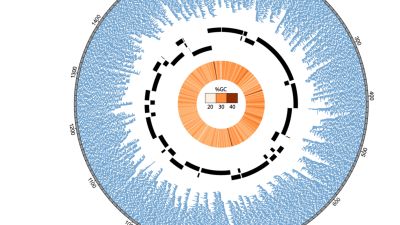
Scaffolding of a bacterial genome using MinION nanopore sequencing
- shared.published_on: April 7 2016
Second generation sequencing has revolutionized genomic studies. However, most genomes contain repeated DNA elements that are longer than the read lengths achievable with typical sequencers, so the genomic order of several generated contigs cannot be easily resolved. A new generation of sequencers offering substantially longer reads is emerging, notably the Pacific Biosciences (PacBio) RS II system and the MinION system, released in early 2014 by Oxford Nanopore Technologies through an early access program. The latter has highly advantageous portability and sequences samples by measuring changes in ionic current when single-stranded DNA molecules are translocated through nanopores. We show that the MinION system produces long reads with high mapability that can be used for scaffolding bacterial genomes, despite currently producing substantially higher error rates than PacBio reads. With further development we anticipate that MinION will be useful not only for assembling genomes, but also for rapid detection of organisms, potentially in the field.