution License CC BY 4.0.
‘This is the largest benchmarking study to date, using 92 clinical Enterobacterales isolates, to demonstrate structural completeness and accuracy of nanopore super-high accuracy long-read-only bacterial genome assembly compared with hybrid assembly’
Nagy, D. et al.2
Explore our nanopore-only microbial isolate sequencing solution (NO-MISS) and watch our knowledge exchange.
Cancer research
3. Concurrent LINE-1 insertions promote structural changes in tumour development (Science)
Some of the most disruptive mutations in cancer may not come from external factors — but from our own genome moving around. In this study, Zumalave et al. used nanopore sequencing to capture the full scale and structure of long interspersed nuclear element-1 (LINE-1; L1) activity, revealing thousands of insertions and a level of genomic reshaping that’s been largely hidden until now.
By spanning these events in single reads, our technology uncovers how multiple L1 insertions can act together to drive complex rearrangements, including reciprocal translocations and inversions that short-read approaches often miss. Strikingly, many of these events occur early in tumour development, positioning L1 activity as a potential driver of chromosomal instability. In practice, our technology could allow you to move beyond what’s visible with short reads to unlock a deeper understanding of how cancer genomes evolve.
‘Approximately three-quarters of the retrotransposon-mediated rearrangements identified in this work would remain undetected or misclassified by short-read sequencing, which underscores how this class of structural variation has been largely overlooked’
Zumalave et al.3
Find out how you can unlock transformative cancer insights.
Biopharma
4. Assessing RNA quality for cell and gene therapy applications using nanopore direct RNA sequencing (Analytical Chemistry)
RNA therapeutics are pushing boundaries, and quality control needs to keep pace. Here, Chatla et al. used nanopore direct RNA sequencing to assess all the critical quality attributes in a single assay. This provides a complete view of the messenger RNA (mRNA) and single-guide RNA (sgRNA) from sequence identity and integrity, to poly(A) tail length and 5′ capping efficiency, all measured directly from native RNA molecules using a streamlined workflow with a fast turnaround time and simple sample prep.
This research also moves into new territory, enabling full-length sequencing of non-polyadenylated sgRNAs by incorporating a 5’ RNA oligo adapter. For future RNA-based therapies, our technology has the potential to support quality assurance — helping you move from fragmented testing to a single, confident view of RNA quality.
‘Nanopore direct RNA sequencing has emerged as a powerful single-molecule analytical approach capable of simultaneously resolving sequence and structural features consistent with regulatory expectations’
Chatla, K. et al.4
Discover our solutions for cell and gene therapies.
Human genetics
5. Integrated transcriptomic profiling identifies mechanisms associated with programmed cell death in arthritis (Frontiers in Immunology)
Ankylosing spondylitis (AS) is an early onset arthritis with no known cause or cure, but what if the missing piece in understanding AS isn’t only which genes are involved, but also how they’re being used? In this study, Cao and Li et al. used nanopore cDNA sequencing to capture full-length transcripts, revealing gene expression, isoform switching, and alternative splicing in unprecedented detail.
They found complex underlying immune dysregulation: over a thousand genes and nearly two thousand transcripts were altered, with strong signals pointing to disrupted programmed cell death (PCD) pathways. Crucially, these changes don’t happen in isolation; coordinated changes across gene expression and isoform usage suggest that tightly linked regulatory networks are driving inflammation. For researchers, it’s a shift from a flat transcriptomic snapshot to a multi-dimensional map — one that could help you pinpoint more precise biomarkers and potentially uncover new therapeutic angles in complex immune diseases.
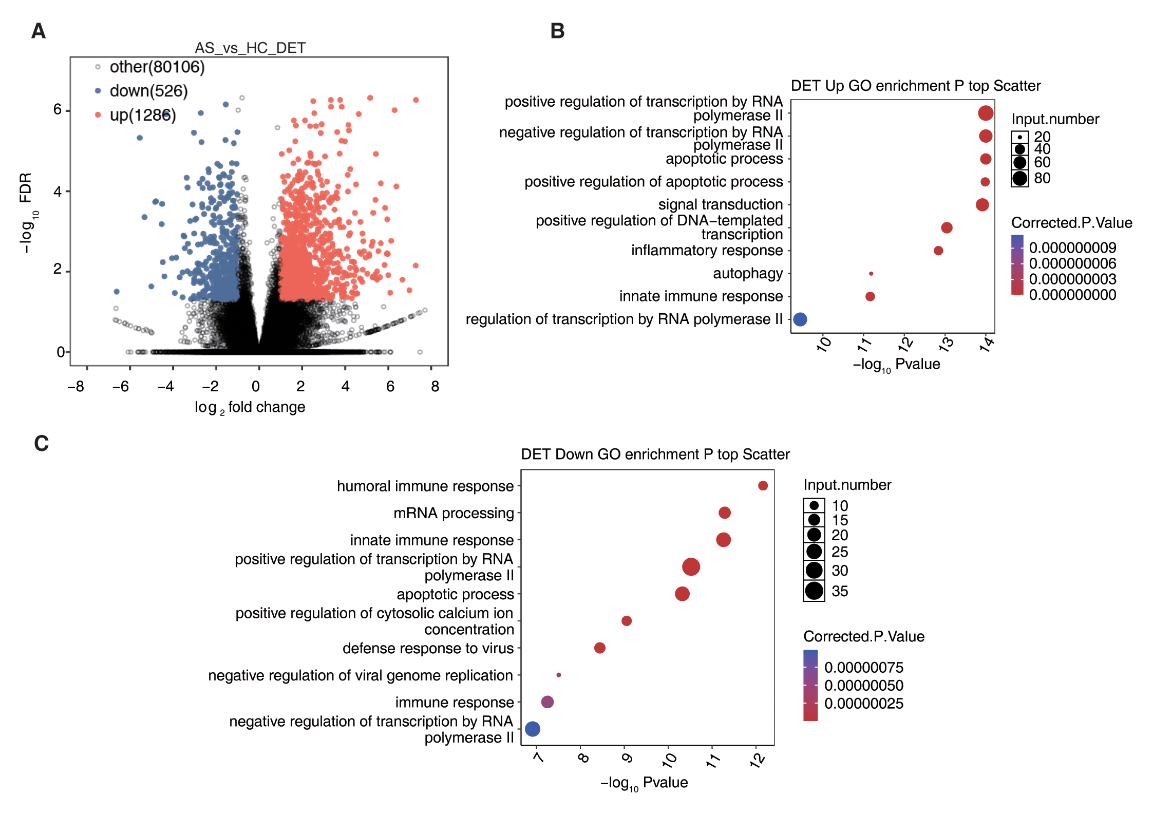
Figure 2: Oxford Nanopore sequencing reveals the transcript expression landscape in AS. (A) Volcano plot showing differentially expressed transcripts (DET), 1,286 were upregulated and 526 were downregulated. (B) Upregulated DETs were significantly associated with transcriptional regulation via RNA polymerase II, apoptotic signalling, inflammation, autophagy, and innate immune activation. (C) Downregulated DETs were enriched in biological processes including humoral immune response, mRNA splicing and processing, antiviral defence, and regulation of cytoplasmic calcium ion levels. Figure redistributed from Cao and Li et al. 20265 under Creative Commons Attribution License CC BY 4.0.
‘Our [nanopore] long-read transcriptomic profiling of AS provides novel insights into the multilayered regulation of gene activity, especially as it pertains to PCD-related pathways’
Cao and Li et al.5
Check out our getting started guide for transcriptomics.
Inspired? Apply Oxford Nanopore sequencing to your own research questions and you'll never see sequencing the same way again. Explore the nanopore sequencing solution.
Oxford Nanopore Technologies products are not intended for use for health assessment or to diagnose, treat, mitigate, cure, or prevent any disease or condition.
Grimes, L. and Fink, J. et al. Identifying intra-hospital norovirus GII transmission using whole-genome sequencing. Genome Med. (2026). DOI: https://doi.org/10.1186/s13073-026-01621-1
Nagy, D. et al. Nanopore long-read-only genome assembly of clinical Enterobacterales isolates is complete and accurate. Microbial Genomics 12(2): 001631 (2026). DOI: https://doi.org/10.1099/mgen.0.001631
Zumalave, S. et al. Concurrent L1 retrotransposition events promote reciprocal translocations in human tumorigenesis. Science 0, eaee4513 (2026). DOI: https://doi.org/10.1126/science.aee4513
Chatla, K. et al. Assessing mRNA and sgRNA quality for cell and gene therapy applications using nanopore direct RNA sequencing. Analytical Chemistry 98 (10), 7452-7461 (2026). DOI: https://doi.org/10.1021/acs.analchem.5c06819
Cao, X. and Li, P. et al. Integrated long-read transcriptomic profiling of peripheral blood from ankylosing spondylitis patients identifies regulatory shifts and core genes associated with programmed cell death. Front. Immunol. 17 (2026). DOI: https://doi.org/10.3389/fimmu.2026.1640271






