Science Unlocked: publication picks from February 2026
In this monthly series, we share a selection of recent publications that use Oxford Nanopore sequencing to unlock novel insights. Spanning colorectal cancer, telomere-to-telomere (T2T) assembly, and sequencing in low-resource settings, these studies showcase the advances in scientific research made possible by Oxford Nanopore technology.
Featured in this edition:
1. Near-T2T genome assembly without ultra-long reads
2. Rapid DNA extraction, purification, and library prep without a pipette
3. A skipped exon with potentially cancerous consequences
4. Removing barriers to pathogen detection
5. Accurate and scalable rare disease resolution with Adaptive Sampling
Bioinformatics
1. Efficient near-T2T assembly of nanopore reads (Nature)
We are one step closer to true T2T de novo genome assembly, thanks to this study from Cheng and Qu et al. Previous methods for delivering T2T assemblies have relied on additional sample prep with high input requirements to generate ultra-long reads, which need around 40 times more input DNA than standard protocols. Instead, hifiasm (ONT) uses phasing-based error correction to determine true variants, enabling near-T2T genome assemblies with standard Oxford Nanopore data without ultra-long reads.
In this study, hifiasm (ONT) outperformed Verkko (plus HERRO for error correction), without the need for complex hybrid assembly methods. Delivering high contiguity at low cost and computational demand, hifiasm (ONT) could broaden access to high-quality assemblies across research applications. In the not-too-distant future, population-scale genome reconstruction from a single dataset could be within reach.
‘By using the potential of [Oxford Nanopore] simplex sequencing fully, hifiasm (ONT) achieves superior performance over other assemblers ... with [Oxford Nanopore] ultra-long simplex reads, hifiasm (ONT) reconstructs more human chromosomes from telomere to telomere than any other approach’
Cheng, H. and Qu, H. et al.1
Read our news summary or catch the talk by lead algorithm designer Haoyu Cheng at London Calling 2025 to find out how we’re helping researchers find the missing piece of the puzzle for T2T assembly.
Cancer research
2. An ultra-fast sample-to-sequencing workflow using a microfluidic cartridge (medRxiv)
Bisogni et al. are rethinking how DNA is prepared for nanopore sequencing with their microfluidic cartridge that integrates extraction, purification, and library preparation. Many people want to sequence long DNA fragments, but traditional library preparation often has multiple steps that can lead to excess fragmentation and DNA loss. Plus, significant hands-on time and training are required, creating a barrier to access and limiting DNA prep to a lab.
By developing a microfluidic cartridge with a robust workflow, the team rapidly produced libraries ready for nanopore sequencing across a range of tissue types — from whole blood to flash-frozen brain tumour research samples. From as little as a drop of blood, the researchers recovered high-molecular-weight DNA for nanopore sequencing with minimal hands-on steps, demonstrating a potentially powerful solution to improve library prep speed and access to DNA sequencing.
‘[The] micropillar workflow [is] particularly relevant for point-of-care diagnostics, pediatric applications, and resource-limited settings where phlebotomy infrastructure may be unavailable’
Bisogni, A.J. et al.2
Learn how you can comprehensively characterise genes implicated in cancer predisposition in our Hereditary Cancer Panel workflow.
3. Exon skipping as a potential diagnostic biomarker in colorectal cancer (BMC Human Genomics)
Colorectal cancer is one of the most common cancers in the world, yet pathogenic mechanisms are poorly understood. Teasing apart how splicing and RNA modifications interact in this disease has been technically out of reach — until now. In this study, Zhang et al. used Oxford Nanopore direct RNA sequencing to resolve both processes simultaneously, revealing a potential new diagnostic biomarker: exon skipping in MYH11, a gene associated with colorectal cancer.
The team also identified widespread variation in RNA modifications enriched in exons and 3′ untranslated regions (UTRs), mapping more than 300 potentially regulatory pairs connecting modification sites to splicing changes. This includes two that may regulate MYH11 splicing. Although the researchers stress that independent verification with large cohorts is needed to confirm the utility of these targets, this is a compelling example of how nanopore reads can shine a light on previously invisible details.
‘Compared to short read sequencing, nanopore sequencing significantly improved accuracy in detecting aberrant [alternative splicing events]’
Zhang, L., Cui, J., and Shi, J. et al.3
Reveal more biology to transform human health with RNA and cDNA sequencing.
Infectious disease
4. Improving local sequencing diagnostic capacity for pathogen identification in Northern Nigeria (Access Microbiology)
Timely identification of bacterial pathogens is essential for effective antimicrobial therapy, but this can be hard in low-resource settings that rely on external, centralised laboratories. However, nanopore sequencing is rewriting the rules of sequencing.
In a hands-on workshop run by Bastug, Umar, Okolo, and Thielen, et al., they demonstrated a low-cost workflow to detect pathogens and antimicrobial resistance (AMR) genes. They used a MinION, a low-cost salt-based DNA extraction method, and open-source bioinformatics tools to develop a practical workflow, generating high-quality data for ~$23.68 per sample. They were even able to reclassify two Escherichia coli isolates that were originally misidentified as Salmonella. Thanks to accessible technology, nanopore sequencing has the potential to deliver faster answers, fewer misclassifications, and stronger local AMR surveillance than other current technologies.
‘Our findings support nanopore sequencing as a practical and promising tool for strengthening diagnostic microbiology and antimicrobial resistance surveillance in low-resource settings’
Bastug, K.A., Umar, U., Okolo, M., and Thielen, B.K. et al.4
Discover sequencing that keeps up with you, from low to high throughput.
Human genetics
5. Genetic characterisation of variants in CYP21A2 with Adaptive Sampling (European Journal of Human Genetics)
Sanger sequencing is often used as a clinical diagnostic test, but it is time-consuming, costly, and doesn’t reliably detect complex structural variants. For example, it is used to diagnose congenital adrenal hyperplasia (CAH), caused by variants in the CYP21A2 gene. However, the gene is highly homologous, recombination-prone, and located next to its inactive pseudogene, so it is challenging to correctly assign disease-causing variants to the functional or inactive gene with Sanger sequencing. Digital panels, powered by Adaptive Sampling — our unique software-based target-enrichment method — could help reveal the hidden causes of rare diseases with easy access to PCR-free long reads.
In this study, Lildballe et al. targeted chromosome six with Adaptive Sampling and long-read sequencing (AS-LRS) and used a custom mapping tool (NanoCAH) to achieve a 94% diagnostic yield across 34 patients previously clinically diagnosed with CAH. Adaptive Sampling was capable of resolving chimeric alleles, large deletions, gene conversions, and previously undetected missense variants, and even reliably phased alleles without parental testing. For complex genomic regions where PCR falters, this study demonstrates Adaptive Sampling as a potentially ‘promising tool not only for CAH diagnosis, but also for genetic testing'.
‘Compared to current gold standard methods, AS-LRS proved to be faster and more scalable, while providing greater accuracy in detecting variants within the CYP21A2 region’
Lildballe, D.L. et al.5
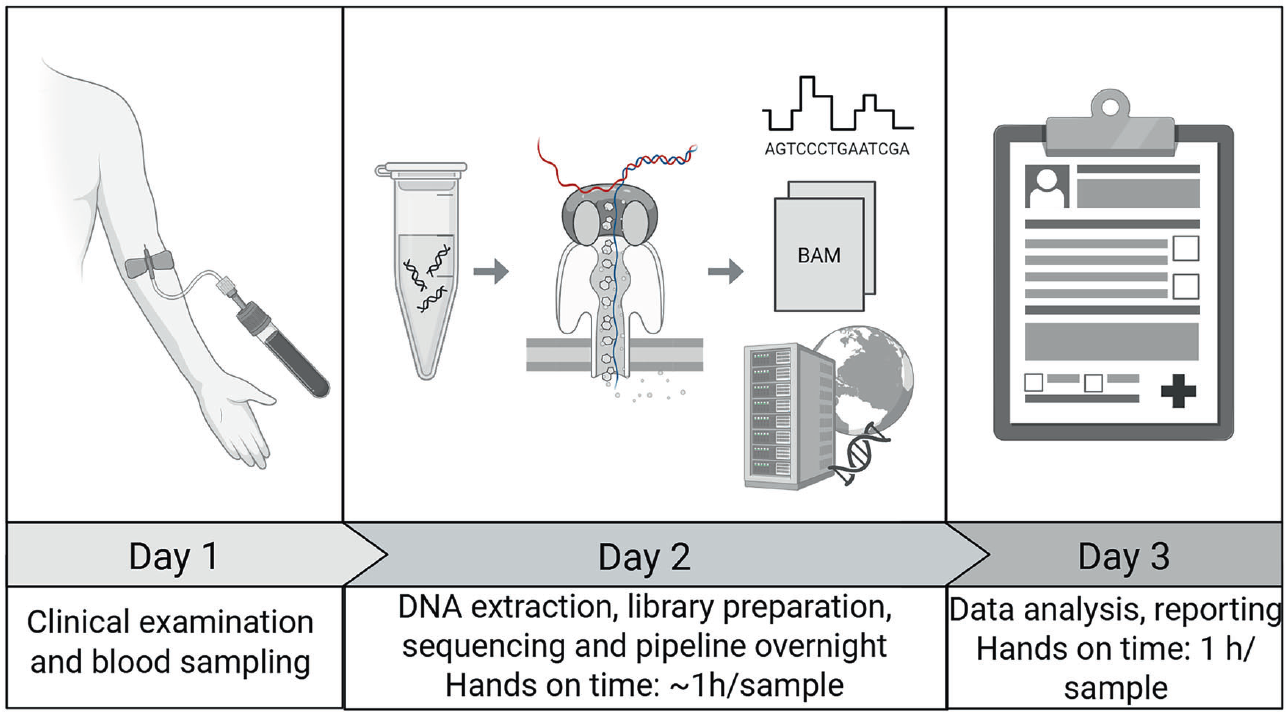
Figure 1: Graphical overview of the process from sample collection to clinical report using Adaptive Sampling for genetic confirmation of CAH. The figure is made in BioRender (Created in BioRender. Gravholt, C. 2026, https://BioRender.com/ddvzox9). Hands-on time per sample depends on the number of samples being processed in parallel. A batch of eight samples was processed in six hours, resulting in an average hands-on time of less than one hour per sample. Basecalling runs in parallel with sequencing, and the analysis pipeline is automatically started once sequencing and basecalling are complete. For comparison, Sanger sequencing and multiplex ligation-dependent probe amplification have a hands-on time of approximately 2.5 hours per sample. Figure redistributed from Lildballe et al under Creative Commons Attribution License CC BY 4.0.
Get clarity for rare disease, without complexity, with our getting started guide to human genomics.
What questions could you answer with powerful sequencing at any scale? Discover our range of nanopore sequencing solutions.
Oxford Nanopore Technologies products are not intended for use for health assessment or to diagnose, treat, mitigate, cure, or prevent any disease or condition.
- Cheng, H. and Qu, H. et al. Efficient near-telomere-to-telomere assembly of nanopore simplex reads. Nature Online ahead of print (2026). DOI: https://doi.org/10.1038/s41586-026-10105-6
- Bisogni, A.J. et al. Ultra-fast sample-to-sequencing workflow for clinical diagnostics using micropillars. MedRxiv 26345156 (2026). DOI: https://doi.org/10.64898/2026.01.29.26345156
- Zhang, L., Cui, J., and Shi, J. et al. Exon skipping as a potential diagnostic biomarker in colorectal cancer: an integrated epigenomic-transcriptomic analysis. Hum. Genomics Online ahead of print (2026). DOI: https://doi.org/10.1186/s40246-026-00931-0
- Bastug, K.A., Umar, U., Okolo, M., and Thielen, B.K. et al. Improving local diagnostic capacity for microbiological identification and antimicrobial resistance gene detection in Northern Nigeria using nanopore whole-genome sequencing. Access Microbiol. Online ahead of print (2026).
- Lildballe, D.L. et al. Genetic diagnosis of CYP21A2-related CAH: Adaptive Sampling long-read sequencing is an accurate and scalable solution. Eur. J. Hum. Genet. Online ahead of print (2026). DOI: https://doi.org/10.1038/s41431-026-02019-8






