Determining the fate of engineered tRNAs in human cells by direct RNA sequencing
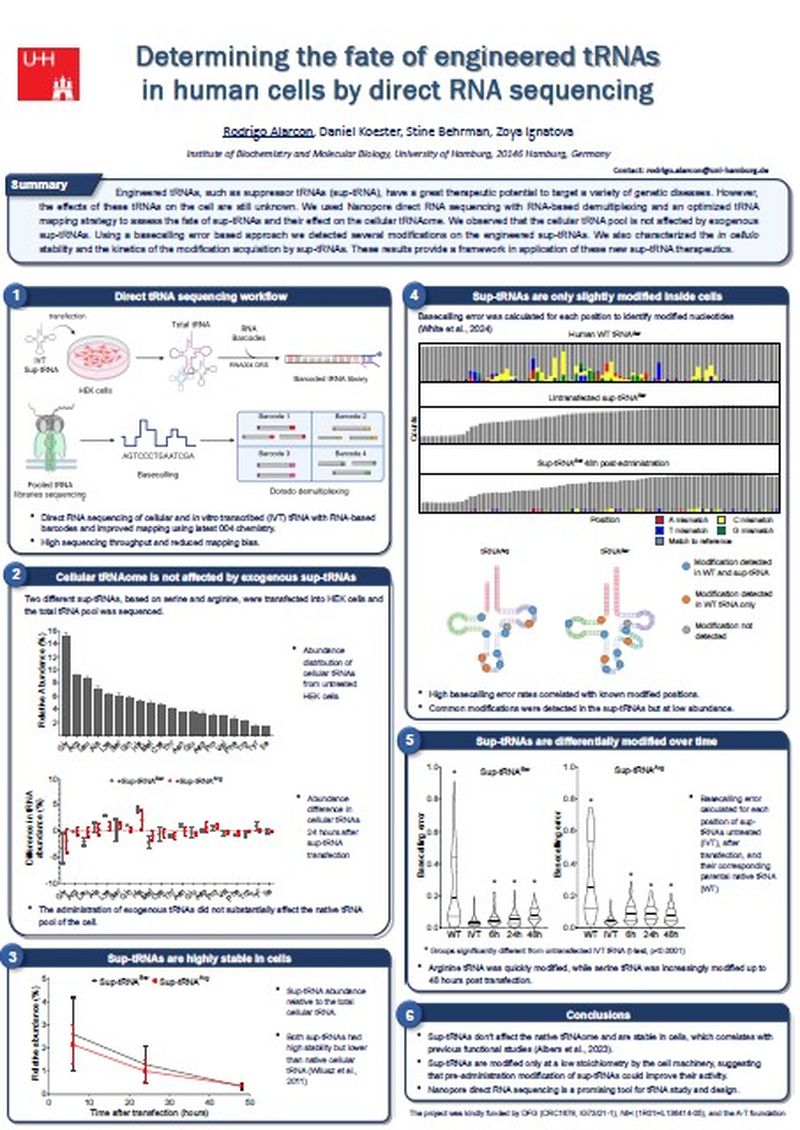
Engineered tRNAs designed to suppress nonsense mutations (sup-tRNAs) have a great therapeutic potential for a variety of genetic diseases. However, the effects of these tRNAs on the cellular translation machinery, particularly their impact on the native tRNA pool, remain unknown.
In this poster, researchers investigate:
- The use of Oxford Nanopore direct RNA sequencing of the tRNAome to study the effect of sup-tRNAs on them and the fate of these sup-tRNAs in human cells.
- How these results provide a framework in understanding the effect of these new sup-tRNA therapeutics.
- New workflows for tRNA analysis to expand the possibilities of RNA sequencing with Oxford Nanopore.







