NCM 2022: To make a short story long: simultaneous short and long RNA profiling on Oxford Nanopore devices
- shared.published_on: December 7 2022
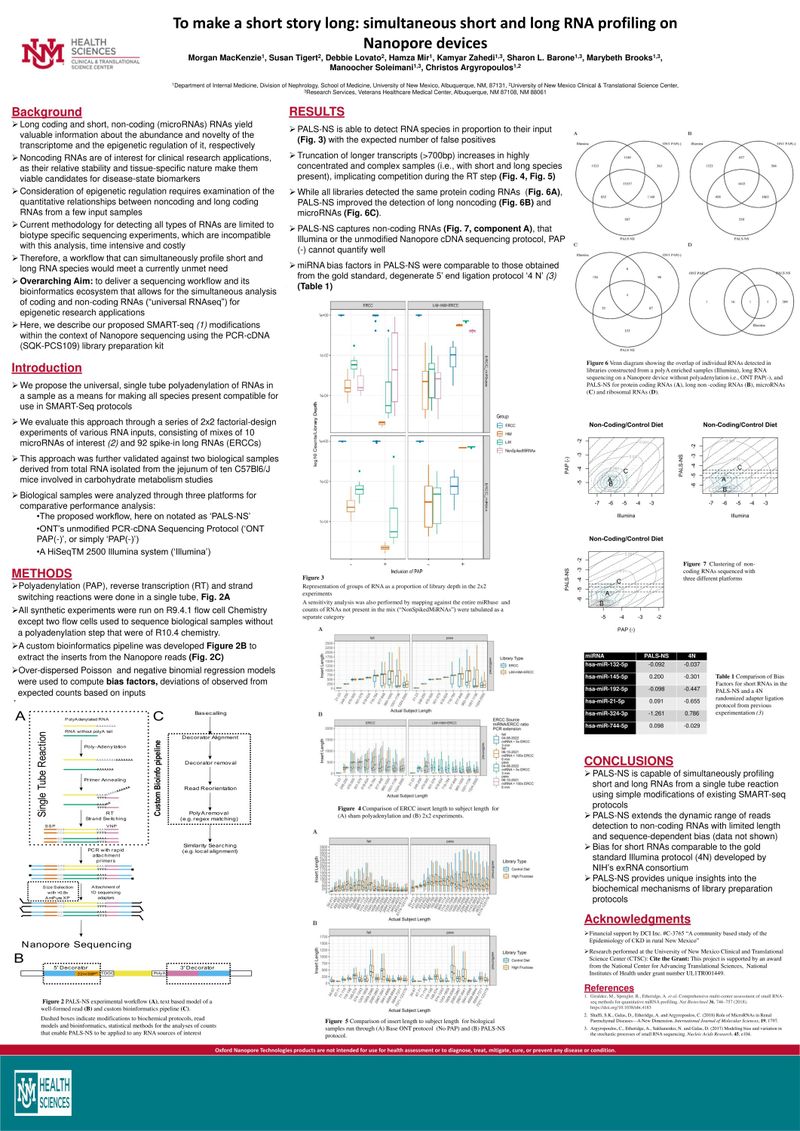
Long coding and short, non-coding (microRNAs) RNAs yield valuable information about the abundance and novelty of the transcriptome and the epigenetic regulation of it, respectively. Noncoding RNAs are of interest for clinical research applications, as their relative stability and tissue-specific nature make them viable candidates for disease-state biomarkers. Consideration of epigenetic regulation requires examination of the quantitative relationships between noncoding and long coding RNAs from a few input samples. Current methodology for detecting all types of RNAs are limited to biotype specific sequencing experiments, which are incompatible with this analysis, time intensive and costly.
Therefore, a workflow that can simultaneously profile short and long RNA species would meet a currently unmet need. Overarching Aim: to deliver a sequencing workflow and its bioinformatics ecosystem that allows for the simultaneous analysis of coding and non-coding RNA's universal for epigenetic research applications. Here, we describe our proposed SMART-seq modifications within the context of nanopore sequencing using the PCR-cDNA (SQK-PCS109) library preparation kit.






