Automated single-platform telomere-to-telomere de novo genome assembly for human, plant, and animal genomes
- shared.published_on: May 20 2025
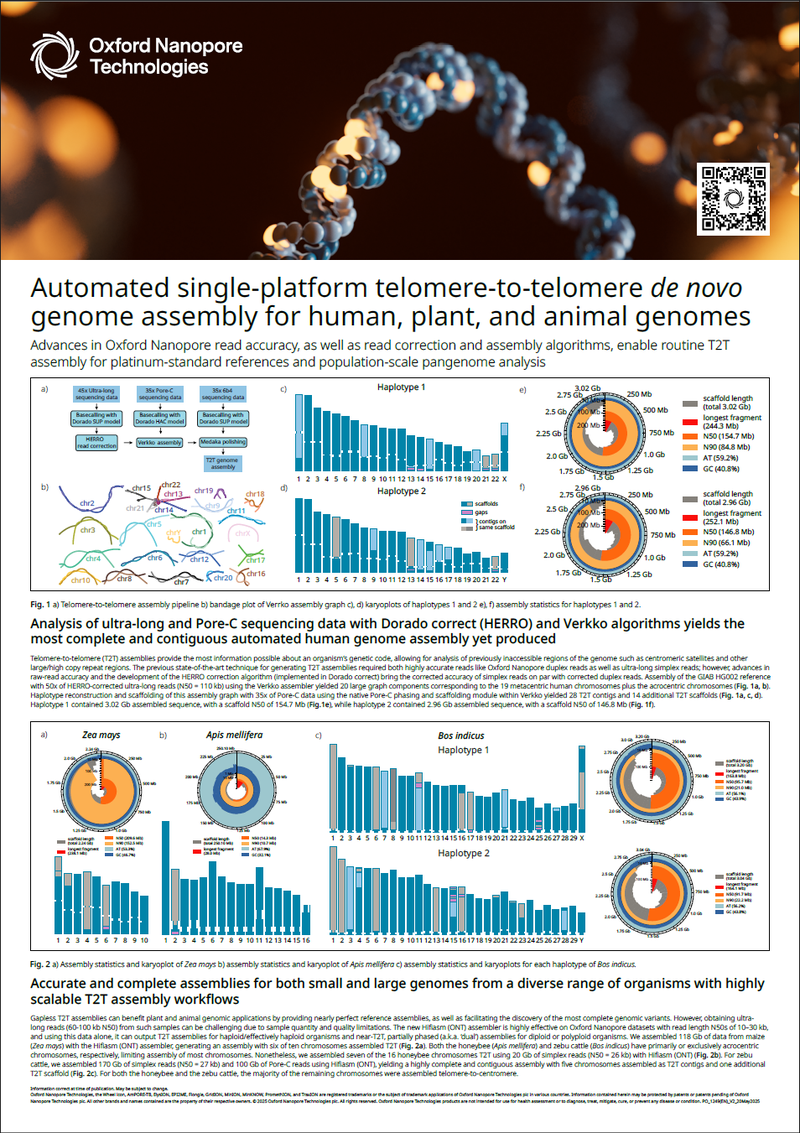
Advances in Oxford Nanopore read accuracy, as well as read correction and assembly algorithms, enable routine T2T assembly for platinum-standard references and population-scale pangenome analysis.
Download the poster to discover:
- That analysis of ultra-long and Pore-C sequencing data with the HERRO, Verkko, and GFase algorithms yields the most complete automated human genome assembly yet produced
- How to achieve accurate and complete assemblies for genomes of any size across a wide variety of organisms, using highly scalable telomere-to-telomere (T2T) assembly workflows







