Improving algorithms for amplicon sequencing
- shared.published_on: May 19 2023
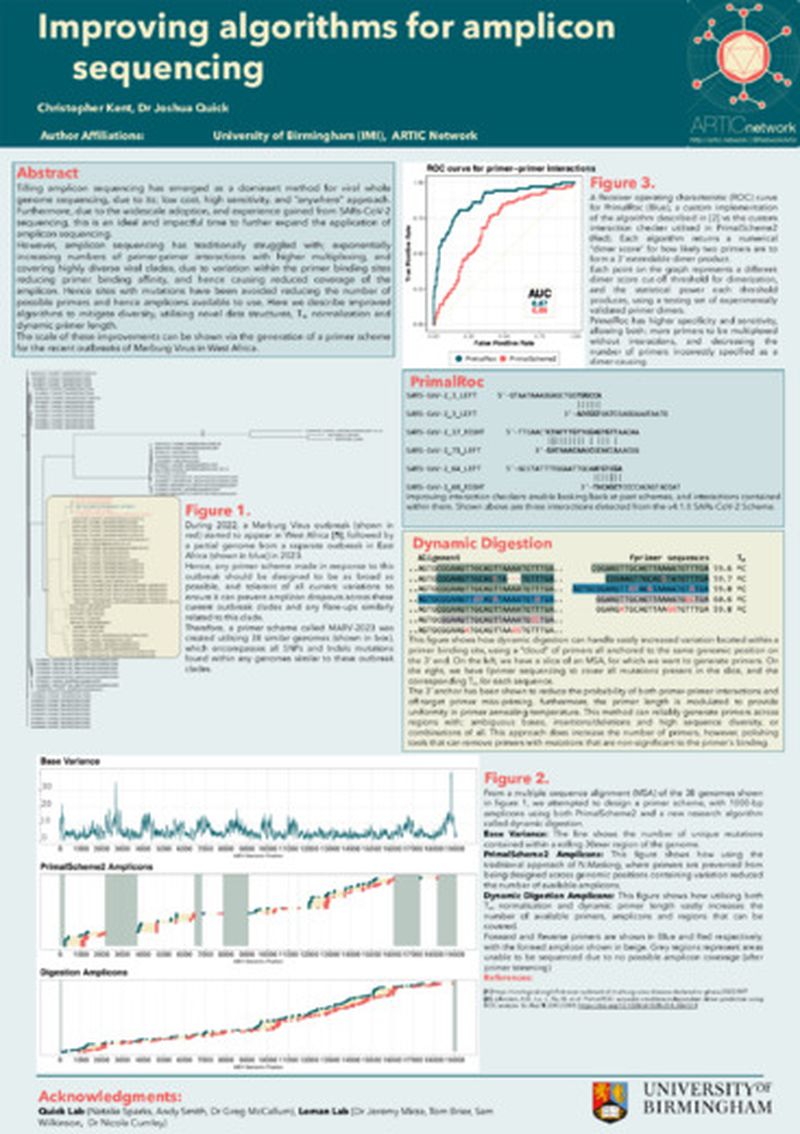
Tiling amplicon sequencing has emerged as a dominant method for viral whole genome sequencing, due to its; low cost, high sensitivity, and “anywhere” approach. Furthermore, due to the widescale adoption, and experience gained from SARs-CoV-2 sequencing, this is an ideal and impactful time to further expand the application of amplicon sequencing.
However, amplicon sequencing has traditionally struggled with; exponentially increasing numbers of primer-primer interactions with higher multiplexing, and covering highly diverse viral clades, due to variation within the primer binding sites reducing primer binding affinity, and hence causing reduced coverage of the amplicon. Hence sites with mutations have been avoided reducing the number of possible primers and hence amplicons available to use.
Here we describe improved algorithms to mitigate diversity, utilising novel data structures, Tm normalization and dynamic primer length.
The scale of these improvements can be shown via the generation of a primer scheme for the recent outbreaks of Marburg Virus in West Africa.






