What is ORG.one?
ORG.one is a project designed to support equitable, faster, and more localised sequencing of endangered species. It enables biologists to rapidly study those species close to the sample’s origin using the latest sequencing approaches. For the project to have the greatest impact, the data generated will be uploaded to an open-access database, to ensure anyone, anywhere can use it to improve and guide their conservation efforts.
Existing projects to sequence threatened species have made good progress, however they can be limited by complex workflows and the requirement to send samples to centralised, often overseas, locations for sequencing.
With its partners, ORG.one is focusing on delivering and demonstrating the impactful benefits of genomics for the conservation of endangered organisms. ORG.one aims to develop an open database of the most comprehensive high quality de novo draft assemblies of genomes to support the conservation of as many endangered species as possible.


How it works
The first step is to register your interest. Accepted participants of the program will receive free-of-charge consumables and support, to enable them to sequence eligible species, classified as Endangered, Critically Endangered or Extinct in the Wild by the IUCN Red List, with the latest Oxford Nanopore tools and methods. Sequencing should be conducted within the species’ country of origin, wherever possible, and the data generated uploaded to the EMBL-EBI ENA open public database within 6 weeks. Researchers must be responsible for any local legal and ethical permits to generate the data and to share or export it. Finally, we stipulate that no animals are harmed or killed for their DNA.
Who can take part?
ORG.one is open to anyone. The project encourages participants of all levels of resources and expertise to join.
Often those with access to samples of endangered species do not have the access, capacity, or required knowledge to sequence them. Those with sequencing capacity may not have the bioinformatic expertise to use their sequencing data to generate a high-quality reference genome. Finally, even if a high-quality reference genome is available, many conservationists are not aware of the potential benefits it can bring to help protect their species of interest.
ORG.one aims to help by developing an active and distributed community of conservationists, scientists, bioinformaticians, and other stakeholders to support the generation and communication of open data for conservation genomics, as well as ensuring local community involvement so those communities can leverage the research to their own advantage.
How does conservation genomics help?
Genomic data can play a pivotal role in addressing key biological questions that are crucial for conserving endangered species. It can be used to inform wide ranging questions from taxonomy and population connectivity to strategies for mitigating disease and the effects of climate change. By generating and providing access to reference data for species facing the highest risk of extinction, ORG.one empowers researchers to conduct impactful studies that enhance conservation initiatives.
Why is research at source so important?
Conservation biology is dependent on community support and local participation. Without it, efforts to protect endangered species have little or no impact.
When local communities are engaged from the beginning, they can share their wealth of local knowledge, as well as have open conversations about their needs and values. Open dialog can help to ensure that the community’s needs are met whilst also discussing with the community the possible benefits, opportunities, and skills (such as ecotourism or learning bioinformatics) that could arise from being involved in conservation. In the long term, communities that have invested in conservation and are seeing the benefits come to fruition are much less likely to revert to practices that negatively impact local conservation efforts.
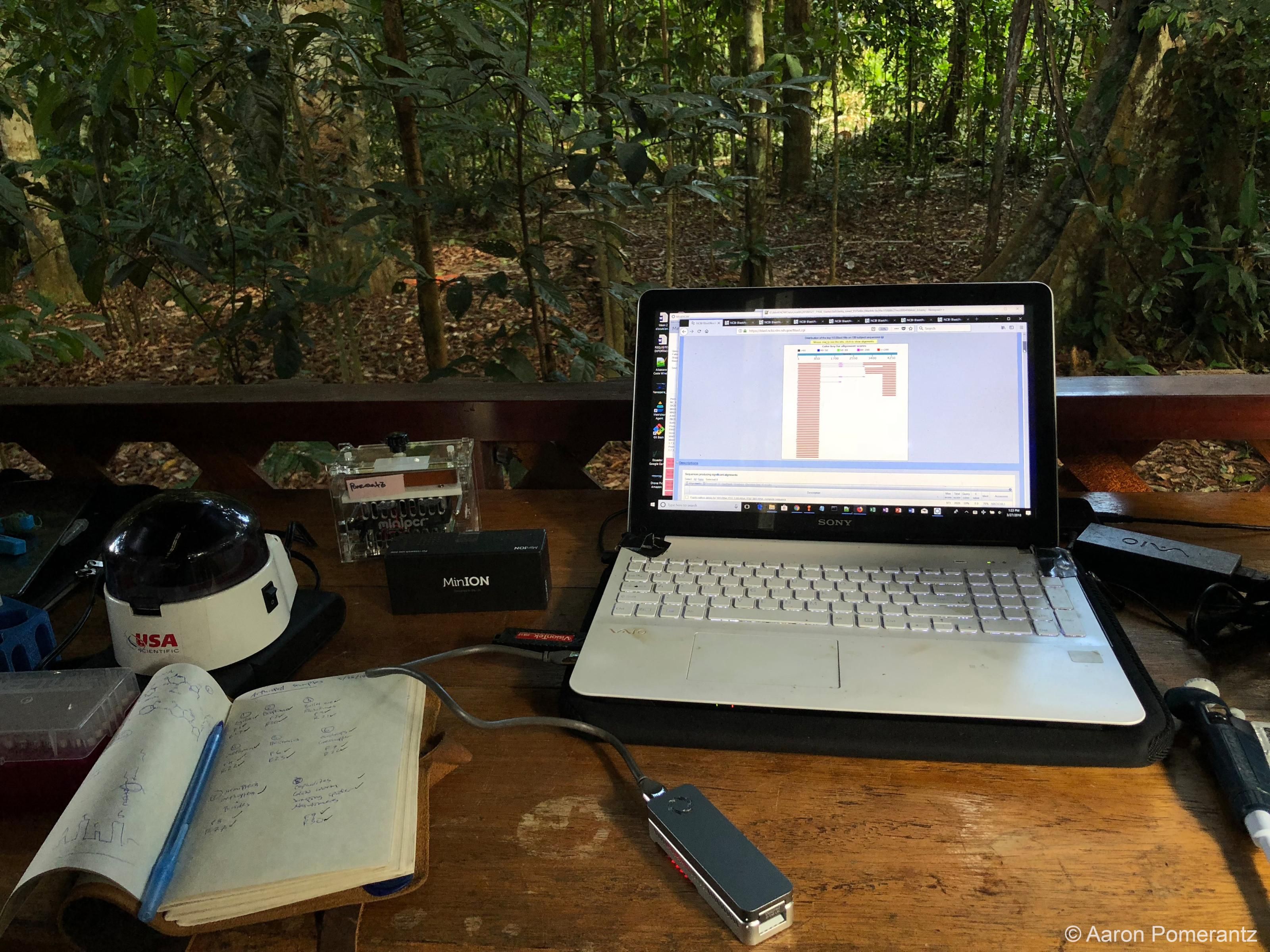
This is why, where possible, ORG.one aims to perform the sequencing in situ and encourages participants from all levels of resources to contribute.
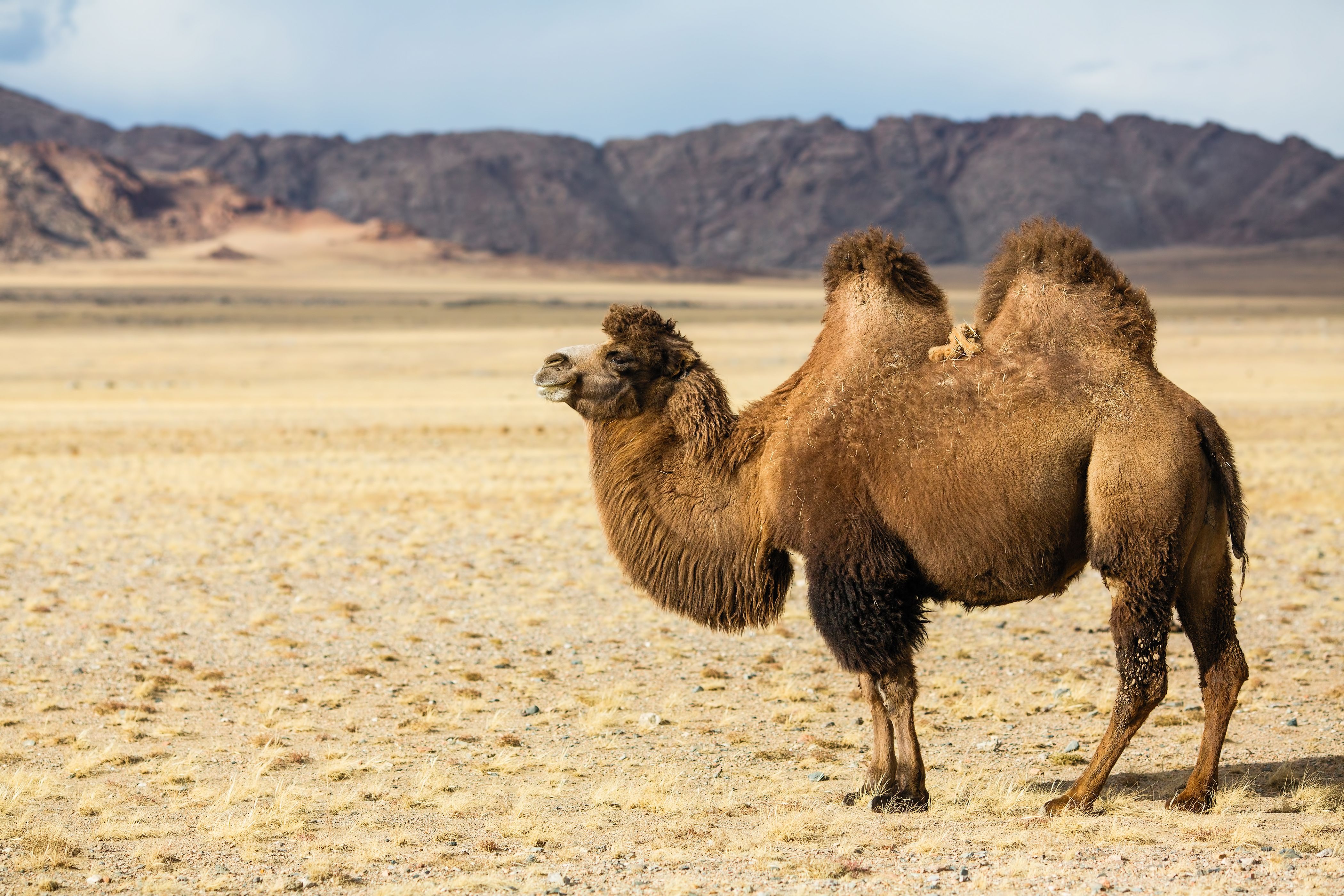


Completed projects
Over 150 species have been sequenced by ORG.one participants!
Case studies
Check out several success stories already part of ORG.one.
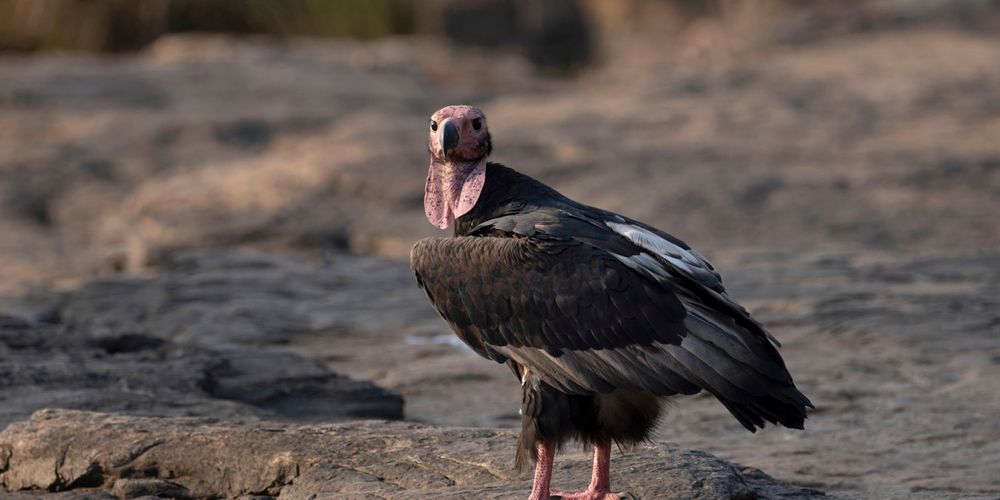

Asian king vulture (Sarcogyps calvus)
Researchers: Gunnaporn Suriyaphol Location: Thailand
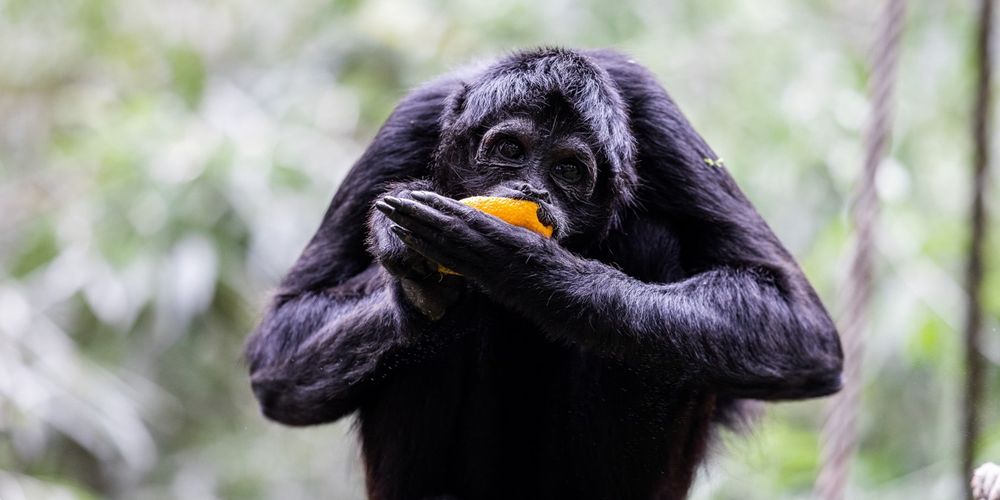

Brown-headed spider monkey (Ateles fusciceps fusciceps)
Researchers: Gabriela Pozo Andrade Location: Ecuador
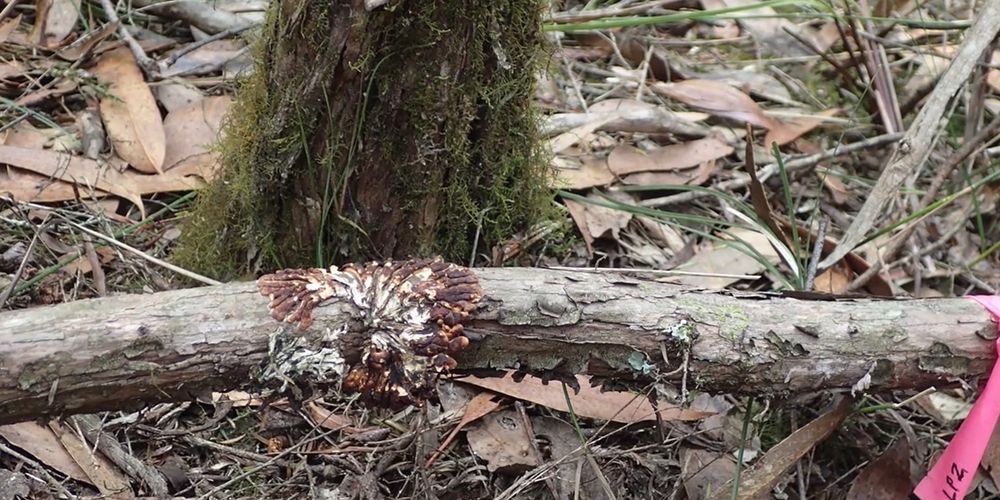

Tea-tree Fingers (Hypocreopsis amplectens)
Researchers: Sapphire McMullan-Fisher & Michael Amor Location: Australia
Our partners
ORG.one was set up by Oxford Nanopore Technologies, the maker of scalable, real-time DNA/RNA sequencing devices. We also have a number of partners across the globe who are helping us to support the rapid sequencing of endangered species, anywhere, by anyone.
