16S vs shotgun nanopore sequencing detected different taxa in the same freshwater samples
- shared.published_on: May 19 2023
Problem:
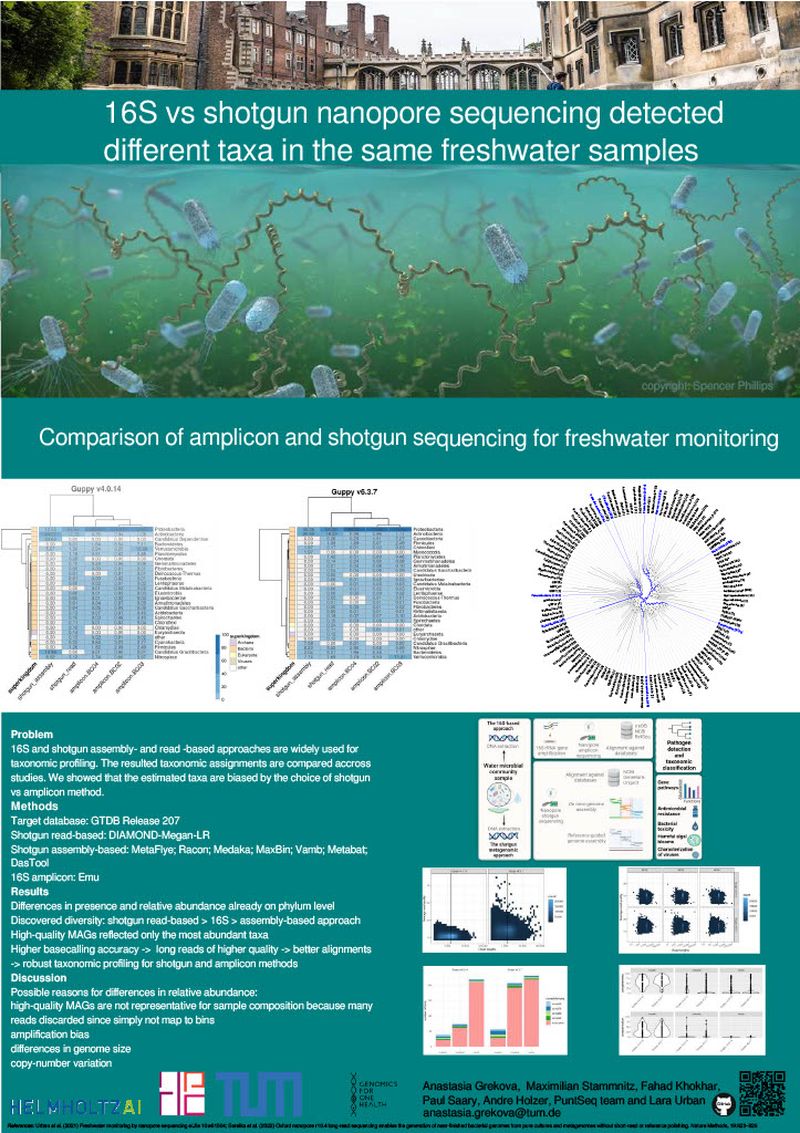
16S and shotgun assembly and read-based approaches are widely used for taxonomic profiling. The resulted taxonomic assignments are compared across studies. We showed that the estimated taxa are biased by the choice of shotgun vs amplicon method.
Methods:
- Target database: GTDB Release 207
- Shotgun read-based: DIAMOND-Megan-LR
- Shotgun assembly-based: MetaFlye; Racon; Medaka; MaxBin; Vamb; Metabat; DasTool
- 16S amplicon: Emu
Results:
Differences in presence and relative abundance already on phylum level Discovered diversity: shotgun read-based > 16S > assembly-based approach High-quality MAGs reflected only the most abundant taxa Higher basecalling accuracy -> long reads of higher quality -> better alignments -> robust taxonomic profiling for shotgun and amplicon methods.
Discussion:
Possible reasons for differences in relative abundance: high-quality MAGs are not representative for sample composition because many reads discarded since simply not map to bins amplification bias differences in genome size copy-number variation.






