Plasmid sequencing: revealing the whole picture, one base at a time
Plasmids are essential in molecular biology, yet they are prone to unexpected mutation. Legacy sequencing methods make whole-plasmid sequencing difficult, meaning that labs often only check the protein-coding regions they have edited. Even in recent years, challenges in capturing complete plasmid sequences have left researchers with incomplete information on what’s in their plasmids.
But technological advances are changing this.
What if you could sequence your entire plasmid in just a few hours, without amplification or primers?
In this Nanopore Know-How blog, we explore the history of plasmid verification and why whole-plasmid sequencing is becoming the new gold standard in labs.
The backbone of molecular biology
Plasmids are small, circular DNA molecules, typically found in bacteria1, separate from chromosomal DNA. As they are mobile genetic elements, they enable bacteria to evolve rapidly by sharing beneficial genes, such as those for antibiotic resistance2,3.
Beyond their evolutionary role in bacteria, plasmids have become a critical tool in molecular biology. Their ability to carry and express genes makes them foundational in many applications, including gene therapy, vaccine development, and genetic engineering4.
For example, in genetic diseases where a functional gene is missing or dysfunctional, researchers can insert a specific gene into a plasmid and deliver it to patient cells5. Researchers are actively investigating novel gene therapy approaches, such as using inhaled lentivirus as a potential treatment for patients with cystic fibrosis6.
In any application, ensuring that the plasmid sequence produced matches the planned design is essential. Any unexpected mutations can significantly impact biological outcomes, derail experiments, and invalidate therapeutic constructs7.
A brief history of plasmid sequencing
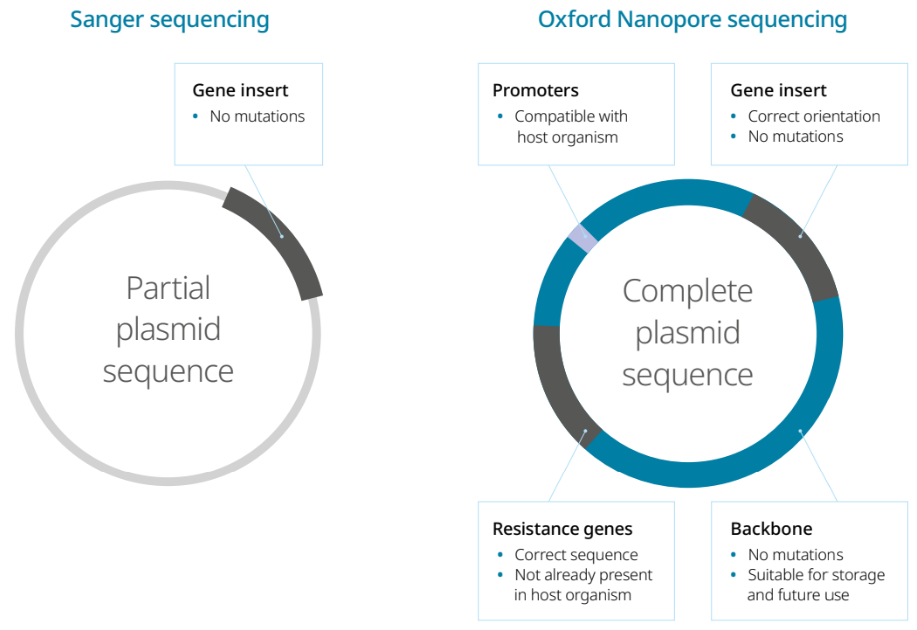
Sanger sequencing has long been the primary method for plasmid verification. While it is widely used, it has limitations, such as requiring PCR amplification and careful primer design; it also only reads sequences of approximately 900 bp at a time. This means that sequencing an entire plasmid requires multiple reactions, leading to extended time in the lab and higher costs4.
Sanger sequencing also struggles with repetitive or structurally complex regions, so researchers often need secondary validation methods to resolve these, such as restriction enzyme mapping and primer walking8. This complexity results in labs often only sequencing engineered sections and missing any mutations in the plasmid backbone4.
Short-read sequencing has been turned to for whole-plasmid sequencing, but it also relies on PCR amplification. As a result, this technique underrepresents GC-rich, low-complexity, and repetitive regions4. Consequently, even whole-plasmid short-read sequencing can still leave blind spots.
Now, Oxford Nanopore Technologies is challenging the norm by overcoming these difficulties.
Why researchers are choosing nanopore plasmid sequencing
Over the past few years, Oxford Nanopore sequencing has become a popular option for whole-plasmid sequencing, with service providers offering overnight plasmid verification. Our technology sequences native DNA of any length, so entire plasmids — not just protein-coding regions — can be read in single reads, without amplification or primers. Plus, it can resolve complex structural variants and is faster and cheaper than short-read sequencing methods4,8.
With advances in our sequencing accuracy, studies show that nanopore per-base error rates are significantly lower than Sanger sequencing8. Using the latest Oxford Nanopore chemistry, researchers are achieving 98.9% read-level accuracy and generating de novo assemblies with almost identical accuracy to those from short-read technology, without the need for a hybrid approach9.
This means that whole-plasmid verification can be performed quickly, simply, and accurately using nanopore data alone.
Raising the bar for plasmid quality
Accurate plasmid sequencing isn’t just good practice, it’s essential. Even single-base-pair changes can significantly alter gene expression or disrupt function, with costly consequences for downstream experiments7.
In a recent large-scale study by a cloning service provider, researchers revealed that plasmid errors are more common than we realise. The team estimated that 45–50% of lab-made plasmids submitted to the service provider contained undetected sequence or design errors10. These errors can arise during construction, propagation, or even storage, and without whole-plasmid sequencing, they often go unnoticed.
These findings highlight the need for robust quality control, as well as a high level of training in vector design and plasmid handling10. Plus, they underscore the importance of verifying the entire plasmid to ensure they are fully error-free, not just the sequences within engineered regions.
Redefining the gold standard
Oxford Nanopore sequencing is transforming plasmid verification by providing a complete view of plasmids in a single workflow. With our method, service labs are routinely uncovering errors that previously slipped under the radar, including mycoplasma contamination10.
Mark Budde, co-founder and CEO of Plasmidsaurus, noted in a presentation at the Nanopore Community Meeting 2023 that customers often receive unexpected results for their synthesised plasmids as mutations often occur during transfer. Whole-plasmid nanopore sequencing exposes these previously hidden mutations, enabling researchers to use their plasmids with confidence.
Nanopore sequencing is fast, accurate, and reveals the whole picture, making it an essential tool for plasmid-based research. Whether you use a service provider or run our plasmid workflow in your own lab, whole-plasmid sequencing is more accessible than ever.
Oxford Nanopore Technologies products are not intended for use for health assessment or to diagnose, treat, mitigate, cure, or prevent any disease or condition.
- National Human Genome Research Institute. Plasmid. https://www.genome.gov/genetics-glossary/Plasmid (2025) [Accessed 13 March 2026]
- Garcillán-Barcia, M.P. et al. The facts and family secrets of plasmids that replicate via the rolling-circle mechanism. Microbiol. Mol. Biol. Rev. 86(1):e00222–20 (2021). DOI: https://doi.org/10.1128/MMBR.00222-20
- Dewan, I. and Uecker, H. A mathematician’s guide to plasmids: an introduction to plasmid biology for modellers. Microbiology (Reading) 169(7):001362 (2023). DOI: https://doi.org/10.1099/mic.0.001362
- Mumm, C. et al. Multiplexed long-read plasmid validation and analysis using OnRamp. Genome Res. 33(5):741–749 (2023). DOI: https://doi.org/10.1101/gr.277369.122
- Bulcha, J.T. and Wang, Y. et al. Viral vector platforms within the gene therapy landscape. Signal Transduct. Target. Ther. 6(1):53 (2021). DOI: https://doi.org/10.1038/s41392-021-00487-6
- Davies, J.C. and Polineni, D. et al. Lentiviral gene therapy for cystic fibrosis: a promising approach and first-in-human trial. Am. J. Respir. Crit. Care Med. 210(12):1398–1408 (2024). DOI: https://doi.org/10.1164/rccm.202402-0389CI
- Hernandez, S.I. et al. Sequencing strategy to ensure accurate plasmid assembly. bioRxiv 586694 (2024). DOI: https://doi.org/10.1101/2024.03.25.586694
- Emiliani, F., Hsu, I., and McKenna, A. Multiplexed assembly and annotation of synthetic biology constructs using long-read nanopore sequencing. ACS Synth. Biol. 11(7):2238–2246 (2022). DOI: https://doi.org/10.1021/acssynbio.2c00126
- Zhao, W. et al. Oxford Nanopore long-read sequencing enables the generation of complete bacterial and plasmid genomes without short-read sequencing. Front. Microbiol. 14:1179966 (2023). DOI: https://doi.org/10.3389/fmicb.2023.1179966
- Bai, X. et al. Prevalence of errors in lab-made plasmids across the globe. bioRxiv 596931 (2025). DOI: https://doi.org/10.1101/2024.06.17.596931